Biomod/2011/TUM/TNT/Project/Theory
<html>
<style>
- column-one { display:none; width:0px;}
.container{background-color: #f5f5f5; margin-top:50px} .OWWNBcpCurrentDateFilled {display: none;}
- content { width: 0px; margin: 0 auto auto 0; padding: 1em 1em 1em 1em; align: center;}
- column-content {width: 0px; float: left; margin: 0 0 0 0;padding: 0;}
.firstHeading {display:none; width:0px;}
- globalWrapper{ width:1280px;}
div.tleft { border-bottom-width: 0em; border-left-width: 0px; border-right-width: 0em; border-top-width: 0em; clear: left; float: left; margin-right: 0em;
}
div.tnone { border-bottom-width: 0em; border-left-width: 0px; border-right-width: 0em; border-top-width: 0em; }
- column-one {display:none; width:0px;background-color: #ffffff;}
- content{ margin: 0 0 0 0; padding: 1em 1em 1em 1em; position: center; width: 800px;background-color: #f5f5f5; }
.container{ width: 800px; margin: auto; background-color: #ffffff; text-align:justify; font-family: helvetica, arial, sans-serif; color:#555555; margin-top:25px; }
- bodyContent{ width: 1267px; align: center; background-color: #ffffff;}
- column-content{width: 1280px;background-color: #f5f5f5;}
.firstHeading { display:none;width:0px;background-color: #ffffff;}
- header{position: center; width: 800px;background-color: #ffffff;}
- footer{position: center;}
div.thumb { border-bottom-color: rgb(255, 255, 255); border-bottom-style: solid; border-left-color: rgb(255, 255, 255); border-left-style: solid; border-right-color: rgb(255, 255, 255); border-right-style: solid; border-top-color: rgb(255, 255, 255); border-top-style: solid; margin-bottom: 0em; width: auto;
}
div.tright { border-bottom-width: 0em; border-left-width: 0px; border-right-width: 0px; border-top-width: 0em; clear: right; float: right;
}
body {
font-family: helvetica, arial, sans-serif; color: black;
background:#f5f5f5;
}
ul#topnav {
margin: 0 0 0 0px ; padding: 0 0 0 0px; float: left; width: 800px; list-style: none; position: relative;
font-family: helvetica, arial, sans-serif;
font-size: 12px; background: #f5f5f5;
height: 5px;
} ul#topnav li { float: left; margin: 0; padding: 0; border-right: 1px solid #ffffff; /*--Divider for each parent level links--*/ } ul#topnav li a { padding: 11px 15px; display: block; color: #333333; text-decoration: none; } ul#topnav li:hover { background: #f5f5f5;} /*--Notice the hover color is on the list item itself, not on the link. This is so it can stay highlighted even when hovering over the subnav--*/
ul#topnav li span { float: left; padding: 15px 0; position: absolute; left: 0; top:30px; display: inline; /*--Hide by default--*/ width: 800px; background: #cccccc; color: #333333; /*--Bottom right rounded corner--*/ -moz-border-radius-bottomright: 5px; -khtml-border-radius-bottomright: 5px; -webkit-border-bottom-right-radius: 5px; /*--Bottom left rounded corner--*/ -moz-border-radius-bottomleft: 5px; -khtml-border-radius-bottomleft: 5px; -webkit-border-bottom-left-radius: 5px; } ul#topnav li:hover span { display: block; } /*--Show subnav on hover--*/ ul#topnav li span a { display: inline;} /*--Since we declared a link style on the parent list link, we will correct it back to its original state--*/ ul#topnav li span a:hover {text-decoration: underline;}
- abstractandlinks{
display:block; width:800px; background-color:#f5f5f5; text-align:justify;font-family: helvetica, arial, sans-serif; color:#555555; margin-top:0px; }
- innerBox
{ display: block; padding: 5px; }
- abstract{
font-family: helvetica, arial, sans-serif; width: 490px; margin-top: 20px; padding:0px 5px; float: left;background-color:#f5f5f5; ; }
- links{
font-family: helvetica, arial, sans-serif; width: 290px; margin-top: 20px; padding:0px 5px; float: left; background-color:#f5f5f5;; } .clear { clear:both; }
- tour{
font-family: helvetica, arial, sans-serif; width: 790px; padding:0px 5px; margin-top: 1em, margin-bottom: 1em; float: left;background:#f5f5f5; font-size: 2em; }
- struktur{
font-family: helvetica, arial, sans-serif; width: 790px; padding:0px 5px; float: left;background:#f5f5f5; }
- toctitle {align: left; text-align: left; }
- table.toc {background-color:"#5f5f5f";align: left; text-align: left;}
- toc #toctitle, .toc #toctitle, #toc .toctitle, .toc .toctitle {
text-align: left;
}
- toc, .toc, .mw-warning {
background-color: #f5f5f5; border-bottom-color: rgb(170, 170, 170); border-bottom-style: solid; border-bottom-width: 0px; border-left-color: rgb(170, 170, 170); border-left-style: solid; border-left-width: 0px; border-right-color: rgb(170, 170, 170); border-right-style: solid; border-right-width: 0px; border-top-color: rgb(170, 170, 170); border-top-style: solid; border-top-width: 0px; font-size: 105%; padding-bottom: 5px; padding-left: 5px; padding-right: 5px; padding-top: 5px; align: left; text-align: left; display: left
}
</style>
<head>
<script type="text/javascript"> var fullDates = new Array('08/11/2011','08/12/2011','08/16/2011','08/17/2011','08/23/2011','08/24/2011','08/25/2011','08/29/2011','08/30/2011','08/31/2011','09/01/2011','09/02/2011','09/06/2011','09/12/2011','09/13/2011','09/14/2011','09/15/2011','09/16/2011','09/19/2011','09/20/2011','09/21/2011','09/27/2011','09/28/2011','09/29/2011','09/30/2011','10/05/2011','10/06/2011','10/07/2011','10/10/2011','10/12/2011','10/13/2011','10/14/2011','10/17/2011','10/18/2011','10/19/2011','10/20/2011','10/21/2011','10/26/2011','10/27/2011','10/28/2011','10/30/2011');
$(document).ready(function() {
$("ul#topnav li").hover(function() { //Hover over event on list item $(this).css({ 'background' : '#cccccc'}); //Add background color and image on hovered list item $(this).find("span").show(); //Show the subnav
} ,
$("ul#topnav li").oneclick(function() { //Hover over event on list item $(this).css({ 'background' : '#cccccc'; 'display' : 'inline'}); //Add background color and image on hovered list item $(this).find("span").show(); //Show the subnav
} ,
function() { //on hover out...
$(this).css({ 'background' : 'none'}); //Ditch the background
//$(this).find("span").hide(); //Hide the subnav });
}); </script>
<script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.3/jquery.min.js"></script>
<meta http-equiv="Content-Type" content="text/html; charset=utf-8" />
<script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.4.2/jquery.min.js"></script> <script type="text/javascript"> $(document).ready(function(){ $('#lside1').mouseover(function() { $('#lside1').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside1').mouseleave(function() { $('#lside1').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); $('#lside2').mouseover(function() { $('#lside2').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside2').mouseleave(function() { $('#lside2').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); $('#lside3').mouseover(function() { $('#lside3').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside3').mouseleave(function() { $('#lside3').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); }); </script>
<style type="text/css">
#side1 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside1 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
#side2 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside2 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
#side3 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside3 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
</style>
<title>
TUM NanU - Home
</title> </head>
<body background-color="#F0F0F0">
</body> </html>
Theoretical Background
http://openwetware.org/index.php?title=Biomod/2011/TUM/TNT/Project/Theory&action=edit
Twist of the arms
Theoretical considerations

