BioSysBio:abstracts/2007/Fernando Ortega
- Add or delete the sections that you require.
Kinetic constraints on the sensitivity of large metabolic responses
Author(s): Fernando Ortega1 and Luis Acerenza2
Affiliations: 1. School of Biosciences. University of Birmingham. UK 2. Laboratorio de Biología de Sistemas, Facultad de Ciencias, Universidad de la República. Uruguay
Contact:email: f.ortega@bham.ac.uk
Keywords: 'metabolic control analysis' 'MCA' 'modular analysis' 'mean control coefficients'
Background/Introduction
The quantitative study of metabolic responses in intact cells is essential for predicting the phenotypic consequences of genetic manipulations. Metabolic responses have been described within the framework of metabolic control analysis (MCA) [1]. One of the central goals of MCA is to determine how the responses of system variables, quantified by control coefficients, depend on the kinetic properties of the component reactions or groups of reactions (modules).
Attempts to extend infinitesimal control analysis to large changes in the variables have been reviewed in previous publications [2,3].
In the present contribution, we analyse how some kinetic features of the modules constraint the sensitivity of large metabolic responses. In particular, we explore the different kinds of flux control profile that can be achieved using modules that follow two different types of rate equations: Michaelis-Menten and Hill.
Results
In this study, we consider that the metabolic network can be split into a supply module ([math]\displaystyle{ v_{1} }[/math]), that produces the intermediate S, and a demand module ([math]\displaystyle{ v_{{2}} }[/math]), that consumes [math]\displaystyle{ S }[/math] (scheme) [4]. This top-down or modular MCA approach has been widely used to analyse the sensitivity properties of metabolic networks [5]. Firstly, the rates and vs. S were chosen to obey a reversible Michaelis-Menten and Hill equation respectively (see Figure for details). The maximal activities of module 1 and 2 are modified by the factor [math]\displaystyle{ r_{{1}} }[/math] and [math]\displaystyle{ r_{{2}} }[/math], respectively (one at a time). The system starts at a reference steady-state o, where [math]\displaystyle{ r_{{1}}=r_{{2}}=1 }[/math]. After the parameter is changed ([math]\displaystyle{ r_{{1}} }[/math] or [math]\displaystyle{ r_{{2}} }[/math]) the variables of the system ([math]\displaystyle{ S }[/math] and [math]\displaystyle{ J }[/math]) freely adjust to the final steady state.
In the Fig. we plot the mean flux and metabolite control coefficient for both modules when subject to large changes in the enzyme concentrations or activities. The infinitesimal control coefficients ([math]\displaystyle{ r_{{1}}=r_{{2}}=r=1 }[/math]) are [math]\displaystyle{ C_{{v1}}^{{J}}=0.58 }[/math], [math]\displaystyle{ C_{{v2}}^{{J}}=0.42 }[/math], and [math]\displaystyle{ C_{{v1}}^{{J}}=|C_{{v2}}^{{S}}|=1.05 }[/math]. But for large changes, which module shows the greatest control depends on the value of [math]\displaystyle{ r }[/math]. In the interval (0.56-1.8) the greatest flux control is in module 1, meanwhile outside this interval the flux is mainly controlled by the second module. Regarding the control of [math]\displaystyle{ S }[/math], for [math]\displaystyle{ r>1 }[/math]a change in module 1 produces a greater effect on [math]\displaystyle{ S }[/math] than module 2, while for [math]\displaystyle{ r<1 }[/math] module 2 has the greatest effect.
On the other hand, if both modules fulfil M-M rate equation it can be demonstrated that for all values of [math]\displaystyle{ r }[/math] one of the modules has the greatest flux control (results not shown).
Figures
![]()
Scheme: Metabolic system constituted by a supply module ([math]\displaystyle{ v_{1} }[/math])
and a demand module ([math]\displaystyle{ v_{{2}} }[/math]) kinked by one intermediate.
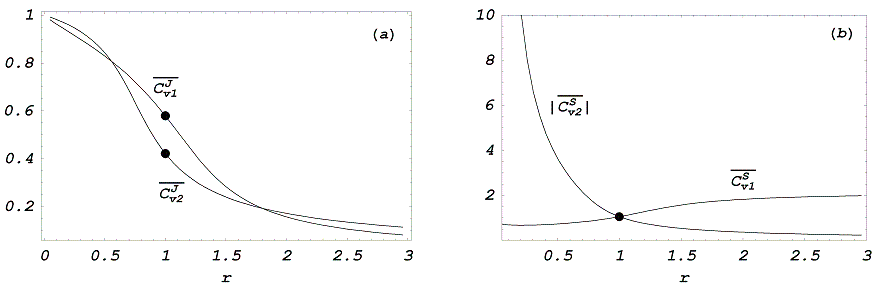
Figure: Mean control coefficients vs module activity, [math]\displaystyle{ r }[/math]. a) flux
mean control coefficients and b) intermediate mean control coefficients.
M-M equation parameters : [math]\displaystyle{ X_{{0}}=10,\, K_{S}=1,\, K_{P}=0.15,\,1/K_{eq}=0,\, and\, V_{m1}=1 }[/math].
Hill equation parameters: [math]\displaystyle{ K_{S}=0.15,\, h=4,\, and\, V_{m2}=0.63 }[/math].
Materials/Methods
The mean control coefficient quantifies the sensitivity of a steady-state metabolic variable ([math]\displaystyle{ W }[/math]) to changes in a parameter ([math]\displaystyle{ p }[/math]). It is defined as [math]\displaystyle{ \overline{C_{p}^{W}}=((W_{{x}}-W_{{o}})/W_{{o}})/((p_{{x}}-p_{o})/p_{o}) }[/math]. It represents the relative change in [math]\displaystyle{ W }[/math] divided by the relative change in p when the system goes from one initial ([math]\displaystyle{ o }[/math]) to one final steady state ([math]\displaystyle{ x }[/math]) [2].
Conclusion and Discussion
We have analysed two different models to see if certain patterns of control for large changes could be, in principle, obtained using modules whose rates are governed by usual rate equations. We have shown that, if the rates of both modules obey hyperbolic kinetics, the flux control distribution is constrained, one module having the greatest control over all enzyme activity range. On the contrary, if the rate of the demand module follows a Hill kinetics, which module has the greatest control on the flux depends on the enzymatic activity change.
References
1. Heinrich R. & Schuster S (1996) The regulation of cellular systems, 1st edn. Chapman & Hall, NY.
2. Ortega F & Acerenza L (2002) Elasticity analysis and design for large metabolic responses produced by changes in enzyme activities. Biochem. J. 367,41-48.
3. Acerenza, L & Ortega, F (2006) Metabolic control analysis for large changes: extension to variable elasticity coefficients. IEE Proc Syst Biol 153, 323-326.
4. Hofmeyr, JS & Cornish-Bowden, A (2000) Regulating the cellular economy of supply and demand. FEBS Lett 476, 47-51.
5. Quant PA (1993) Experimental application of top-down control analysis to metabolic systems. Trends Biochem Sci 18, 26-30.
For further help on editing, please see [ http://en.wikipedia.org/wiki/Help:Editing here]