Bertilsson
Welcome to the Bertilsson Lab at Uppsala University
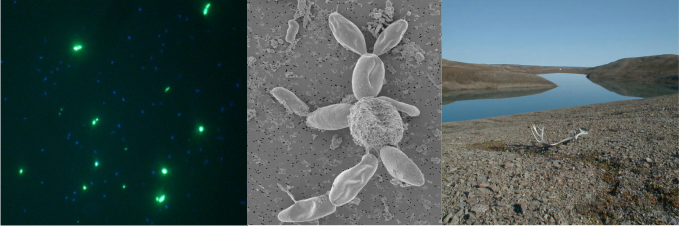
Fieldwork in Mosquito-land |
News!
|
Research
The Microbial Ecology group is part of the Limnology program at the Department of Ecology & Evolution, EBC, Uppsala University. The research carried out in the group is focused on understanding the roles of microorganisms in aquatic food-webs and biogeochemical cycles. We address the role of microbial diversity in controlling processes at the community and ecosystem-level using an array of culture-independent molecular methods. We also design and perform laboratory and field experiments to address these questions and our continued research aims at revealing relationships between function and identity at the level of individuals, populations and communities.
Selected publications
Bertilsson, S., Eiler, A., Nordqvist, A., and N.O.G. Jørgensen. 2007. Links between bacterial production, amino acid utilization and community composition in productive lakes. ISME Journal 1:532-544
Eiler, A., and S. Bertilsson. 2007. Flavobacteria blooms in four productive lakes: Linking population dynamics of freshwater bacterioplankton to resource availability. Applied and Environmental Microbiology 73:3511-3518
Polymenakou, P., Tselepides, A., Stephanou., E., and S. Bertilsson. 2007. Organic Matter Preservation and microbial community accumulations in deep-hypersaline anoxic basins. Geomicrobiology Journal 24:19-29
Eiler, A., Johansson, M., and S. Bertilsson. 2006. Environmental influences on Vibrio populations in northern temperate and boreal coastal waters (Baltic and Skagerrak Sea). Applied and Environmental Microbiology 72:6004-6011
Jarvius, J., Melin, J., Göransson, J., Stenberg, J., Fredriksson, S., Gonzalez-Rey, C., Bertilsson, S., and M. Nilsson. 2006. Digital quantification using amplified single molecule detection. Nature Methods 3:725-727
Recent updates to the lab wiki
List of abbreviations:
- N
- This edit created a new page (also see list of new pages)
- m
- This is a minor edit
- b
- This edit was performed by a bot
- (±123)
- The page size changed by this number of bytes
21 May 2026
|
|
21:46 | Hu:Publications 3 changes history +405 [Hugangqing (3×)] | |||
|
|
21:46 (cur | prev) +1 Hugangqing talk contribs | ||||
|
|
21:46 (cur | prev) +56 Hugangqing talk contribs | ||||
|
|
21:42 (cur | prev) +348 Hugangqing talk contribs | ||||
|
|
21:40 | Hu 2 changes history +195 [Hugangqing (2×)] | |||
|
|
21:40 (cur | prev) +105 Hugangqing talk contribs | ||||
|
|
03:26 (cur | prev) +90 Hugangqing talk contribs | ||||
19 May 2026
|
|
02:49 | Hu:Members 2 changes history +178 [Hugangqing (2×)] | |||
|
|
02:49 (cur | prev) −563 Hugangqing talk contribs | ||||
|
|
02:47 (cur | prev) +741 Hugangqing talk contribs | ||||
| 02:45 | Upload log Hugangqing talk contribs uploaded File:Naved.png (Naved) | ||||