BioBuilding: Synthetic Biology for Students: Lab 4
|
Eau That Smell Lab |
Lab 4: What a Colorful World
Acknowledgments: This lab was developed with materials from the University of Cambridge 2009 iGEM team, as well as guidance and technical insights from Drew Endy and his BIOE.44 class at Stanford UniversityObjectivesBy the conclusion of this laboratory investigation, the student will be able to:
IntroductionOne potential use of engineered bacteria is as indicator of toxic substances. Bacterial sensing systems have been designed for arsenic and lead. Bacteria are cheap and easy to produce and store. This reduces the need for expensive and technologically complex chemical tests. The bacteria are also much more sensitive to the toxin levels. However, there is one potential drawback. The bacteria respond to the toxin metabolically. This means we may be able to detect a change in pH or other indicator of metabolism. This requires further equipment such a pH indicator. Sensors have been linked by synthetic biologists to other forms of output such as the green fluorescent protein. However, this also requires further equipment such as a fluorescent light. This reduces the practicality in impoverished areas of the world, the very areas most at risk for arsenic or lead contamination. The 2009 Cambridge iGEM team took up the challenge to design an indicator that could be used without additional technology. They designed color generator devices that could be linked to sensors. E. coli are naturally colorless, but other bacteria make pigments and so do appear colored. The iGEM team designed “e chromi,” engineered E. coli capable of producing colors through the synthesis of pigments. One pigment they used is Violacein, a pigment produced by a handful of genes originally found in Chromobacterium violacein. These genes were re-engineered and combined to produce purple and green in E. coli. The violacein operon consists of five genes which metabolize L-tyrosine. Expression of all five genes will produce a purple pigment. However, removal of the third gene in the sequence will cause the cell to metabolize the L-tyrosine into a green pigment. These pigments are easily visible to the naked eye. This device could be linked to a biosensor for a toxin and the bacteria will turn color in response to the toxin concentration. It's reasonable to wonder: Why didn't the team just use the Chromobacterium? Synthetic biologists like to use E. coli because it is well understood and easy and safe (if proper strains are used) to work with. But it's important to realize that this was a choice! Synthetic biologists refer to the host cell as the chassis, and just as you'd carefully design a genetic program to encode, you'd also need to carefully choose the chassis that will run it. For an engineered genetic system to function in a chassis, the chassis must supply the cell with energy, materials for protein synthesis and materials those proteins will use when they function. The chassis will take care of all the material needs to meet the engineer’s specifications. The better the chassis is understood, and the better it can provide materials for the engineered system, the better the results. By primarily using one chassis, synthetic biologists are managing complexity. A standard chassis allows engineers from many labs across the world to compare results. Note how we also manage complexity in our everyday life. When we buy bananas or bell peppers, we simply call them bananas or bell peppers. In actuality, many varieties get mixed together in the store. But is it really important that we are aware of this when we shop? As long as the taste is similar, does it matter what variety of peppers you use? Cars, however, are a different story. A car is a highly engineered system of interconnected parts. While many of these parts are similar, they must be tailored to the size and function of the car. So, while the chassis of a truck, a GTO muscle car and a Toyota hybrid are different, so are many of the internal parts that make up the engine and the drive train. We might be able to move a radio from a truck chassis to a sports car chassis, but not much else. The car manufacturers are comfortable with this complexity and it has little effect on the user of the car. What about your computer? You can think of your computer and its operating system as a chassis, making Macs and PCs different chassis (though in computer lingo they are known as platforms). There was a time in the past when word processing files written on one platform could not be viewed or edited on the other. But interoperability was clearly needed and so the computer companies have agreed on certain standards. Through re-engineering of the programs and the chassis/platforms, users no longer get lost in the complexity. Thankfully, files written on one platform can be viewed and edited on the other. Synthetic biologist George Church is working to further remove the complexity from engineered systems by creating what are known as minimal cells. The idea is to design a cell that contains just the minimum genome to maintain its existence. These cells will only be able to survive on special media and all of their metabolic functions will be well characterized. Another example of research into this idea was published by Craig Venter in May of 2010. His lab replaced the genome of a bacterial cell with a fully synthesized genome and were able to produce bacteria that expressed the synthetic genome. As appealling as these chassis are for synthetic biology, the work has a way to go before they can be in general use. So, until minimal cells or synthetic cells are a viable option, researchers continue to use E coli and other domesticated cells as chassis for experiments. Mostly, the strains of E. coli that are used in research labs are one of two kinds. One strain is known as K-12 and the other B. Both strains are known to be safe and have been effectively used for genetic experiments for almost 100 years. The differences between these strains seem to be minor. Most are related to metabolism and none would seem likely to affect the color generator system. You can read about the interesting history of these strains here. So now imagine that a group of engineers is manufacturing an arsenic sensor in E. coli. This group would like the intensity of purple color to vary as a function of arsenic level. Now imagine that a second group of engineers are also doing this but they use a different strain of E. coli. How sure can we be that the pigment will be expressed the same in a different chassis? Thinking back to our analogy with car chassis: would an engineer put a V-8 engine from a Lexus into a Mercedes chassis? Would the engine behave the same? Would the car? In this lab you will transform bacteria from two different strains of E. coli, in other words, two different chassis. Strain 4-1 is a K-12 strain, while strain 4-2 is a B-type strain. Into each strain you will insert plasmids containing violacein-pigment devices. One plasmid, pPRL, has the purple version of this device while the other plasmid, pGRN, has the dark green version. Otherwise, the plasmids are the same. Can we expect the devices to behave the same in each strain or will the chassis have an effect on the intensity of color produced? ProcedurePart 1: Preparing Strain 4-1 and 4-2 for transformation
Part 2: Transforming Strains 4-1 and 4-2 with pPRL and pGRNThe cells you've prepared will be enough to complete a total of 6 transformations. You will transform the purple-color generator into each strain, and also the green-color generator into each strain. You will also use the last bit of competent cells as negative controls for the transformation.
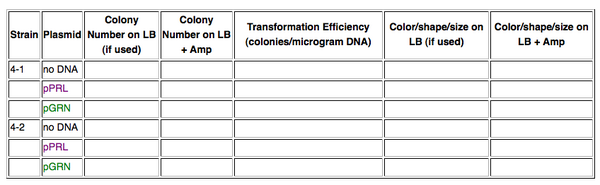
Next dayIn your lab notebook, you will need to construct a data table as shown below. These may be provided. Also be sure to share your data with the BioBuilder community here.
CalculationsHere is a sample calculation for transformation efficiency
Calculation:
Lab ReportI. Introduction
II. Methods
III. Results
IV. Discussion
Navigation
|