Berk2006-Samantha's Notebook
12-12-06
- Assayed all tRNA-like linker keys
TGI 90 1122 109 1022 13189 J23201 738 J23202 107 J23203 374 J23204 82 J23205 231 J23206 439 J23207 153 J23208 214 J23262 206 J23270 712
12-08-06
- miniprepped J23202 and transformed in into J01122 (in incubator now)
- sequenced J23202, J23204, J23205, J23206, J23207, and J23208 (check J23205 and J23207 carefully)
- Assayed all keys except for J23202
TGI 93 1122 89 1022 9765 J23201 510 J23203 386 J23204 281 J23205 478 J23206 342 J23207 323 J23208 313
12-06-06
- sequencing results for the old keys came out nicely - J23201 & J23203 were perfect, but J23205 and J23207 were switched. Not too sure if the minipreps are actually reversed or if I just messed up while sequencing...any way to do figure out besides sequencing again?
- Assuming all the even keys are most likely right as well (will sequence them later), I transformed all the keys into J01122 (lock3) except for J23202 because the miniprep has disappeared for some reason
- Grew up a culture of J23202 to miniprep - will send all the even keys out for sequencing once I get J23202.
12-01-06
- a little scared that the backbone digest of J23061 may have been wierd because Chris has been getting wierd results with it
- miniprepped one clone of each new key
- set the odd keys out for sequencing - J23201, J23203, J23205, and J23207.
11-29-06
- ran the colony PCRs for 4 clones of each new key - they all looked pretty good with bands around 200bp
- grew up two cultures of each key to miniprep and sequence
11-27-06
- Colony PCRed the new keys again (all 8 this time)
- Ran gels on the PCRs from Monday, only 2 bands that were way too big - hmm...redo
- Looking for bands of around 200 bp (J23205 is 216)
- Updated Registry
yo sam, can you update these new keys in the registry? happy thanksgiving. thanks -bryan
11-20-06
- Colony PCRed the 8 new keys (J23201-J23208) except for J23206 and J23207, which did not grow any colonies.
- Religated and transformed J23206 and J23207
- transformed key J23086 into new lock J23093.
11-17-06
- Tecan Results of new lock:
J01122 J23090 J23077 J23090 x J23086 J01022
367 559 392 2995 39624
99 166 105 812 10873
- Made another digest of J23061 (discovered that it only has about an 8 base pair sequence between BamHI/HindII sites - never had GFP inserted into it.
- Subcloned the 8 new keys with different spacings (J23201-J23208) - digested and cleaned up PCR products and then ligated it with J23061 and transformed them into TGIs.
11-15-06
- Did a larger-than-usual-scale digest on the J23061 (BamHI/HindIII, in NEB2)
- Picked a J01022 colony to make a -80 stock, and also freshly restreaked it
- Make a small-scale (1mL culture) KCM Comp cell prep of pSB3C6-J23090 and
transform with pJ23006-J23086, plate on Carb/Cam (J23090 is a new lock variant which has the extended 5' end and the 'corrected' rbs spacing.
- set up the extension reactions and cleaned up the PCR products
10-25-06
- Miniprepped pJ23006-J23066
- Then transformed J23066 into J23077
- replated KOs at 43C after dilution
- Tecan of several Riboregulator transformations gave the following results:
- J23077 only 31
- J01022 only 1817
- TGI only 34
- J23066xJ01122 149
- J23077xJ01122 43
- J23059xJ01122 53
- J23060xJ01122 146
- J23029xJ01122 30
- J01122 only 19
10-16-06
TECAN RESULTS
CULTURE 120 gain 130 gain 140 gain
J23077 x J23086 (mated) 503 861 1630
J23077 x J23086 (transformed) 475 794 1427
J23077 x J0103 74 117 202
J01022 x J23086 4745 8308 14699
J01022 x J01003 5913 10514 18598
TGI 61 94 168
J01022 1774 3170 5580
J23077 63 93 175
Why won't the wiki show more than one space between the numbers even though thats how I'm formatting them? And why is the fluorescence of J01022 by itself so small? The mated and transformed values of J23077xJ23086 are close and look pretty good though.
