Beauchamp:iELVIS
Beauchamp Lab wiki
|
To search for things on the wiki, use Google's site search feature. For instance, to find an Experiment Sheet, type
site:openwetware.org/wiki/Beauchamp ExperimentSheet
Overview
This page describes how to use iELVis, as documented by Buffy Nesbitt in August 2019.
Updates:
Nov. 15, 2019: added alternate version of FLIRT command.
What is iELVis?
iELVis stands for Intracranial Electrode Visualizer. We use the iELVis software package to visualize electrode locations in our research patients along with a number of other programs. iELVis is not a self-contained program but runs in a combination of BASH script, BioImage Suite, and MATLAB and can be downloaded from GitHub at:
https://github.com/iELVis/iELVis
Setting up iELVis
General Notes
Setting up and using iELVis requires a little bit of knowledge about general programming in a UNIX-type system and in an advanced language like MATLAB. For iELVis to work properly, multiple programs need to be able to access each other and work in harmony, which may require a little bit of troubleshooting depending on the setup of your individual system. This document contains the information needed to run iELVis on a BCM beauchamplab computer, but another system may require different commands or even different programs.
Currently, iELVis is only developed for MacOS. While it may be able to run on Linux, it is not developed for Windows systems. Sections marked in brackets are variable by computer, like the installation folders of certain programs or patient codes. The correct input will depend on your system and the process you're trying to run.
Prerequisites
FreeSurfer and FSL
FreeSurfer is a software package for analyzing and visualizing neuroimaging data. Install the program according to instructions at the FreeSurfer wiki:
http://freesurfer.net/fswiki/DownloadAndInstall
Installing FreeSurfer should also set up FSL, its viewing program, but if it doesn't install automatically, FSL can be downloaded and installed separately at:
https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/FslInstallation
FreeSurfer requires a license to use, available for free on the FreeSurfer site. The license.txt file must be in the /Applications/freesurfer folder for the program to run.
BioImage Suite
BioImage Suite is a software package for visualization of bioimaging data, mainly developed for abdominal/cardiac imaging and neuroimaging. Currently, the version we use is BioImage Suite 35; a web version is available but is set up differently from our preferred version. The required version of this software is stored in the BCM ECoG server and can be installed using terminal commands:
cd /Volumes/ecog/Foster_Lab/CODE/Toolboxes/Bioimagesuite_35 sudo sh ./bioimagesuite-35_0b1_06_Apr_2017-MacOS-c++-x86_64.sh -prefix= ~/
MATLAB
MATLAB is a statistical analysis program that requires a license to use. BCM has campus-wide licenses for students and faculty that provides access to the latest versions of MATLAB though iELVis can be run on any a/b version from 2016 or later. Downloads and licensing information can be found at:
https://www.mathworks.com/products/matlab.html
To connect MATLAB and FreeSurfer, you'll need to add the FreeSurfer pathways to a startup.m file that will run upon opening. Lines that begin with a percent sign (%) are comments; you can enter whatever text you'd like to remember that these lines are for setting up FreeSurfer. Create a new script in MATLAB and enter:
%FreeSurfer Setup
fshome = getenv('FREESURFER_HOME');
fsmatlab = sprintf('%s/matlab', fshome);
if (exist(fsmatlab) == 7)
path(path,fsmatlab);
end
clear fshome fsmatlab;
FreeSurfer's FsFast is a toolbox for analyzing MRI/fMRI data and should also be set up in the startup.m file as follows:
%FreeSurfer FAST
fsfasthome = getenv('FSFAST_HOME');
fsfasttoolbox = sprintf('%s/toolbox', fsfasthome);
if (exist(fsfasttoolbox) == 7)
path(path,fsfasttoolbox);
end
clear fsfasthome fsfasttoolbox;
Save the file as startup.m and MATLAB should run it automatically upon launching.
Dcm2niix and MRIcroGL
Dcm2niix is a console command that converts CT and MRI data stored in .dcm files to NIfTI (read: nifty) .nii format which is readable by FreeSurfer and other viewing programs. Avoid the older version dcm2nii; it is no longer supported.
The dcm2niix package is available here and is installed through Python 2.0+:
https://github.com/rordenlab/dcm2niix
MRIcroGL is a neuroimaging viewing program that contains the latest version of dcm2niix as well as a number of image editing options for CT and MRI data. This is our preference because it allows immediate viewing of the converted .nii and gunzipped .nii.gz files.
Download and install MRIcroGL according to the instructions here:
https://www.nitrc.org/plugins/mwiki/index.php/dcm2nii:MainPage
BASH
BASH commands are UNIX-type commands used in the MacOS terminal window. BASH uses profiles which can be saved with file pathways and setup commands that will run upon opening that profile. We generally recommend .bashrc, .profile, or .bash profiles and this document will refer to first as the default. While .tcsh can be used, be aware that the syntax it uses is slightly different. To access a profile, enter the terminal and enter:
nano ~/.bashrc
You may need to use sudo nano ~/.bashrc if admin permission is required, in which case you will need a password. iELVis requires that all necessary files be on the same pathway and that that pathway can be accessed by BASH, MATLAB, and BioImage Suite alike. Entering these lines in .bashrc will allow these programs to setup the correct pathway. These lines should be marked by a comment line (to write a comment in BASH, enter an octothorpe (#) followed by your text) to denote what pathways they're setting up.
