BioSysBio:abstracts/2007/Martin Hemberg
A study of the properties of stochastic genetic oscillators
Author(s): Martin Hemberg and Mauricio Barahona
Affiliations: Imperial College London
Contact:email: martin dot hemberg at imperial dot ac dot uk
Keywords: 'oscillator' 'simulation' 'Perfect Sampling' 'add_keyword_4'
Introduction
Oscillations are of great importance in biological systems as they are found in a number of key proceses, such as the circadian rhythm. This type of periodic behavior at the level of genes has attracted significant attention from both experimentalists and theoreticians for a long time [1]. In recent years there has been a surge of interest in stochastic models of gene regulatory networks. This type of description allows one to investigate how the systems perform in a noisy environment. Investigations of stochastic chemical reactions usually take their starting point in the Chemical Master Equation (CME). The CME is a difference-differential equation describing the probability distribution of the system.
We present a comparative study of three genetic oscillators: the repressilator [2] the Atkinson oscillator [3] and a simplified models of a transcriptional oscillator by Guantes and Poyatos [4]. The first two examples are synthetic oscillators that have been implemented in E. coli. The last one is an example of an enzymatic oscillator and should be considered as a simplified model of natural systems. A crucial advantage of the last models is that it is two-dimensional making it easier to visualize the full solution.
Predicting the behavior of a stochastic oscillator based solely on the underlying equations is very difficult. Furthermore, solving the CME is often very hard and computationally intensive. An important aspect of the stochastic description is that the deterministic description can be obtained as a limiting case. Thus it is important to be able to relate the properties of the stochastic oscillator to properties of the corresponding deterministic formulation. These are often easier to compute and are easier to understand and interpret than the stochastic quantities. Moreover, we investigate the use of methods based on a stochastic differential equation (SDE) description of the stochastic system and show how they compare with the CME and the deterministic results.
Results
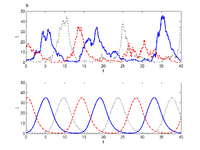
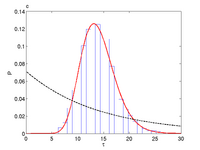
We investigate several properties of stochastic genetic oscillators by the use of a perfect sampling algorithm (see Methods below). These include, the distribution of period lengths, a characterization of the noise level affecting the period, the autocorrelation, the effect of the system size, the shape of the oscillations and the sensitivity to parametric perturbations. Furthermore, we show how high-dimensional noisy oscillators can be robustly embedded in a lower dimensional space. We also investigate how the noise level can be connected to the divergence of the deterministic system. These properties are important as they allow us to characterize genetic oscillators in terms of reliability, precision and tunability.
Methods
Solving the CME analytically is very difficult and the most popular method today is to use a numerical method; Gillespie's Stochastic Simulation Algorithm (SSA). The SSA is a kinetic Monte Carlo algorithm which provides an exact realization of the underlying CME. In principle, the stationary distribution of the CME can be obtained by running the SSA for an infinite amount of time and in practice one usually runs the for 'a very long time'. We have developed the Dominated Coupling From The Past Stochastic Simulation Algorithm (DCFTP-SSA) which guarantees that the simulation of the CME will not be terminated before the stationary distribution has been reached (paper in submission, please email MH for preprint). By employing the DCFTP-SSA, we can start the simulation of the oscillators from the stationary distribution. This ensures that the data will not have any biases due to sampling from the transient.
References
- Goldbeter A. Computational approaches to cellular rhythms. Nature. 2002 Nov 14;420(6912):238-45. DOI:10.1038/nature01259 |
- Elowitz MB and Leibler S. A synthetic oscillatory network of transcriptional regulators. Nature. 2000 Jan 20;403(6767):335-8. DOI:10.1038/35002125 |
- Atkinson MR, Savageau MA, Myers JT, and Ninfa AJ. Development of genetic circuitry exhibiting toggle switch or oscillatory behavior in Escherichia coli. Cell. 2003 May 30;113(5):597-607. DOI:10.1016/s0092-8674(03)00346-5 |
- Guantes R and Poyatos JF. Dynamical principles of two-component genetic oscillators. PLoS Comput Biol. 2006 Mar;2(3):e30. DOI:10.1371/journal.pcbi.0020030 |