The origin of z is located at the bottoms of the arms where the base ends as depicted in the figure to the left. The angle [math]\displaystyle{ \alpha }[/math] of the torsion of the arms shows a linear dependency on z [math]\displaystyle{ \alpha_0 }[/math] defines a position in the undeformed state, [math]\displaystyle{ \Delta \alpha + \alpha_0 }[/math] is the maximum torsion at the end of the arms, B is the length of the base and L is the total length of the structure):
[math]\displaystyle{ \alpha (z) = \frac{z}{L-B} \cdot \Delta\alpha + \alpha_0 }[/math]
The distance between two points on the mantle of the cylinders that describe the arms can be easily derived by vector analysis and results in:
[math]\displaystyle{ d = 2 \cdot \sqrt{R^2 + \left( \frac{D}{2} \right)^2 - R \cdot D \cdot cos \left( \alpha (z) \right)} }[/math]
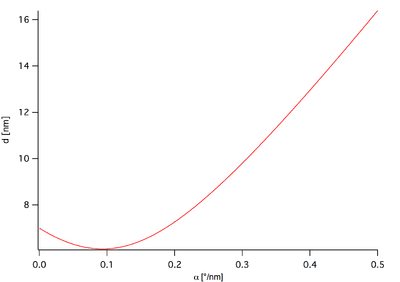
The following diagram shows a relation between [math]\displaystyle{ \Delta\alpha }[/math] and d for R=4nm, D=13nm and [math]\displaystyle{ \alpha_0 }[/math]=0°.