#SETUP iELVis Freesurfer export FREESURFER_HOME=/[location of freesurfer folder] source $FREESURFER_HOME/SetUpFreeSurfer.sh
#SETUP iELVis BASH export PATH=$ PATH: [location of iELVis folder]/iELVis_master/iELVis_MAIN/iELVis_BASH/ export PATH=$PATH: [location of iELVis folder]/iELVis_master/iELVis_MAIN/iELVis_MATLAB/
Starting Electrode Localization
Converting .dcm to .nii
Before visualizations can start, the CT and MRI scans need to be in .nii or .nii.gz format. Most CT and MRI scans are developed as .dcm (read: DICOM) images, one for each slice, and need to be compiled by a conversion program like dcm2niix.
In our lab, we work with patients receiving care at St. Luke's Episcopal; the hospital stores patient scans on disk in the Radiology Film Library on floor B1. It can take a day or so for the Film Library to get the scans, so be prepared by calling ahead at (832)355-2081 and asking for the pre-operative MRI and the most recent post-operative CT scan. If you have a disk, be sure to copy the files into a designated folder in the subject's name on the beauchamplab and/or ecog servers.
We use OsiriX Lite to sort, export, and anonymize patient DICOM files. (See the "Useful Links" section for the website.) OsiriX reads in the DICOMs from disk and sorts them by header information into individual scan types. Often multiple scans from the same session are included; only the T1 and post-op CT are necessary for the iELVis pipeline. The T1 should be named "T1_3D_FFE" and contain about 150-250 DICOMs (note: T1_3D_post_FFE is not the same--the "post" refers to the application of a contrast dye that will make the image unusable by FreeSurfer--nor is any scan labeled with "Cor" (coronal), "Ax" (axial), or "Sag" (saggital), which are partial scans. On some modern machines there may be an e3D T1 that is similarly problematic for reconstruction). OsiriX also includes an anonymization feature that removes patient information from the headers of the DICOM files and exports them as a single folder to a target location. We perform both of these processes before converting the files to NIfTI format.
If using dcm2niix as a console command, open a terminal window and enter:
dcm2niix [folder with DICOM images]
This is the minimum requirement to run the command; to see what other functions or personalizations the command can perform, enter:
dcm2niix -help
If using MRIcroGL, open the software, select "Import" from top right menu, and select "Convert DICOM to NIFTI". This will open a new window. In this window, select "Edit" from the top right menu. In the popup window, find the folder containing the MRI or CT .dcm files and select "Open."
Both of these methods should automatically convert the files into a .nii.gz file which can be unzipped to get the .nii file or, for many programs, read as-is. It will also create a .json file of the scan. Be sure to appropriately name these files with the format subID_CT.nii/.gz or subID_MRI.nii/.gz as the next command utilizes this naming scheme.
Surface Reconstruction (aka The Long Step)
The first step of electrode localization is to reconstruct a 3D image of the patient's brain from their pre-operative MRI. There are very few programs that do this, and even fewer so time-tested and simple as FreeSurfer's recon-all command, which will run the complete process with a single line. That said, recon-all is not a simple program; it is an enormous script that has no less than 31 sub-steps, some of which call to multiple commands, and can take as long as 24 hours on a well-equipped machine (or longer if there are errors). More detailed information about the recon-all command is available on the FreeSurfer Wiki (see Useful Links).
The bare minimum needed to run recon-all is a target folder, a subject's name, and an input MRI in .nii format within the target folder. The target folder is the folder in which all the subject's reconstruction and later localization data will be stored. This location is saved as the variable $SUBJECTS_DIR. By default, FreeSurfer uses its own /subjects folder as the target folder, located at /Users/[username]/Applications/freesurfer/subjects if installed in the default location, but can be changed to any folder by exporting the SUBJECTS_DIR variable as a new location (see the commands below). Recon-all accepts the flags -s, -sid, -subjid, and -subject to identify the subject's name which, in the Beauchamp lab, is generally a three-letter code beginning with Y such as YAI or YCK. The command can also accept more than one reference MRI as input flagged with -i [filename].nii, should more than one T1 scan exist, and can even utilize T2-type scans in later processing steps to improve the final surfaces. An MRI in .nii.gz format does not need to be unzipped to use as input.
With all of this said, the minimum call to recon-all is as follows:
cd /[target folder]/ export SUBJECTS_DIR=/[target folder]/ recon-all -s [subject ID] -i [subject's MRI].nii -all
FreeSurfer breaks the recon-all command into three main portions: autorecon1, autorecon2, and autorecon3. Autorecon2 is broken down further into autorecon2-cp, autorecon2-wm, and autorecon2-pial. Using the flag -all or -autorecon-all will run all of these steps in order. To run or re-run a section of the command, flag the section to be run (i.e., recon-all -s [subject ID] -autorecon2). If the subject has a previous reconstruction, do not include the -i flag; FreeSurfer will return an error that it will not overwrite the data.
The call we use for iELVis is as follows:
cd /[target folder]/ export SUBJECTS_DIR=/[target folder]/ recon-all -s [subject ID] -i [subject's MRI].nii -all -parallel -localGI
There are only two additions to the minimum command that we use, the flags -parallel and -localGI. The -parallel flag commands FreeSurfer to parallelize its actions, utilizing more memory and CPU power but significantly speeding up the process. This isn't a necessary flag; on a computer with less than 32GB RAM, or on a computer that will be running other tasks while the reconstruction takes place, it may be better not to use it. The flag -localGI refers to the Local Gyrification Index, a calculation performed after autorecon3 that provides a ratio of how much of the pial surface is gyral vs. sulcal. iELVis will utilize this ratio in projection steps. This step requires MATLAB to run, so be sure that both the BASH pathway and the MATLAB startup.m files are correctly set up before running.
