BioSysBio:abstracts/2007/Philip Jones
EMBL EBI Services Supporting the Proteomics Biocomputing Community
Author(s): Philip Jones, Richard Cote, Lennart Martens, Henning Hermjakob
Affiliations: EMBL European Bioinformatics Institute, Wellcome Trust Genome Campus Hinxton, Cambridge, UK, CB10 1SD
Contact:email: pjones@ebi.ac.uk
Keywords: 'proteomics' 'ontology lookup' 'mass spectrometry' 'database' 'HUPO PSI'
Background/Introduction
The European Bioinformatics Institute EBI is a centre for research and services in bioinformatics, operating as a non-profit academic organisation that forms part of the European Molecular Biology Laboratory EMBL. Part of the work of the EBI includes services for the proteomics community, through the work of the Proteomics Services Team. The work of this team includes support for specific collaborative research efforts such as Transfog and Enfin together with the development, curation and maintenance of public data repositories for proteomics, including the Intact database of molecular interactions. This presentation describes two resources that are made available from the Proteomics Services team, the PRIDE PRoteomics IDEntifications database and OLS, the Ontology Lookup Service.
Results
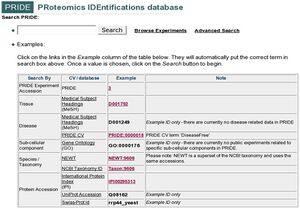
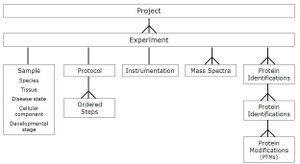
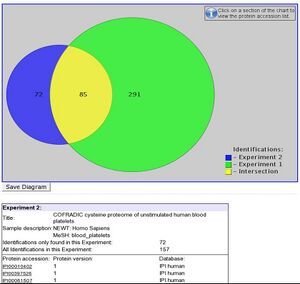
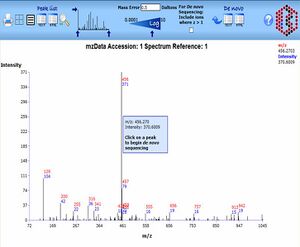
The PRIDE PRoteomics IDEntifications database PRIDE is a centralized, standards compliant, public data repository for proteomics data. It has been developed to provide the proteomics community with a public repository for protein and peptide identifications together with the evidence supporting these identifications. PRIDE currently implements the HUPO PSI mzData schema (version 1.05) that captures mass spectra peak lists, analysis sample details and spectrometer settings. PRIDE extends this schema with a data model designed to capture the output of search engines, including details of protein and peptide identifications and protein modifications that have been identified from MS. The PRIDE development team are committed to implementing relevant standards from the HUPO Proteomics Standards Initiative as they are ratified. It is expected that the PSI analysisXML standard that is being designed to capture search engine output will replace much of the existing PRIDE-specific schema within the next twelve months.
The Ontology Lookup Service OLS is a spin-out project and dependency of PRIDE that provides a single point of query for over 40 controlled vocabularies and ontologies that are updated daily from their sources. OLS provides interactive and programmatic interfaces for queries on term names, synonyms, relationships, annotations and database cross-references.
Images/Tables
The images below are displayed as 'thumbnails'. Please click on an image to view it full size.
PRIDE Screen Shots

OLS Screen Shots



Materials/Methods
Both PRIDE and OLS are pure Java applications that are made available under the Apache 2 Open Source Software Licence and both can be obtained by clicking on the green 'Download PRIDE' icon on the PRIDE Sourceforge project page. Both applications include web applications for query, reporting and in the case of PRIDE, data submission. These web applications are based upon the Struts framework.
Conclusion
Both PRIDE and OLS will be presented, including the dependency of PRIDE upon OLS and the relationship between PRIDE and the HUPO Proteomics Standards Initiative. Time and space permitting, future developments of the PRIDE user interface (especially developments of the query and reporting interfaces) will be described.
References
Philip Jones, Richard Cote, Lennart Martens, Antony Quinn, Chris Taylor, William Derache, Henning Hermjakob, Rolf Apweiler. (2006) PRIDE: a public repository of protein and peptide identifications for the proteomics community Nucleic Acids Research Vol 1 Issue 34 (Database issue) D659-D663
View Full Text (Open Access Publication)
Lennart Martens, Henning Hermjakob, Philip Jones, Chris Taylor, Kris Gevaert, Joel Vandekerckhove, Rolf Apweiler. (2005) PRIDE: The PRoteomics IDEntifications database Proteomics Vol 5 Issue 13 Pages 3537-3545.
View ePublication abstract
Cote RG, Jones P, Apweiler R, Hermjakob H.The ontology lookup service, a lightweight cross-platform tool for controlled vocabulary queries.
BMC Bioinformatics. 2006 Feb 28;7(1):97
Acknowledgements
PRIDE is supported through BBSRC iSPIDER and HUPO Plasma Proteome Project funding as well as an EU Marie Curie fellowship.
For further help on editing, please see here