Observing at TEM pictures
In TEM images a dark line can be observed between the two six helix buldles of each arm. If the arms are twisted, this line is no longer exactly in the middle of one arm but "moves around" it. There should also be a variation in the visualized width of the arms due to the asymetric cross-sectional area.


Twist of the base
Theoretical considerations

To transform the measured arm-twist into a deformation of the structure's base one can assume "the U" as tree cylinders of the same radius. Two of length L (the arms) and one of length B (the middle part of the base). To describe the deformation another cylinder of radius R = radius of the base-cylinder + radius of one arm cylinder and length B is constructed in a way that the center-lines of both arm cylinders are located on its mantle. The torsion of this cylinder (shown in red on the image above) can be directly related to the arm twist.
[math]\displaystyle{ \alpha }[/math] describes the torsion and is divided into [math]\displaystyle{ \alpha = \beta + \gamma }[/math] to include the different projections of the structure on the grid within the theory. The angle [math]\displaystyle{ 2\delta }[/math] between the arms is related to an angle [math]\displaystyle{ \phi }[/math] due to an easier measurement (tree characteristic points are defined as shown in the following picture, one in the middle of the end of the base and one in the middle of the end of each arm)

[math]\displaystyle{ ( L- x ) sin \delta = L sin \frac{\phi}{2} }[/math]
[math]\displaystyle{ x = B \frac{sin \beta}{sin \beta + sin ( \alpha - \beta )} }[/math]
The correlation between [math]\displaystyle{ \delta }[/math] and [math]\displaystyle{ \alpha }[/math] including the different projections described by [math]\displaystyle{ \beta }[/math] is:
[math]\displaystyle{ sin \beta + sin(\alpha - \beta) = \frac{B}{R} sin \delta }[/math]
[math]\displaystyle{ L ( sin \beta + sin (\alpha - \beta )) - B sin \beta = \frac{B L}{R} sin \frac{\phi}{2} }[/math]
The following equation is the arithmetic average of [math]\displaystyle{ L ( sin \beta + sin (\alpha - \beta )) - B sin \beta }[/math] for [math]\displaystyle{ 0 \lt \beta \lt \alpha }[/math]:
[math]\displaystyle{ \frac{1}{\alpha} \int_0^{\alpha} L ( sin \beta + sin (\alpha - \beta )) B\beta = \frac{( cos \alpha -1 ) ( B - 2 L )}{\alpha} }[/math]
[math]\displaystyle{ \frac{cos \alpha -1}{\alpha} = \frac{B L}{R (B - 2 L)} sin \frac{\phi}{2} }[/math]
This equation relates [math]\displaystyle{ \alpha }[/math] and [math]\displaystyle{ \phi }[/math] for an average projection.

This graph shows the result plotted for radius R=6nm, base length B=35nm and total length L=98nm. For this calculation [math]\displaystyle{ \phi }[/math] was calculated for [math]\displaystyle{ \alpha }[/math] from 0 to 133° because of the maximum of [math]\displaystyle{ \epsilon }[/math] at this angle. It can be shown in TEM images that the maximum torsion is below this maximum angle [math]\displaystyle{ \alpha }[/math] by observing the change of the base width. The maximum change refers to a torsion of 180° and can easily be recognized by thinner base. However, none of the structures examined show a big change of base width.
Observing at TEM pictures
The twist of the base can be observed at structures that are shown from the side. The twist of the base (in this case the middle part of the base) can be related to a speading of the arms. The following picture shows a structure in a control measurement of BM2 without DNA binders on the left and a twisted structure of the reference BM24 that is designed as a twisted structure on the right.