Freeview, the imaging program that comes with FreeSurfer, can be used to check the output of the command and identify problems as they occur. Recon-all is a lengthy, complex process with numerous sub-steps; while the command above is perfectly acceptable and will provide everything needed to continue localization through iELVis, it can also be broken down into smaller steps for easier troubleshooting.
CT alignment
Coregistering the CT and MRI
iELVis uses a function called ct2mri.sh to coregister the CT and MRI that is based on FreeSurfer's flirt command. To run iELVis's coregistration, enter the commands below:
cd /[folder_location]/ export SUBJECTS_DIR=/[target_folder]/ ct2mri.sh [target folder name] subID_CT.nii
This step can be done while the recon-all step is still running as long as the T1.mgz file has been created (found in the subject's "surf" folder). The aligned CT will be saved as postInPre.nii.gz and two .gif files will pop up showing the coregistration. Check the alignment of the CT and MRI in these files. If they're misaligned, the postInPre.nii.gz is also misaligned and the file won't be useful the next steps.
An important note: part of the ct2mri.sh process is the creation of a T1.nii.gz from the T1.mgz. This new T1 is de-obliqued, meaning that it has been shifted from the straight X, Y, and Z axes. The alignment will be done with this version of the T1. Keep in mind that, if at any point you need to re-align the images or reconstruct the MRI again, that there is an axial difference between the subID_MRI.nii input into recon-all and the T1.nii.gz that that process outputs into the subject's "surf" folder.
The ct2mri.sh command will also create the elec_recon folder in the subject's FreeSurfer folder that iELVis will use to store a number of files with the patient's electrode localization info, including electrode coordinates and labels, and later steps will use this folder as both storage and an access point for those information files. However, if you'd like to perform a coregistration of two .nii images without setting up the elec_recon folder, you can run FreeSurfer's flirt command on its own:
flirt -in [subID_CT].nii -ref [subID_t1].nii -out [output file name].nii -omat ct2t1.mat -interp trilinear -cost mutualinfo -dof 6 -searchcost mutualinfo -searchrx -180 180 -searchry -180 180 -searchrz -180 180
These are the parameters used by ct2mri.sh during its call to flirt but they can be adjusted as needed; the general flirt command can also be used to coregister multiple MRIs or CTs based on the input and reference file. Similarly, the degrees of freedom can be adjusted from 6 and the angle range from -180 to 180 as desired. A less strict coregistration for problematic alignments can be done by increasing the degrees of freedom to 12 and decreasing the angle range from -90 to 90.
For quick localizations, we use FLIRT with the following parameters as a faster command:
flirt -in [subID]_CT.nii -ref [subID]_MRI.nii -out [output file name] -bins 256 -cost corratio -searchrx -90 90 -searchry -90 90 -searchrz -90 90 -dof 12 -interp trilinear
Electrode Localization
Patient Files and iELVis Naming
Before you begin the actual localization process, you'll need some information about the patient to correctly identify the electrodes in their scans. At a minimum you'll need the patient's montage file listing the inserted electrodes and the aligned CT from the previous step. If surgical files, implant drawings, electrode diagrams, intra-operative photographs, or any other information is available it should be gathered and used for the next steps to ensure the localization is correct.
At BCM, we keep patient files with the surgical diagrams, CT/MRI discs, clinical and research montages, and other data in folders under lock in room S 104 F. Contact Dr. Brett Foster and/or Dr. Bill Boskings for access. Much of the information in these folders is also available on the beauchamplab and ecog servers with identifying information removed.
We use both AdTech and PMT brand electrodes for our electrocorticography experiments. These brands produce very similar electrodes of equal quality, but there are sometimes differences in numbering, electrode size/shape, and spacing between electrodes, depending on the model used. Diagrams of the electrodes can be useful in localization and are available at their websites:
AdTech: https://adtechmedical.com/ PMT: http://www.pmtcorp.com/index.html
iELVis refers to all electrodes, be they grid, strip, or depth, as "grids" and each individual electrode channel as a "contact"; the rest of this section will do the same. iELVis uses a naming system that gives each grid a unique name as well as a tag identifying its location on the left or right hemisphere and whether the model grid is a grid, strip, or depth electrode. This is written as an "L" or "R" for left/right, a "G", "S", or "D" for grid/strip/depth, an underscore, and the electrode name from the patient's montage (e.g., LG_Grid, RS_PTO, LD_LIns). Each grid must have a unique name not including the LS/RD tag-- i.e., if there are two grids with the same name on each hemisphere, their names should include the left/right designation, not just the tag (e.g., for two amygdalal probes, use LS_LAMY and RS_RAMY, not LS_AMY and RS_AMY)-- two grids with the same name will cause errors later in MATLAB when trying to plot these electrodes.