MATING RESULTS
Conjugation experiments with J23077 or J01022 as the recipient
Number of Titer Colonies for EC100D::pir+ = 4
| Doner | Recipient | # of Colonies on AmpCamp | Dilution Factor | Comments |
|---|---|---|---|---|
| J23086 x BW20767 | J23077 | 1096 | 100 | looks good |
| J23066 x BW20767 | J23077 | 3 | 0 | looks good |
| J01003 x BW20767 | J23077 | 1408 | 100 | looks good |
| TGI | J23077 | 0 | 0 | looks good |
| J23086 x BW20767 | J01022 | 1600 | 100 | looks good |
| J23066 x BW20767 | J01022 | 4 | 0 | looks good |
| J01003 x BW20767 | J01022 | 1472 | 100 | looks good |
| TG1 | J01022 | 0 | 0 | looks good |
Conjugation experiments with EC100D::pir+ as the recipient
Number of Titer Colonies for EC100D::pir+ = 4
| Culture | # of Colonies on TriK | # of Colonies on TriA | Dilution Factor | Comments |
|---|---|---|---|---|
| WT pOX38 | 3968 | 0 | 100 | looks good |
| pOX 38+DH10B/J01064 | 2880 | 1760 | 100 | results from an old experiment |
| J23015 | 0 | 0 | 0 | looks good |
| J23015xJ01064 | 10 | 1264 | 100 | looks good |
| RP1 | 112 | 2016 | 100 | weird that there are more on TriA than TriK (ignore TriA maybe) |
| Rlambda | 1 | 0 | 0 | looks good |
| Rlambda+JO1003 | 0 | 1664 | 0 | looks good |
Number of Titer Colonies: 5 (done for J23066xBW20767 - assumed to be the same for all starter cultures)
10-13-06
HUGE MATING FEST - all were plated undiluted and also diluted 100x
AMPCAM PLATES:
- J23086xBW mate with J23077
- J23066xBW mate with J23077
- J01003xBW mate with J23077
- TGI mate with J23077
- J23086xBW mate with J01022
- J23066xBW mate with J01022
- J01003xBW mate with J01022
- TGI mate with J01022
TRIK & TRIA PLATES (All mated with EC100D):
- pox38
- J23015
- J23015xJ01064
- RP1
- Rlambda
- RlambdaxJ01003
NOTE: pOX38xJ01064 did not grow so we are using old results which were 2880 colonies on TriK, and 1760 colonies on TriA with a 100x dilution, with a titer that gave 3 colonies. Undiluted - there were lawns. (this comes from the conjugation notebook)
10-11-06
- mated J23086xBW20767 and pSB3C6 - J23077
- transofrmed J23066 into BW
- transformed J01003 into BW
- transformed J01064 into J23015
- cotransformed J23086 and J23077 into TG1s (for fluor. testing)
- restreaked pox38x1064, Rlambdax1003 and pox38
- streaked from -80, 1022, J01003, RP1, Rlambda, J23015, and TG1s
10-09-06
- subclone pSB1AKG0-J23075 - plated on AmpKanGen
- mapped J23086 (4 clones) and J23088 (2 clones)
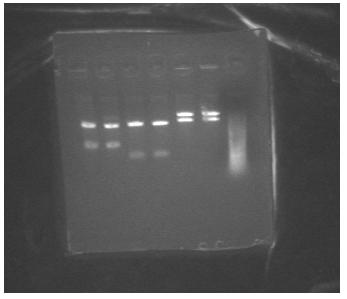
Lanes 1-4 are j23086, lanes 5-6 are J23088 and lane 7 is the ladder. The ladder came out pretty badly and John thinks it's because someone got enzyme in the ladder and it's been slowly chewing away at it because his gels have been getting wierd ladders too. But clones 1&2 of J23086 look good and so do the J23088s.
- transformed pJ23006-J23086-1 into BW20767 to mate wednesday.
10-06-06
- Mated J23016/J23074 with Ec100D, plate on TriK to test for complentation of trbC
- Grew up J23087 (the trbC/oriTF double knockout, it may be in the 42)
- miniprepped pSB3C6-J23074
- subcloned J23086
- subcloned J23088
10-04-06
- Retransformed to TGI to stock pSB1A2-J23038#4
- Transformed pSB1AK3-J23074 into J23016 trbC KO to do assay
- Scraped, diluted, and replated at 42 degrees the TrbC KO of J23055
- analyzed pSB1AG0-J23038 sequence data - clone #2 is correct, clone #1 is not
- miniprepped 4 clones of pSB1A2-J23086, mapped them with E/P
- Subclone pSB3C6-J23074
- mapped the remaining three clones of pSB1A2-J23075 - they're all wrong
- sequence J23074 and J23075
JCAnderson 22:38, 2 October 2006 (EDT)
I ran Sam's gel of J23074 and J23075, but overran it. They all looked much better than when we transformed into TG1--all gave normal bands, no extra bands, all samples were mostly similar. Note that the correct sequences of things are:
- pSB1AK3-J23074
- pSB1A2-J23075
(They aren't both pSB1AK3).
