Biomod/2011/MIT/Origami/Tat Protein
From OpenWetWare
Jump to navigationJump to search

Geometry, Structure and Location
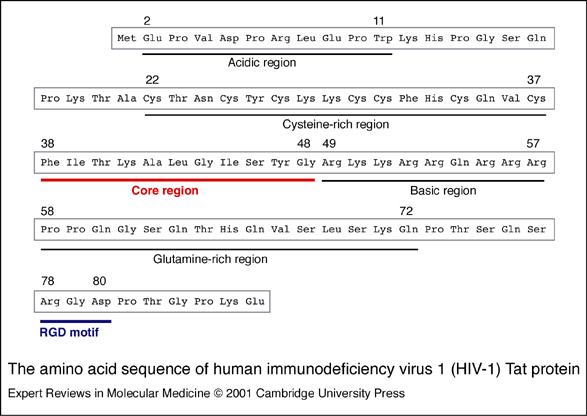
- The tat protein is composed of an N-terminal, a cysteine-rich hydrophobic core, a basic region, a glutamine-rich region, and a C-terminal.
- The 16-amino acid core region is strictly conserved.
- It is expressed very early in the infection of T cells from multiply spliced viral mRNAs and is then localized to the nucleus to play an active role in the reverse transcription complex.
- The protein later leaves the cell and has several roles in HIV pathogenesis, as will be discussed later.
- The protein folds such that the N-terminal, from Val to Pro, is in the center of the molecule, interacting with the core, basic, and G-rich regions.
- The cysteine-rich and basic regions are highly flexible.
Role in Reverse Transcription Complex of HIV-1 Virus
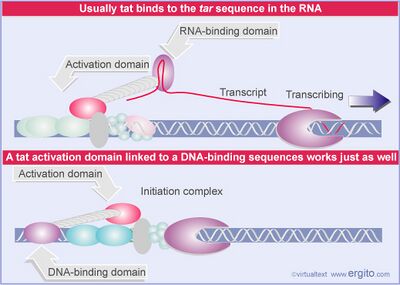
- Usually there is a latency period in reverse transcription during which RNA polymerase disengages prematurely from the complex. This results in the production of shorter and much less productive transcripts.
- Tat protein interacts with the Tar region at the 5' end of the viral RNA and is recruited to the promoter, where it keeps the transcription complex intact and allows for continuous transcripts to be produced.
Interactions with Viral RNA
- Tar RNA folds into a bulged beta-hairpin loop. The hairpin loop is located in the major groove of the RNA.
- Hydrophobic interactions in the widened major groove are important in the interaction of the Tat protein with the RNA.
- The hairpin turns between arginine73 and arginine77, and is facilitated by glycine74 and glycine76. Thus, substitutes in those regions are not permitted.
- Cationic amino acids in Tat bind to phosphate residues in the Tar region and anchor the beta sheet of the protein to the major groove. This is contrary to the alpha helix interactions that were thought to occur between Tat and Tar.
Application/Treatment
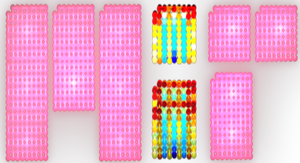
- CGP64222 is a peptidic compound that is able to bind to Tar RNA and prevent Tat from binding to it, thus providing selecting inhibition against Tat.
- The image below shows the synthetic peptide-RNA complex. There is high occupancy of the RNA groove, which contributes to the stability of the complex. Negative charges associated with backbone phosphates are highlighted in yellow.
- A DNA Origami device with more affinity for this or another competitive synthetic peptide than for the Tat protein itself could serve as a shuttle or delivery mechanism to the nucleus, or at least the T cells. The problem here is that the synthetic peptide must be similar to Tat in order to bind to the viral RNA, and so creating a device that has much greater affinity for the synthetic peptide will be difficult.
Extracellular Role in Pathogenesis
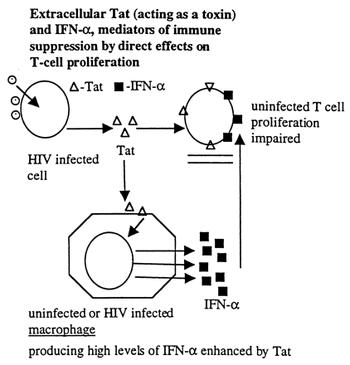
- After reverse transcription, Tat is released from infected CD4+ T cells. Its release is dependent on temperature and on serum concentration (19). However, the release is not inhibited by drugs that inhibit classic secretion pathways or endo and exocytosis, such as brefeldin A and methylamine.
- Tat activates interferon-alpha production from macrophages. IFN-alpha is present in high levels in patients infected with HIV, and is immune suppressive.
- Tat also inhibits T-cell proliferation, and causes apoptosis of some surrounding cells.
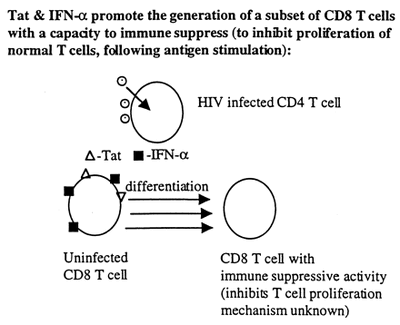
- Together, Tat and IFN-alpha promote the production of a type of CD8+ T lymphocytes that have immunosuppressive properties.
- Full recovery of normal T cell proliferation is achieved when peripheral blood mononuclear cells are cultured in the presense of antibodies for both Tat and IFN-alpha. Only partial recovery is seen when cultured with each antibody alone (20).
- Tat and IFN-alpha also inhibit the production of C-C chemokines that, when abundant, have an effective inflammatory response to HIV-1.
Potential Origami
- A DNA origami device that can enhances affinity between Tat and its antibody, or IFN-alpha and its antibody, would create more reliable and stable targets for immune cells because of massive aggregation, and may increase the efficiency of the immune response.
- The device would probably be small, and could possibly be administered with a Tat vaccine.