BioImageSuite Settings
The actual localization process is done in BioImage Suite. The version we use is launched through the terminal with the command:
start_bioimagesuite
The BioImage Suite menu should appear with a left-hand column of menus and options within each appearing in the larger right column. Electrode identification and placement is done through the Electrode Editor, accessed from the "Editors" menu to the left, and launches in two windows called the Electrode Editor and the Electrode Control windows. The Electrode Editor window will display the subject's aligned postInPre.nii.gz and the right will be used to create the patient's .mgrid file containing the names and locations of the electrodes.
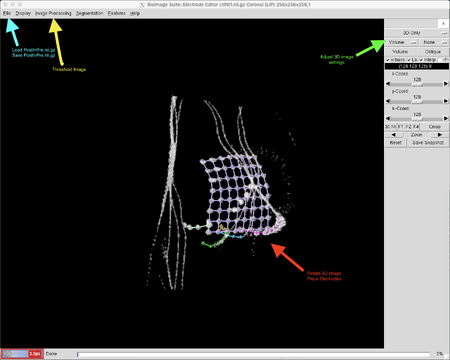
First, load the subject's postInPre.nii.gz into the Electrode Editor window. Go to "File" then "Load" and use the directory to select the correct file. There are a number of views in which BioImage Suite can display this 3D image, accessible from the right-hand menu "General View" and the options menus below it. To view a rotatable, black-and-white 3D image of the skull and electrodes, set the "General View" to "3D only" and the "View Options" below it to "Volume." You can remove the green orientation box around the image by selecting "Orientation Options" and setting that to "None." This will display the electrode contacts as small white dots in the thin grey-white skull as well as the white teeth and any wires in the CT scan as grey-white lines. Click and drag the image to rotate it and click "Reset" at the bottom of the left menu to return the image to its original orientation. Hitting "Reset" will only reset the image's position; it will not erase any electrode data.
In some cases, especially with depth electrodes, it may be more useful to view the electrodes in sagittal, coronal, and axial views aligned side-by-side; to do this, set the "General View" to "Simple Mode" and the "View Options" to "3-Slice Mode." The background will be displayed in cyan and electrodes and bones in bright red. To navigate this 2D image, click and drag the crossbar through one view to adjust the displayed slices in the others.
For any view, the image will need to be processed to adjust the contrast and make the electrodes more visible. Using the top menu, select "Image Processing" and then "Threshold" to open the Image Threshold window. Most patients in 3D view have a high threshold of about 3000 and a low threshold of about -1000 though these numbers, especially the former, can vary greatly by scan. Adjusting the low threshold to about 1000 will reveal the electrodes and the surrounding skull; adjusting it to about 2000 will show only the electrodes, wires, and teeth. It's worth playing with these settings a little when first loading a patient into BioImage Suite to get the clearest view possible. You can adjust these settings at any time from the Image Threshold window to make individual contacts more visible.
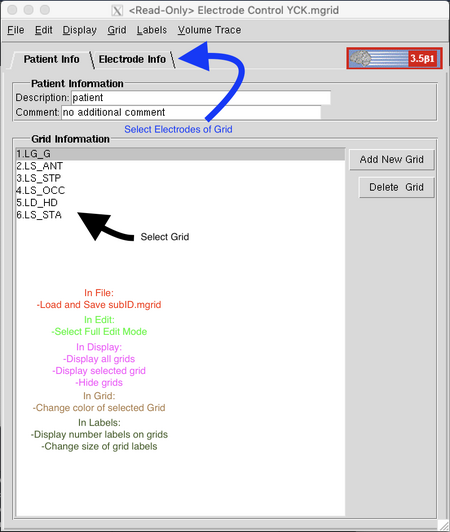
Settings must also be adjusted in the Electrode Control window. At the top menu, select "Edit" and then "Full Edit Mode" to remove the Read-Only restriction. Only one electrode strip/grid will be shown at a time in the Electrode Editor window unless "Display All" is selected from the "Display" menu. If loading a patient with a pre-existing .mgrid file, load that file by selecting "File" then "Load" from the upper left menu and using the directory. This window has two tabs, Patient Info and Electrode Info. The first will display a list of electrode strips, grids, or depth electrodes by name; the second allows you to edit the information and contact placement of the selected electrode.
Electrode Selection
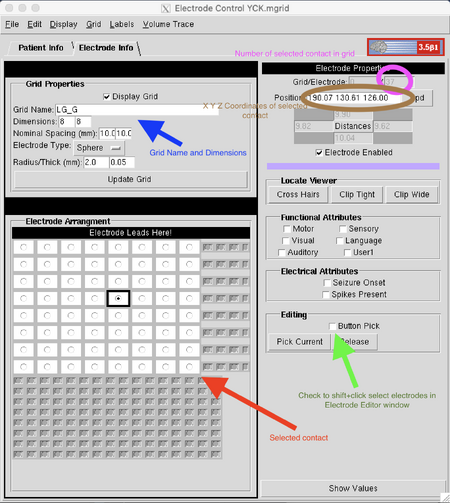
To begin electrode selection, select a grid from the Grid Information box of the Patient Info tab or click "Add New Grid" to the right to add another grid to the list. With the grid selected, enter the Electrode Info tab. This tab is divided into three boxes, "Grid Properties" to the upper left, "Electrode Arrangement" to the lower left, and "Electrode Properties" to the right. In the Grid Properties box, enter a name for the grid that matches the patient's montage and includes the LS/RD tag. Below that, enter the dimensions of the grid with the smaller number first (i.e., 4x8, not 8x4) and click "Update Grid." A dialog box may pop up warning that changing the grid dimensions will erase all placement information and asking to confirm this choice; all this means is that changing the grid dimensions will remove any placed contacts. Hit "yes" to save the changes. You can also change the color of the grid in the "Grid" menu at the top by dragging the arrows beneath the Red, Green, and Blue sliders or entering a value between 0 and 255 into the boxes for each bar. This is useful for creating contrast between multiple grids and between grids and a colored brain image.