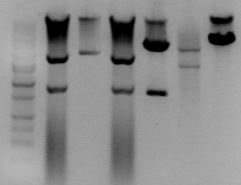 I remapped clone 1 of each and set up the EcoRI/SpeI digests to transfer things to pSB3C6 (above) The lanes go:
I remapped clone 1 of each and set up the EcoRI/SpeI digests to transfer things to pSB3C6 (above) The lanes go:
- pJ23019 (ES)
- pSB1AK3-J23074 (ES)
- pJ23019 (ES)
- pSB1A2-J23075 (ES)
- pSB1AK3-J23074 (AvrII/AlwNI)
- pSB1A2-J23075 (AvrII/SpeI)
The J23074 clone 1 is correct, and I cut the slices to do ligations. J23075 was wrong, we need to remap the other 3 clones.
10-02-06
- Miniprepped J23074 and J23075, digested in AvrII and SpeI to map
- Miniprepped pSB1AG0-J23038s, and sent them for sequencing
- Results from conjugation assay:
J23057xJ01003 on TriA ---- has >1000 colonies, is problematic because supposed to have a lot less efficiency since traG is knocked out
J23057xJ01003 on TriK ---- no colonies, which is good
J23055xJ01064 on TriA ---- only 4 red colonies on plate after a 1 mL dilution - good results
J23055xJ01064 on TriK ---- no colonies, good
- plate of pJ23006-J23086 key3.oriTr looks good. need to grow up cultures and miniprep eventually
9-30-06
- Genomic mini of j23016
- PCR j23016 with JL16 and JL17 for D/W
- mate J23055/J01064, mate J23057x1003
- resub key3.oriTr - jen said pJ23006-J23006 but actually pJ23006-J23066 and J01003
- grow up two clones of pSB1AG0 - J23038 , 4 clones of pSB1A2-J23074 and pSB1A2-J23075 each (Maybe John will do the growing up tomorrow so that they will be fresh on monday )
9-29-06
- Mini, map, J23074 and J23075 - the gel came out pretty crazy, none look right
- move to pJ23018 (ES)(p15A version ofTraG) when ready) - digested it today
- (Transform J23016 x pSB1A2-J23075 (Show we can rescue TrbC function))
- Religate J23074 and J23075 and put them into DH10B instead
- Subclone J23038 with Gen marker in front)
- Sub the key3.oriTr with pJ23006 and J01003
- Grow to gen prep J23016 (trbC KO, CmR)
- Grow to D/W J23055 #1 (OriTf KO) /pKD46 (x trbC cass)
- Grow to test mate J23055/J01064 (OriTf) (Show we can move transfer function to new plasmid)
- Grow to test mate J23057/1003 redo (later x pAC-traG) (Same as above but in the R system)
9-22-06
- resub J23074 and J23075
- map pSB1AG0-J23078 (the analytical digest is in the incubator)
- comp J23055 #1 from starter x pSB1A2-J01064, and xpkd46 (30degrees)
9-20-06
- made competent cells of Rlam/J23057
- transformed pSB1A2-J01003 into Rlam/J23057
- Subcloning pSB1AG0-J23078 by digesting pSB1AKG0-b0015 (small) and pSB1A2-J23078 (big) in E/X to drop the Gen marker from b0015 into J23078. Got up to the digestion clean up step - ligation and transformation left to do.
- Ran a gel from the analytic digests of J23074, J23075, J23026, J23028. They came out negative and inconsistent
- gel on PCRs of J23055 (2 clones) - clone 1 showed a band, clone 2 did not
- replate to comp J23055 clone#1 on Kan (transofrm J01064, pKD46 later)
- Colony PCR J23075 and J23074
9-18-06
- grew up cultures for tecan for J23077xJ23066 assay including J23077 from -80
- ran a gel on PCRs for traGr knockout strain
9-15-06
- miniprepped and made a -80 of pSB1AT3
- miniprepped J23028 (running low)
- Grew and spotted MC1017C from CP20 and J23069 from CP20 on Amp and Cam
- Grew up 4 clones of pBCa1020-J23038
- Grew up 4 clones to screen of pSB3C6-J23077 and pSB1A2-J23078
- made LB stabs and LB-Carb plates
- Scraped J23015-pCP20 and grew a culture in LB only
JCAnderson 13:25, 15 September 2006 (EDT)
We got sequencing for J23072 clone 5 (iG139), it is correct. The cells were visibly pink, so we aren't pursuing this further. Putting the "correct" spacing between the rbs and start codon seems to bump the background in J23071 and J23072 too high. The tecan data for J23071 was:
TG1 50
1122 60
1122/1129 90
1022 6563
J23071 367
J23071/1129 272
J23071/J23066 844
850
1061
1247
Going to focus instead on J23077 (and J23078, the basic part), which are lock3d variants with the 5' extension.
