Beauchamp:MedNSLab
Beauchamp Lab wiki
|
|
This is the web page for information about downloadable MRI neuroanatomy teaching materials. The instructions below describe how to download MRI datasets and a freely available viewing program to look at them. REQUIRED Step 1: Download MRIcroN (free software for viewing MRI datasets)Download the correct installer package for your computer. Windows, Mac and Linux are supported; detailed installation instructions are provided for Windows and Mac but not Linux. For Windows: The following instructions were tested with Windows 7 running Internet Explorer 9. For different version of Windows or other web browsers, the steps may be slightly different.
After successfully starting the program, you should see the following window. To make the images easier to see, you can enlarge the window. Go to Step 2, below. 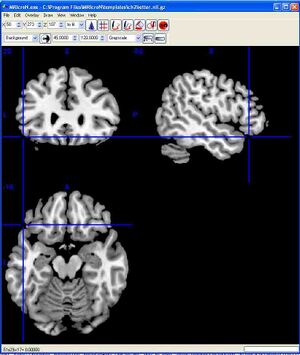 For Macintosh: The following instructions were tested with Mac OSX 10.8.2 running Safari. For different versions of OSX or other web browsers, the steps may be slightly different.
You should see the following window. To make the images easier to see, you can enlarge the window. To see only one plane enlarged, click View/Axial Only. 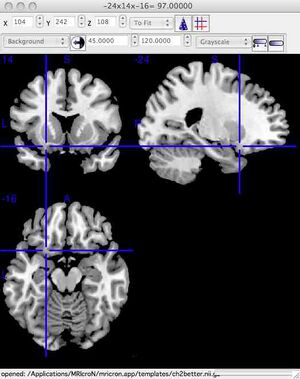 For Linux (or if the above instructions don't work): please go to http://www.mccauslandcenter.sc.edu/mricro/mricron/install.html You are on your own for installation! REQUIRED Step 2: Explore the brain and learn structures found in labs 8 and 9Regardless of the platform (PC or Mac), the top left of the window shows a coronal view of the brain. The top right of the window shows a sagittal view of the brain. The bottom left of the window shows an axial (horizontal) view of the brain. Click the mouse at any brain location to move the blue viewing crosshairs. The dataset name is listed at the top of the MRIcroN window in Windows, at the bottom of the window for Mac. The correct brain to view is the "ch2" file (ch2.nii.gz). Load this file by selecting "File/Open/Templates/ch2.nii.gz". At the top of the program window are three boxes labelled "x", "y" and "z" that show the brain location. Enter each of the co-ordinates (x,y,z) listed below into MRIcron. This will take you to the structure whose name is on the first column. This material will make up part of the testing materials for these laboratories (lab exams). Name of structure / Coordinates of structure (enter into x,y,z boxes in MriCron)
this table was created with the converter at http://excel2wiki.net/index.php Structures in lab 11Limbic System
Teaching Aid: Colored Overlay to show different anatomical regionsWhile the above co-ordinates show examples of single locations and their label, it can also be nice to see the complete extent of an area. This can be done as follows:
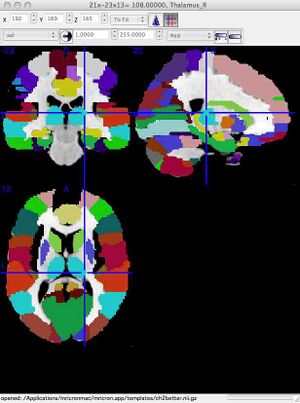 Surf around the brain. When clicking on a colored region, you will see the anatomical label for that colored area of the brain. The label (e.g. Putamen_L for left putamen) will appear in the bottom of the viewing window on PC, in the top of the viewing window on Mac. OPTIONAL Step 3: Different kinds of brain MRIMRI is clinically very useful because it is non-invasive and safe. Because no exposure to radiation is involved, patients can receive many different kinds of MRI imaging procedures without risk. The MRI scanner can be programmed to create different kinds of images that reveal different tissue properties. The three main kinds of images are T1, T2 and Proton Density (FLAIR). The names of the image types refer to the physical priniciples used to create the images. The three links below link to three datasets, all collected from the same normal subject, that show the three different image types. Click on each link and save the file to your computer.
After you have downloaded the datasets, you can load and view them in MriCroN by clicking File/Open. Navigate to the directory where you saved them and select the dataset name. for more information on MRI imaging priniciples, see http://spinwarp.ucsd.edu/NeuroWeb/Text/br-100.htm OPTIONAL Step 4: Different kinds of spine MRIBecause the spine is so long, it is typically imaged in three separate scans covering the cervical, thoracic, and lumbar+sacral portions of the cord. The following archive file contains T1, T2 and IR datasets for each spine portion. Download the file and unzip the 12 datasets that it contains. It is easiest if they are unzipped to the same directory as the datasets in Step 3. The datasets can then be opened in MRIcroN. OPTIONAL Step 5: Cerebral VasculatureHere are two movies that show the results of a Magnetic Resonance Angiogram (MRA). Click to download the movies to your desktop. Then double-click on the movies to view them. They have been tested with Apple Quicktime, free software downloadable from www.apple.com. After the movie is loaded, press the left and right arrow to scroll through the frames of the movie.
License Information for MRIcron software. Chris Rorden's MRIcron, copyright 2012, all rights reserved. Redistribution and use in binary forms, with or without modification, are permitted provided inclusion of the copyright notice, this list of conditions and the following disclaimer is provided with the distribution: Neither the name of the copyright owner nor the name of this project (MRIcron) may be used to endorse or promote products derived from this software without specific prior written permission.This software is provided by the copyright holder "as is" and any express or implied warranties, including, but not limited to, the implied warranties of merchantability and fitness for a particular purpose are disclaimed. In no event shall the copyright owner be liable for any direct, indirect, incidental, special, exemplary, or consequential damages (including, but not limited to, procurement of substitute goods or services; loss of use, data, or profits; or business interruption) however caused and on any theory of liability, whether in contract, strict liability, or tort (including negligence or otherwise) arising in any way out of the use of this software, even if advised of the possibility of such damage. |