Now you can align the model grid with the electrodes on the patient's scan. Under Electrode Properties there is a small box labeled "Editing" and a checkbox labeled "Button Pick." Check this box and then select a contact from the grid in the Electrode Arrangement box. Notice that, once a contact is selected, the upper section of the Electrode Properties menu will fill in with the number of that contact out of the total number of contacts on the grid, coordinates of that contact, and its distance to its neighbors in the grid. With that contact selected, Shift+click the corresponding contact in the Electrode Editor window. The grid contact should snap to the center of the shape. If it snaps incorrectly, click away from and back to the contact in the Electrode Arrangement box and Shift+click the desired place again, or adjust the Image Threshold settings and try again. You can also manually adjust the coordinates of the contact by altering the X, Y, and Z coordinates in the Electrode Properties box. Click the next contact in the Electrode Arrangement grid and repeat until the entire grid has been placed. Do this with all the grids listed in the patient's montage.
When localization is complete, save the .mgrid file in the target folder from the Electrode Control window and name the file [subID].mgrid so that it can be found later by MATLAB scripts expecting this name. Remember that saving the postInPre.nii.gz will NOT save the information in the Electrode Control window; the .mgrid file needs to be saved separately.
Electrode Projection and Visualization
The iELVis projection and visualization steps are run in MATLAB. While the basic projection steps only require built-in features, more advanced projection and analysis may require scripting by the user. The iELVis package installed from GitHub contains all of the commands that are used in the Basic Projection steps below. For those steps of the Advanced Projection section that require scripts not included in the original iELVis package, there will be a link provided for each script. You can also find these in the "Scripts" section.
Basic Projection
Open MATLAB and globalize the subject's iELVis folder to set the patient's directory:
global globalFsDir globalFsDir = '/Location of iELVis localization folder/'
The next script creates eight files--seven will be named [subID] with the extensions: .electrodeNames, .INF, .LEPTO, .LEPTOVOX, .PIAL, .PIALVOX, .POSTIMPLANT and the last will be [subID]PostimpLoc.txt--based on the BioImage Suite localization .mgrid file. If the .mgrid file has been edited since the last projection, this step will need to be run again to overwrite these files with the updated information. All of these files can be found in the subject's elec_recon folder.
makeIniLocTxtFile('subject ID')
Projection and Brainshift Methods
There are two methods of electrode projection built into iELVis, the Dykstra et al. method and the Yang, Wang et al. method, that correct for brain shift. Both will project the post-operative electrode locations to the pre-operative brain. To launch the Dykstra et al. method, enter:
dykstraElecPjct('subject ID')
The Dykstra et al. method projects each electrode to a smoothed pial surface (functionally a dural surface) before projecting to the regular pial surface to maintain the original distance between each contact and its nearest neighbors. When running this command, a figure will appear showing the optimization process of the function followed by the shift distribution of the electrodes on the subject's brain.
To launch the Yang, Wang et al. method, enter:
yangWangElecPjct('subject ID')
If there is an electrode tagged as a grid, the command window will prompt for its dimensions and orientation. Enter the smaller dimension first (i.e., 4x8, not 8x4) and enter the order of the grid's corners as specified. If there are multiple grids, the function will require the dimensions and orientations of each. This method projects strips to the nearest point on the dural surface and projects grids to the dural surface by inverse gnomic projection. This should produce two figures. The first will have on the right a plot of Euclidean differences between each grid contact and its neighbors and on the left the standard deviation of contact distances. This function includes a number of tested parameters to produce the lowest difference in known distance between contacts and the lowest standard deviation of those differences. If none of those parameters work (which may indicate an error in electrode placement or grid orientation), it will prompt the user to elect the best parameter by selecting the contact from its displayed list with the lowest standard deviation. The second figure it will produce is the shift distribution map.
After running either projection function, display-ready images of the electrodes on the subject's brain can be generated by running:
plotMgridOnPial('subject ID', 1)
Advanced Projection
AFNI and SUMA
While not a requirement for localization in iELVis, AFNI and its viewer SUMA are long-time standards of the neuroscience world and almost ubiquitous in their use. AFNI contains its own electrode localization programs and being able to convert between the two pipelines is a useful feature. For this reason, we've included a few steps to make iELVis output data viewable in AFNI and SUMA.
To install AFNI and SUMA, follow this link:
https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/background_install/download_links.html#download-current-precompiled-afni-binaries
Documentation for AFNI, SUMA, and a number of other useful software packages for neuroscience can be found here:
https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/background_install/main_toc.html
Converting Files between iELVis and AFNI/SUMA
If using AFNI/SUMA to view patient data, you'll need to create a set of specification files for those programs to read. These files will appear in a folder called "SUMA" within the subject's folder. Open the terminal and enter:
cd [subject's folder] @SUMA_Make_Spec_FS -NIFTI -sid [subID]
This should also make specification files for the AFNI/SUMA's standard 141 brain for comparison.
