BME103:W930 Group10
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OUR TEAMLAB 1 WRITE-UPInitial Machine Testing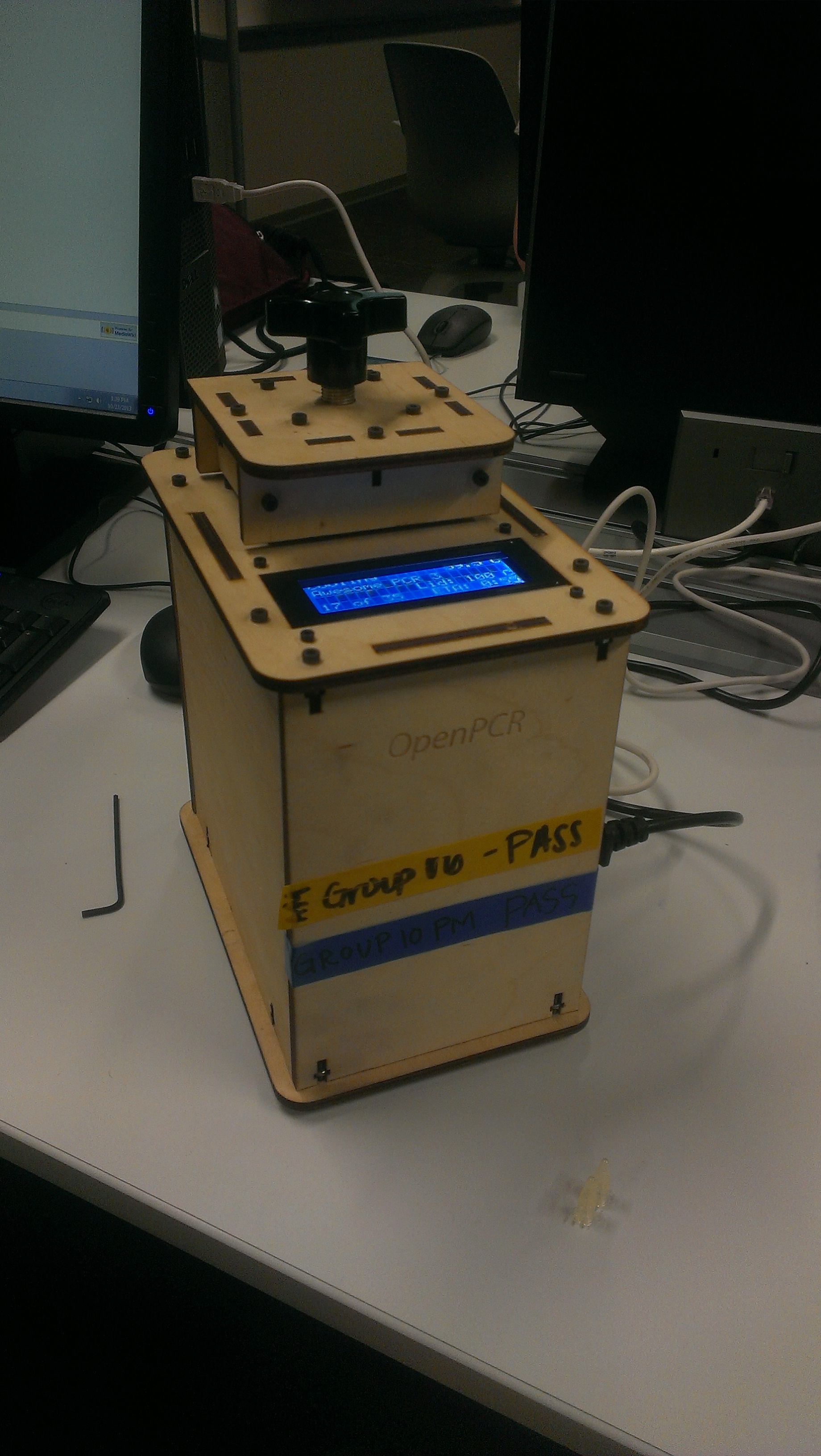 This is the OpenPCR machine utilized to automate polymerase chain reactions. This reaction allows for the amplification of specific DNA which is useful for detecting different markers, such as those indiciating an increased risk for cancer, presence of HIV, etc. Experimenting With the Connections When we unplugged the LCD screen from the Open PCR circuit board, the machine's LCD screen did not turn on. When we unplugged the white wire that connects Open PCR circuit board to the heat sink, there appeared to be no effect, however, it is likely that the heat sink would not function during a trial.
On Oct. 24th, 2012, we first used the Open PCR machine number 10 to run 25 cycles on eight samples which included two sets of three samples and a positive and negative control. The process was successful, taking about 90 minutes for the reaction to complete. Initial testing of the device indicated that the machine and software were synced in regards to the temperature during each cycle.
