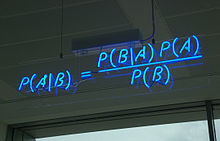BME103:W930 Group6
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |
OUR TEAMLAB 1 WRITE-UPInitial Machine Testing The Original Design For the lab, we used an Open PCR machine to amplify parts of the DNA through polymerase chain reaction. The OpenPCR machines are used to amplify a particular segment of the DNA. Multiple copies are needed to properly understand and work with it. Not much DNA is needed to make the replications.
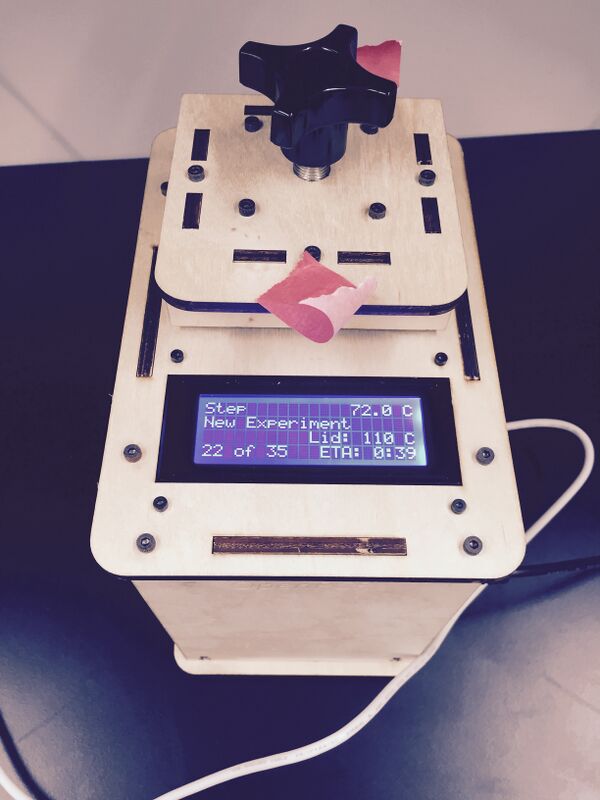
The first step is to extract DNA and place it in a PCR tube. Next, add Primer 1 and 2, which attaches to the part of the DNA that we want copied. Next add nucleotides and DNA polymerase. The DNA polymerase is the part that creates the copies of DNA. The PCR machine has a storing dock for DNA samples, which is also a heating plate. A laptop connected to the PCR machine controls the different temperatures and how long they need to last. The heating plate raises and lowers the temperature depending on what is needed for a particular step in the DNA replication process. This process of replication is repeated thirty times. By the end of the thirtieth cycle, there are millions of copies of the sequence needed for analysis. (Content by Sairah F, edited and posted by William L).
Experimenting With the Connections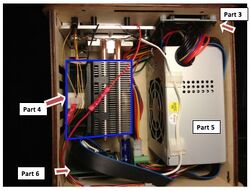  Part 3: LCD screen Part 4: Heat Sink Part 5 Rsenda ATX power supply Part 5 OpenPCR circuit board (Content by Sairah F When we unplugged the LCD screen (part 3) from the Open PCR circuit board (part 6), the screen went blank the machine had no visual output.
We first tested the PCR machine on Wednesday October 24, 2012. The easiest part was dealing with the set up of the cycles for the reaction. The program was pretty self explanatory and very easy to use once you knew all of the parts. The PCR machine itself was fairly easy to deal with as well. One problem that we encountered was that the lid seemed somewhat hard to open and close because we weren't sure how stable it was going to be or if it would break. Also during the last cycle the LCD screen suddenly went blank but this was most likely do to a faulty power strip.
ProtocolsPolymerase Chain Reaction 1. A Polymerase Chain Reaction(PCR) is used to amplify a single piece of DNA. The steps that lead up to the replication of a DNA sequence are denaturation, annealing, and extension which involve several different cycles ranging in terms of both length of time and temperature. This results in the exponential replication of a certain sequence of DNA.