Alternatively, to use AFNI's GIFtI format, the same command can be entered with the flag -GIFTI instead of -NIFTI.
Converting Electrode Coordinates between iELVis and AFNI/SUMA
iELVis does not display brains in the same orientation space as AFNI/SUMA, so to view brains in the latter (or in any program that uses the same orientation) the coordinates must be converted. We use the script afni2elvis.m to convert input coordinates from native FreeSurfer/iELVis space to AFNI/SUMA space.
In addition, the iELVis outputs, especially for grids, may not be in the same order as the patient's montage; in particular, iELVis will list grid contacts by column, not numerical order. An updated form of the afni2elvis.m script called afni2elvis_reorder.m will both convert the coordinates between FreeSurfer and AFNI/SUMA space and reorder the labeled coordinates according to the patient's montage. The electrode names in iELVis must match the names in the montage (not including the contact number or the LS/RD tag) for the script to work.
Mapping Electrodes to Average Brains
The iELVis wiki has detailed instructions on plotting electrodes on an average brain in MATLAB here:
http://ielvis.pbworks.com/w/page/117733770/Mapping%20Electrode%20Locations%20to%20Average%20Brains
Built into iELVis is the command sub2AvgBrain (see above for details) that will take the electrode names and coordinates stored in a patient's [subID].PIAL file and plot them on the fsaverage brain. We have a slight variation on this command named sub2SubBrain.m that will also output a patient's coordinates in their original (untransformed) space instead of the fsaverage space and can plot them on their own brain.
Projecting to Brains in RAVE
RAVE (R Analysis and Visualization of Electrodes) is a statistical analysis program for electrocorticography data developed by members of the Beauchamp lab. It contains the ability to show electrodes on a patient's brain in 3D in coordination with the analysis of that electrode data, but it requires a few prerequisites to be able to do this.
Currently, RAVE requires the output of @SUMA_Make_Spec_FS to create 3D surface files for viewing. See above for the full command and its output.
Once the SUMA folder is populated, enter the subject's RAVE folder and copy in the /label, /mri, /SUMA, and /surf folders from the patient's reconstruction. In the /meta folder, there is a file named electrodes.csv; this file contains the coordinates, labels, and other tagging information for each electrode displayed on the brain. If you have the subject's electrode coordinates in FreeSurfer space (as output by the scripts in the section above), add that information to this file. RAVE supports the addition of a "Label" column with the patient's montage information to identify each electrode as well as user-added columns with further information, such as anatomical group or white/grey matter labels.
In the subject's rave/fs/SUMA folder, you'll need an ASCII version of the sulcal dataset for RAVE to read. Launch the Terminal and run the following commands:
cd /[sub's RAVE folder]/fs/SUMA ConvertDset -o_1Dp -input std.141.rh.sulc.niml.dset -prefix std.141.rh.sulc ConvertDset -o_1Dp -input std.141.lh.sulc.niml.dset -prefix std.141.lh.sulc
Launch RAVE through RStudio and load the patient's experimental data. RAVE creates and caches the files it requires when a subject is first loaded, a process that may add a few minutes to the initial loading. Once this is complete, however, all subsequent loadings will use the same cached data and load within a matter of seconds. If any information is updated/changed between loadings, expect a short delay when loading a subject's information.
Scripts, Programs, and Other Links
iELVis requires a lot of components and at the Beauchamp lab we have developed our own scripts atop these to further work with the electrode localizations. For ease of access, this section contains links to all of the programs, scripts, and links provided earlier in this page.
Programs
iELVis package: https://github.com/iELVis/iELVis
FreeSurfer: http://freesurfer.net/fswiki/DownloadAndInstall
FSL: https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/FslInstallation
BioImage Suite (web): https://bioimagesuiteweb.github.io/webapp/
MATLAB: https://www.mathworks.com/products/matlab.html
MRIcroGL: https://www.nitrc.org/plugins/mwiki/index.php/dcm2nii:MainPage
dcm2niix console command: https://github.com/rordenlab/dcm2niix
RAVE: https://github.com/beauchamplab/rave
OsiriX: https://www.osirix-viewer.com/osirix/osirix-md/download-osirix-lite/
Scripts
The scripts provided are in compressed .zip format to allow upload to the OpenWetWare site. Be sure to decompress them before use!
afni2elvis.m-- converts electrode coordinates between FreeSurfer and AFNI/SUMA space and vice-versa.
afni2elvis_reorder.m-- runs afni2elvis.m and reorders electrode coordinates according to subject's montage. Requires afni2elvis.m to run.
sub2SubBrain.m-- can convert coordinates between a subject's native space and the FreeSurfer average brain space and display converted/unconverted electrodes on both.
Useful Links
PMT electrodes: http://www.pmtcorp.com/index.html
Ad-Tech electrodes: https://adtechmedical.com/
iELVis Wiki: http://ielvis.pbworks.com/w/page/116347253/FrontPage
FreeSurfer Wiki: https://surfer.nmr.mgh.harvard.edu/fswiki/FreeSurferWiki
FreeSurfer License: https://surfer.nmr.mgh.harvard.edu/registration.html
Recon-all on the FS wiki: https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all
Recon-all Dev Table: https://surfer.nmr.mgh.harvard.edu/fswiki/ReconAllDevTable
AFNI/SUMA Documentation: https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/background_install/main_toc.html
Troubleshooting
General Troubleshooting
Missing Files/Commands
This is perhaps the most common problem in setting up and running iELVis. The required files may exist in multiple places, on a local machine, on one or both servers, or as part of a program; it can be difficult to get all the required components of iELVis to work harmoniously. First, search for the file name in your computer and the servers. If it appears, check that the folder it's in has been added to the pathways in BASH and MATLAB. If it doesn't, check the name of the file itself to make sure that it's been named correctly (this often occurs when iELVis is searching for a file with a specific name format, such as subID.mgrid, and the file has not been named according to its expected structure). Sometimes commands and files are missing when a program isn't installed correctly; if this is the case, download and re-install the program.