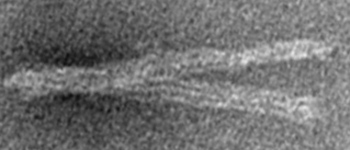
Total twist
The calculations of the base twist result in a modified D for the arm twist. Focusing on the base twist theory the distance between two points on the center lines of the arm cylinders can be described by:
[math]\displaystyle{ D(z) = 2 \cdot R_B \cdot \left( 1 + \frac{z}{B} \right) \cdot \sqrt{2-2 \cdot cos (\alpha_B)} }[/math]
Now the arm twist theory can be applied using D(z) instead of the fix value D:
[math]\displaystyle{ d = 2 \cdot \sqrt{{R_A}^2 + \left( \frac{D(z)}{2} \right)^2 - R_A \cdot D(z) \cdot cos \left( \alpha_A (z) \right)} }[/math]
The angles for base and arm twist can be normalized to [math]\displaystyle{ \alpha }[/math] as an angle per length: [math]\displaystyle{ \alpha = \frac{\alpha_A}{L-B} = \frac{\alpha_B}{B} }[/math]. [math]\displaystyle{ R_B }[/math] is half the distance between the center lines of the arms at the base and [math]\displaystyle{ R_A }[/math] is the distance of the cylinder center to a point on the cylinder mantle in x-y-direction.
[math]\displaystyle{ D(z) = 2 \cdot R_B \cdot \left( 1 + \frac{z}{B} \right) \cdot \sqrt{2-2 \cdot cos (\alpha \cdot B)} }[/math]
[math]\displaystyle{ d = 2 \cdot \sqrt{{R_A}^2 + \left( \frac{D(z)}{2} \right)^2 - R_A \cdot D(z) \cdot cos \left( z \cdot \alpha + \alpha_0 \right)} }[/math]
The graph on the left corresponds to FRET-pair positions C1→C2 as described in the article about FRET-pair positions and to the structure BM14_5/20.
The parameters describing the FRET-pair position are:
- [math]\displaystyle{ R_A }[/math] = 3.5nm
- [math]\displaystyle{ \alpha_0 }[/math] = 17°
- z = 55nm
- B = 35nm.
For a Förster-radius [math]\displaystyle{ R_0 }[/math] = 6.5 nm a maximum distance [math]\displaystyle{ d = \sqrt[3]{3} \cdot R_0 \approx 9.4nm }[/math] can be evaluated. This corresponds to a maximum angle of 0.3° per nm or a maximum torsion of 28° over the whole length of the structure.
Thermal fluctuation of the arms
Introduction
In order to estimate how broad a broad spread of the data points around the mean angle in TEM angle measurements must be expected, we calculated the width of a Boltzmann distributed fluctuation of the arms. Hereby we assumed the DNA helices to be cylinders with a certain stretch modulus E, persistence length p and to fluctuate independently. Within these measurements we always quantified the angle between the end points of the arms end the other end (at the base).
Derivation
Fluctuation of one arm
The probability of finding a rod with a 10 helix bundle cross section at a certain angle [math]\displaystyle{ \phi }[/math] is given by a Boltzmann distribution
[math]\displaystyle{ p(\phi - \phi_0) = \frac{1}{Z} e^{- \frac{E(\phi - \phi_0)}{k_B T}} }[/math]
where the energy is given by
[math]\displaystyle{ E(r ) = \frac{1}{2} \frac{E_{DNA} I_{10 hb} L}{r^2} }[/math]
with r the radius of curvature, L the contour length of the arms and I10 hb the area moment of inertia of a 10 helix bundle. If we assume a constant curvature of the arms, the radius of curvature can be translated in an angle [math]\displaystyle{ \phi }[/math] (see Fig. 1).

[math]\displaystyle{ 2 \phi = 2π \frac{L}{2\pi r} }[/math]
[math]\displaystyle{ E(\phi- \phi_0) = \frac{1}{2} \frac{4 E_{DNA} I_{10 hb} (\phi - \phi_0)^2}{ L } }[/math]
The area moment of inertia of a rod is given by
[math]\displaystyle{ I = \frac{R^4}{4} \pi }[/math]
Now we're able to calculate the area moment of inertia of the 10 helix bundle with the parallel axis theorem (see Fig. 10).
[math]\displaystyle{ I = \sum_n (I_n + {z_n}^2 A_n) = I_{DNA} (10 + 320 sin (\frac{\pi}{6})) }[/math]

and therefore the energy needed to bend one arm by the angle [math]\displaystyle{ \phi }[/math] as well as the probability.
Fluctuation of the measured angles
Since the fluctuation of the arms is stochastically independent, we have to take the convolution of the two probabilities:
[math]\displaystyle{ p(\Delta\phi) = \int p_1 (\phi_1 ) p_2 (\Delta\phi - \phi_1) d\phi_1 }[/math]
with [math]\displaystyle{ \Delta\phi = \phi_2 - \phi_1 }[/math]
With all this follows
[math]\displaystyle{ p(\Delta\phi) = \frac{1}{Z^2} \sqrt{\frac{π k_B T}{E_{eff}}} e^{-\frac{E_{eff}}{2 k_B T}(\Delta \phi - \phi_0)^2} }[/math]
with
[math]\displaystyle{ E_{eff} = \frac{2 E_{DNA} I_{DNA} (10 + 320 sin(\frac{\pi}{6}))}{L} }[/math]
Therefore we get a standard deviation of the measured angles of
[math]\displaystyle{ \sigma = \sqrt{\frac{k_B T}{E_{eff}}} = \sqrt{\frac{L}{2 p (10 + 320 sin(\frac{\pi}{6}))}} }[/math]