Patient ID 72537 74083


Research and DevelopmentPolymerase Chain Reaction This experiment heavily employs the use of PCR, or polymerase chain reaction. This process is used to amplify a segment of DNA for in-depth analysis. The process starts by adding a mix of DNA primers and DNA florescent tags that bind to a specific segment of DNA. After these primers bind, DNA polymerase binds to the primer-DNA strand bond and incorporates the florescent tags into the molecule. The polymerase adds base pairs to the base DNA strand and creates an antisense copy of the original strand. Since this process happens for both strands of the DNA, each process essentially duplicates the DNA strand. In order for this reaction to occur, the solution needs to be heated up to allow the double stranded DNA to melt into single strands. Then the solution needs to cool down and let the DNA anneal. Each one of these heating and cooling steps makes up the process of thermal cycling. Each cycle the DNA is replicated once. The sample is run through anywhere between 30 and 50 cycles in order to amplify the DNA and the targeted signal enough to be picked up by an imaging machine. Since the primers only bind to specific segments of DNA, the sample will only light up when the segment of interest is duplicated. In this lab, the SYBER green dye colored the DNA in solution, only fluorescing when bonded to double-stranded DNA. As a result, PCR can be used to identify the presence or absence of a particular DNA sequence. Specific Cancer Marker Detection - The Underlying Technology A single nucleotide polymorphism is a variation in a single DNA nucleotide. The four DNA nucleotides are represented using the letters A, T, C and G. These variations occur normally throughout DNA and represent the most common form of genetic variation among people. They occur at a rate of 1 per every 100 to 300 bases along the 3-billion-base human genome. SNPs are point mutations that have been evolutionarily successful enough to recur in a significant proportion of the population of a species. In other words, SNPs are evolutionarily stable, meaning they do not change much from generation to generation. This allows SNPs to be considered highly conserved within the population and therefore serve as ideal biological markers for genetic research. In order for a sequence variation to be classified as a SNP it must occur in at least 1% of the population. Millions of SNPs have been identified in the human genome and cataloged in accessible databases. SNPs can occur with a gene, which is the coding region of DNA, or in a non-coding region. Because only about 3-5% of DNA actually codes for the production of proteins, most SNPs are found within non-coding regions. Since SNPs can be located near a gene associated with a certain disease, or occasionally within that gene, researchers have been able to pinpoint various diseases on the genome map. SNPs found within a gene, or somewhere in the regulatory region of a gene, are of particular interest because they are more likely to alter the biological function of the gene and therefore, the function of the protein. It is important to remember that SNPs do not cause or identify a disease state directly, but allow for the possible diagnosis or assists in determining the likelihood that someone will develop a particular illness. They also have the ability to help predict an individual’s response to certain drugs, environmental factors, chemicals, toxins, etc. In fact, since SNPs occur most frequently in the non-coding regions of DNA, they do not produce physical changes in people and have no effect on health or development. (Written by Kim L, Posted by William L).
Single Nucleotide Polymorphisms and this Lab In this lab, the PCR diagnostic is testing for the presence of the cancer-linked rs17879961 SNP mutation. Since this SNP creates a new DNA sequence, primers can be made to only bind to the mutated DNA sequence. As a result, the PCR test can be run with a unique primer that will only bind if the target DNA shows rs17879961 mutation, allowing the test to identify the presence or absence this cancer-linked SNP. In rs17879961, there is an anomalous A-T pair that creates a new and unique DNA sequence specific to this cancer-linked mutation. Thus, only a specific primer will bind to the targeted sequence and, as a result, the PCR will only react if the specific mutation is present. rs17879961 Mutation and Primer Sequence The backward primer that attaches to the other strand of DNA has the following sequence:
Bayes' Rule Bayes' Rule is an exceedingly helpful tool when measuring the accuracy of a certain test. for a more in depth summary of Bayes' Rule lets conclude that P(B/A) are users and P(B/A-) are non users. then the new formula for testing the validity of the test would look like P(A/B)=((P(B/A)(P(A))/((P(B/A)(P(A)+(P(B/A-)(P(A-)). for a example reference, if 2.5% of the tested individuals were positive for cancer and the test was 95% accurate the equation would look something like this, and 97.5 individuals were negative, P(user|+)= (P(+|user)(P(user))/(P(+|user)(P(user)+(P(+|non-user)(P(non-user)). Thus with implementing the numbers for the variables the equation would look like ((.95)(.025))/((.95)(.025)+(.05)(.975))=.33=33%. With the information provided out of a study of 100 individuals 48 would be tested as a false positive and 16 would be tested as actual positive, and 36 would be negative. so the percentage of accuracy of the cancer test would be 33% as calculated. Bayes' Rule is exceedingly useful when calculating the false positive as well as the actual accuracy percentage, a needed tool when dealing with test that could potentially save an individual.
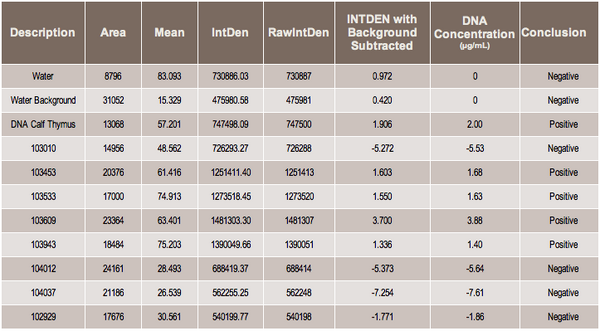
Results 
KEY
|
|