FreeSurfer Errors
Export: command not found
Export is a command inherent to BASH, not FreeSurfer. If this error occurs, it is because the BASH shell being used does not contain this command, as is the case for shells like .tsch and .zsh. Open the Terminal and select "Preferences" from the upper window. Set the default shell to a BASH shell such as .bashrc and be sure that shell contains the FreeSurfer setup as described in the Prerequisites section.
Missing License File
FreeSurfer's installation does not automatically include the license.txt file. It can be found on the FreeSurfer Wiki (see Useful Links) and is free. The file should be stored in the /freesurfer folder which, if the default installation location is used, will be /Users/[username]/Applications/freesurfer.
Recon-all Exited with Errors
When run, recon-all will automatically write and store a .log file in the target folder's /[subject ID]/scripts folder. If the Terminal announces that recon-all has exited with errors, check this file first. It should identify which step of the command encountered a problem and what that problem was. Generally speaking, if this happens, recon-all will have to be re-run in whole or in part.
Troubleshooting FreeSurfer
The benefit of running recon-all as an automatic process is that it requires very little user oversight, but the resulting deficit is that without oversight of the individual steps the program is difficult to troubleshoot. The first thing to do is to check the .log files in the subject's /scripts folder. These list the written output of recon-all as it runs, errors included. If there's an error at any step of the process, even if it did not force recon-all to exit, it should be listed here.
The second thing to do is to check the FreeSurfer Wiki and the recon-all Dev table (see Useful Links). These will list every command called to by recon-all and the inputs, outputs, and parameters of each. If you can identify the error from the .log files, find that command on one or both of these sites for more information and troubleshooting.
If the subject's brain comes out lopsided, jagged, or missing parts, the issue generally has to do with the T1 input. FreeSurfer uses the visible contrast between grey and white matter found in a T1 scan to determine the location of the inner white matter surfaces and "balloons" the pial surface out to the dura from there. If the T1 has poor contrast, unusual anatomical structures (e.g., polymicrogyria, lesions, etc.), or low resolution, or if the autorecon1 process failed to remove parts of the dura or skull from the brainmask image, then the error is most likely related to how FreeSurfer interpreted the T1. Troubleshooting this type of error is complex; it requires using FreeView, FreeSurfer's built-in viewer and editor, to make edits to the T1 and files based on the T1 slice by slice. The FreeSurfer wiki includes detailed instructions on correcting these errors here:
https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/TroubleshootingData
CT/MRI Alignment Errors
No MRI or CT DICOM files
If the individual .dcm images are not available for a subject, it may be possible to recreate them using .BRIK and .HEAD files if the subject has previously been processed in AFNI/SUMA. Check the beauchamplab server for the FreeSurfer folder (generally labeled "fs") in the subject's directory. If there is a SUMA folder therein, check to see if it has .BRIK and .HEAD files. Open terminal and enter:
cd /location of .BRIK and .HEAD files/ 3dAFNItoNIFTI [file_name]+orig.HEAD -prefix [subID CT/MRI].nii
Open the resulting file in your viewing program of choice to check quality.
Permission Denied
If you forget to enter the SUBJECTS_DIR=[target folder] line, you might get this error.
FLIRT: command not found
The FLIRT alignment command is not a part of the base FreeSurfer package, but rather part of FSL. If the FLIRT command can't be found, try updating or reinstalling FSL; it may not have installed correctly with FreeSurfer.
No Brainmask
If an error appears that the ct2mri.sh command cannot find the subject's brainmask.nii.gz, it's likely that you're trying to run the command a little too early into the recon-all process (or, if recon-all has completed, that there was an error). The command will still work as long as the T1.mgz file has been created; the ct2mri.sh command only copies the brainmask.nii.gz into the elec_recon folder. That being said, check to see that the brainmask was, in fact, created by the recon-all step and is saved in the patient's elec_recon folder. If not, recon-all may need to be rerun.
Image Exception #63/22
Generally, this means that the ct2mri.sh command can't find the T1.mgz file created by recon-all. The T1.mgz can be found in the subject's freesurfer "[subID]" folder, in the "mri" subfolder. If the file does not exist, recon-all may have encountered an error and should be run again. If you don't want to overwrite the information, enter a new name for the target folder or move/delete the folder from the original recon-all. If still encountering this error, take the T1.nii.gz file and copy it to the elec_recon folder. Make sure to use the most updated form of dcm2niix to create this and be sure that the T1 is in .nii.gz format, not just .nii.
Misalignment
Misalignment is usually very visible in the .gif files created by ct2mri.sh. One of the easiest ways to tell if the coregistration is correct is to look at the eyes and nose of the patient in the .gif files. If they're crooked, incomplete, or not in the same orientation, the postInPre.nii.gz is also misaligned.