9-13-06
- PCRed pJ23006-J23032 and pJ23019-J23049(aka pSB3C6-J23032) for later construction of J23077 and J23078
- ran a gel for a bunch of PCRs of J23074 and J2307
5s
9-11-06
- transformed J23075 clones 1 and 2 into J23016 for Jen (plated on Amp)
- assayed J23071x1129 and J23072x1129 - the J23071 looked promising, almost 10% fluorescence as the OnRFP, the J23072 was obviously wrong, because flurorescence was even lower than the lock by itself and the TG1 cells.
- J23072 could have a pink phenotype and too high of a background to use. Grew up another 4 colonies of J23072 to screen.
- Grew up tecan cultures J23071 only and J23071x1129 to check the background versus gain for j23071. (Chris will do the tecan tomorrow)
- digested pJ23076 with SpeI/PstI, the backbone to make J23073 with a PCR on j23072 just in case that does end up working in the future. Ran a gel and cleaned up the digested product. (In cardboard riboregulaters box)
- co-transformed pSB3C6-J23071 (best lock so far) and pJ23006-J23066-1 (best key so far) into TGI cells to see what happens before proceeding with J23071 and adding a 5 prime end.
9-10-06
- grew up tecan cultures
9-08-06
- culture of pSB3C6-J23072 clone #1 was slightly pink, so I proceeded only with clone #2 which was white and should be correct.
- Miniprepped, made a -80, and sequenced pSB3C6-J23072-2. Also threw out pSB3C6-J23071-1 from -80 stocks, because clone 2 was the right one.
- transformed J23072-2 and J23071-2 into J01129 (Chris will put plates in fridge tomorrow)
- all cultures of pSB3C6-I13521 appear white, even after pelleting, they still look white, so something must have gone wrong. Will remake later.
- made a bunch of LB Agar and LB Broth (Reza and Melissa will autoclave for me)
9-06-06
- grew up two white colonies of pSB3C6-J23072
- sequencing for J23071 clone#2 looks good and is in the -80
- pSB3C6-I13521 plate has all white colonies it seems unfortunately. Growing up four clones just in case they turn pink in culture.
For Friday:
- miniprep clones of pSB3C6-J23072, make -80s, and send them in for sequencing
- transform J23072 and J23071 into J01129
- Pick colonies Sunday and Assay on Monday
8-25-06
Riboregulators:
- cultures of new locks J23071 and j23072 look a little red. Miniprepped 2 white cultures of pSB3C6-J23071, made -80 stocks of them, and sent them for sequencing with ca1000. Pick colonies of J23072 again when we get back from the break, plates are in the fridge.
- ligated and transformed pSB3C6-I13521. (John will put in fridge tomorrow)
Conjugation:
- purified the PCR product of pSB1A2-J23038 and digested with BsaI/SpeI/dPN1, then purified again.
- digested pSB1AK3-J23026 (traG+TT) and pSB1AK3-J23028 (trbCf+TT) with BsaI/XbaI, ran gel and gel purified.
- ligated and transformed to get products pSB1AK3-J23074 ([pCon][RBS][traGr][TT]) and pSB1AK3-J23075 ([pCon][RBS][trbCf][TT]). (John will put in fridge tomorrow)
8-24-06
Riboregulators:
- grew up 4 white colonies each of new locks J23071 and J23072.
- Started to make pSB3C6-I13521 to replace pSB3C6-J01022, which just removes the upstream Ptet to do future measurements relative to a more-similar standard. Made a construction file and cleaned up digestions of I13521 and pAC997 (found in "Good" Plasmids box). Ligate and Transform tomorrow.
- sequenced pJ23006-J23070 #1 and pJ23006-J23062 #1 (the tRNA-based keys)with ca998.
Conjugation:
Finishing up the traG and trbC complements
- PCRed pSB1A2-J23038 with G00101 and ca641R to get the short pCon+RBS fragment. Tomorrow, digest this fragment with BsaI/SpeI/dPN1. Also, digest pSB1AK3-J23026 (traG+TT) and pSB1AK3-J23028 (trbCf+TT) with BsaI/XbaI. Purify, ligate and transform to make new pSB1AK3 complements. Make construction files. BsaI is not in the APE enzyme list it is GGTCTC.
--Samanthaliang 17:04, 24 August 2006 (EDT)