ProtocolsPolymerase Chain Reaction Polymerase Chain Reaction(PCR) works by using a mix of enzymes that transcribe sections of DNA. The enzyme mix is combined with patient DNA. Then, the sample is heated and cooled in regular cycles to match the ideal temperatures for the different enzymes. This will result in replication of the specific section of DNA that is being tested. Steps to amplify a patient's DNA sample: 1. Add 50 microliters PCR master mix of enzymes to patient DNA sample 2. Put in PCR machine 3. Run for 25 cycles at 95 degrees C for 30 seconds, 57 degrees C for 30 seconds, and 72 degrees C for 30 seconds
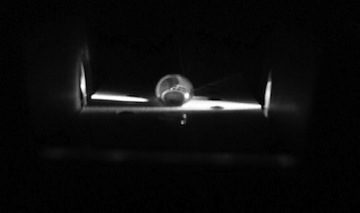
Flourimeter Setup: Camera Setting: The camera had the flash off, ISO at 800, auto white balance, high exposure, high saturation, and low contrast. ImageJ Procedure:
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology The NCBI database allows for genes to be searched through in order to determine different mutations and information about them. In this lab, we looked up CHEK2 as it related to rs17879961, a human gene, in the Short Genetic Variations database. CHEK2 is checkpoint kinase 2 which happens in response to DNA damage. Through the database, it was found that a missense mutation occured in the 22nd chromosome. The primer of the cancer sequence is GGAAGTGGGTCCTAAAAACTCTTACA[C/T]TGCATACATAGAAGATCACAGTGGCand the reverse primer to this would be AACTCTTACACTGCATACAT. The mutation for cancer changes the codon "ATT" to "ACT" in the DNA sequence. This change codes for cancers such as breast and colorectal cancer.
The reason why cancer mutations give a positive PCR signal, while a non-cancer sequence gives no signal, is a result of the primer that attaches to the sequence. The primer is coded for the specific codon "ACT" and will only attach if the sequence is such, which then allows the Taq Polymerase to bind and replicate the DNA exponentially. If the primer sees that the codon is "ATT," it will not bind and therefore will not replicate and cause the PCR signal to be positive. In this lab, the samples that exhibited the fluorescent dye were the ones in which the PCR signal was positive and therefore had cancer.
Baye's Rule is used to determine the probability that a person has cancer or not. In a study of 180 people, 1.1% have the mutation for cancer while 98.9% do not. Using Baye's rule, it was found that 7.8% should have cancer. The formula for Baye's Rule is p (hc|C) = p(C|hc) p(hc) / p(C)
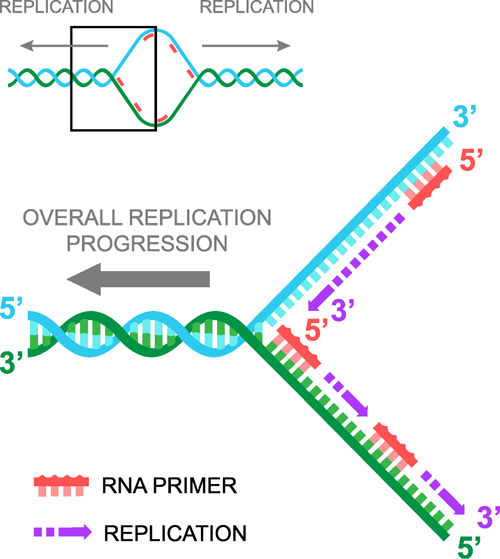 [2] [2]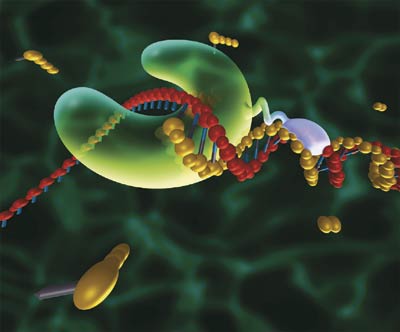 [1] [1]For an animated walkthrough of the process, check out this PCR Virtual Lab from the team at the University of Utah's Genetic Science Learning Center ResultsData Measured via ImageJ
Green Channel With DNA Green Channel With Water
Data Measured via ImageJ
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Works Cited
[1] "Microbial Chatter." - Thermus Aquaticus. N.p., n.d. Web. 31 Oct. 2012. <http://docp.edublogs.org/thermus-aquaticus/>.
[2] "Replication." Shmoop. N.p., n.d. Web. 31 Oct. 2012. <http://www.shmoop.com/dna/dna-replication.html>.