Check the CT and MRI .nii files for errors first and make sure they're in the correct format, then check the MRI to be sure it's a T1 weighted scan. A T1 will be high resolution with lighter white matter than gray matter. While other MRI scan types may align properly with the CT, it will cause problems later in BioImageSuite and MATLAB that depend on the contrast found in a T1-type scan. If the pre-op MRI is not a T1 scan, one can be made from the .BRIK and .HEAD files (see above).
Re-run the alignment with less strict parameters. This will require stepping out from the ct2mri.sh command and using the flirt command directly, then adding the resulting alignment to the elec_recon folder. See the alignment section above for the full flirt command and its parameters.
Multiple Output Files
The dcm2niix command, whether run through MRIcroGL, the console, or another program, will take all of the .dcm images of a folder and sort them by series according to information in each file's header. Most MRI sessions include more than one scan and at least one localizer. Programs like OsiriX can identify this information and export each scan as a single folder, each of which will output its own single .nii.gz/.json pair, so it is highly recommended for time's sake to process the DICOMs in this way before running dcm2niix.
If the T1 or CT are split among multiple scans, dcm2niix has a merge flag (-m y) that will combine multiple scans with different header information into a single output. Further details on this and other helpful flags can be found in the command's help output (dcm2niix -help).
Output Includes ctINt1.nii.gz
If browsing through subjects on the Beauchamp Lab server, you may see that many patients have this file included in their elec_recon folders. A previous version of iELVis outputs this file along with the postInPre.nii.gz and postImpRaw.nii.gz files in its call to ct2mri.sh. There is no difference between ctINt1.nii.gz and postInPre.nii.gz (though postImpRaw.nii.gz is different and should not be used) and, if both occur, both can be used for electrode placement in BioImage Suite. Be aware, however, that if ctINt1.nii.gz is being output, then you are not using the most recent release of iELVis.
Electrode Placement
Can't Find Electrodes/Images Do Not Match Surgical File
First check that you've loaded the correct postInPre.nii.gz and .mgrid file. We recommend naming both files with the subject's identifier and making sure that they match each time you open a new file or switch patients in BioImage Suite. Also check that you have the correct patient's files for reference (surgical files, etc.) and that the scans used to create the postInPre.nii.gz file were the most up-to-date. If using intra-op images for reference, be sure to know the orientation of the patient's head in relation to the camera to avoid rotation errors.
Sometimes the surgical files include a diagram of the electrodes used. Often subdural electrode grids will have a platinum marker on them that will show up in a CT scan and in BioImage Suite that can be used to ensure the orientation of the grid. The electrode producer's website should have information about each product including a diagram with numbered contacts, marker locations, and wire placement.
Electrodes shift after placement; some electrodes may slide along the surface of the brain or move within the tissue. This can cause electrodes to end up a few millimeters from their original implant location and to a limited extent should be expected. The electrode projection methods described above correct for this type of brainshift.
In the case of depth electrodes, improper tightening of the anchor bolt that holds the electrode in place can cause the entire electrode to shift outward from the brain, sometimes leaving contacts in the anchor bolt itself. Often if it seems a depth electrode is missing a contact, it may have slid outward slightly.
If there is a major issue with any element of the patient's setup--mislabeled electrodes, a loose anchor bolt, multiple missing contacts, etc.--contact the patient's surgical team immediately. Provide evidence of the issue's nature and location so it can be quickly identified. This issue is extremely rare but should never be dismissed lightly; if there seems to be a problem, report it or check it out just to be safe.
MATLAB Projection
Errors in makeIniLocTxtFile
Check that the globalFsDir was indeed globalized. This step is dependent on the directory entered as globalFsDir; it cannot find any files not on that path. Check that the file or folder for which it's searching is correct.
Errors in yangWangElecPjct
Sometimes the Yang, Wang et al. projection method will display a warped grid with high standard deviations. If this is the case, re-run the function with slightly different parameters. All grids need to be entered with the smaller dimension first. Accept the default orientation of the grid's corners; it should only affect the labeling of the grid in future images.
If the error "String X# not found in cell array" appears, it means that MATLAB can't find electrode # of grid X. Check labeling first as the issue may be as simple as mixing up the names of the grids. Then check BioImageSuite grid placement that all electrodes have been placed correctly and that the names of the grids are correct.
Sometimes the function will fail to produce the shift distribution maps. Generally, when this occurs it is because the grids were not given unique names or were improperly tagged. If this is the case, rename the electrodes in BioImageSuite and rerun makeIniLocTxtFile to update the files created with the new names.
If the electrodes are projecting to the wrong hemisphere, check that the LS/RG tag is referring to the correct hemisphere. A good warning that this will occur is that the number of iterations of the projection is high (>20) or that the means of the iterations appear as "NaN." If the problem does not lie in the tag, check the parameters of the yangWangElecPjct script and the labeling of grids in BioImageSuite.
Errors in plotMgridOnPial
An odd error reading "Error using *" may appear if you try to run this step after the Yang, Wang et al. function does not produce the shift distribution maps. See above for the correction.
If the brain looks lopsided, jagged, or not like a normal brain should, it may be that the CT/MRI alignment was not completely aligned or not done with a T1 weighted scan. Check the original .nii files used in ct2mri.sh and if necessary re-run the alignment. It is also possible that the FreeSurfer reconstruction encountered errors, in which case recon-all will need to be re-run in steps to troubleshoot the issue.