BME103:T930 Group 6
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||||||||
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingThe Original Design 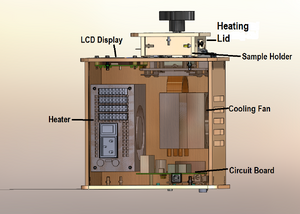 Experimenting With the Connections When we unplugged the LED interface cable from the processing chip, the LED screen went blank. When we unplugged the white wire that connects the processing chip to the heating block, the displayed temperature was no longer correct. Test Run We first tested the OpenPCR around 25 October 2012. The first experience was a simple graphic user interface and extremely easy physical set-up. The only issue was that the directions in the manual were not as clear as they could have been, and the machine ran very slow. Despite the speed, this is an effective machine to amplify and replicate DNA.
ProtocolsPolymerase Chain Reaction Polymerase Chain Reaction (PCR) is a biochemical process the replicates and amplifies a desired sequence of DNA. The original DNA strand is pulled apart and primer marks the location of the targeted DNA sequence for polymerase to begin replication of the complementary strands. This process is continued until there are an exponentially increasing amount of the desired strand, allowing it to be closely examined.The PCR reaction relies on thermal cycling, in which repeated cycles of cooling and heating the samples enable the enzymes to appropriately replicate the targeted strand, and carry out the processes previously described. The Steps: 1. Obtain DNA sample and label them clearly. 2. Using a micro-pipette, transfer the proper amounts of forward primers, reverse primers, dNtP's and Taq Polymerase master mix to each the 8 labeled DNA sample tubes. Be sure to replace the pipette tip after every transfer to avoid contamination. 3. Place the DNA sample tubes into the Open PCR machine. 4. Close lid and tighten screw until it touches the tops of the tubes. 5. Program the machine to run the following cycles: - Stage 1: 1 cycle, 95 degrees Celsius for 3 minutes - Stage 2: 30 cycles, 95 degrees for 30 seconds, 57 degrees for 30 seconds, 72 degrees for 30 second - Stage 3: 72 degrees for 3 minutes - Final Hold: 4 degrees
GoTaq® DNA Polymerase is supplied in 2X Colorless GoTaq® Reaction Buffer (pH 8.5), 400μM dATP, 400μM dGTP, 400μM dCTP, 400μM dTTP and 3mM MgCl2.
Patient Information The eight samples that were tested consisted of a positive and negative control. The other six had three DNA samples from one patient and three DNA samples from the second patient. The first patient has an ID of 51919, and is a female that is fifty-four years of age. The second patient has an ID of 60627, and is also a female, but with an age of sixty-three.
 Fluorometer Assembly Procedure 1. Remove lid from black box and invert the box, open the front flap. 2. Place glass slide into holder. 3. Using pipette, squeeze two drops of SYBR green dye inside the round glass windows on slide. 4. Add DNA sample to dye. 5. Align the sample so the blue LED shines directly though it focusing the light on the other side. 6. Move holder apparatus inside the box so that no light reaches it. 7. Place smartphone with the flash off onto holder and direct camera lens at the slide apparatus. 8. Take photo with smartphone and process on computer using ImageJ.
1. E-mail image to computer 2. On the computer open the image file using ImageJ 3. In ImageJ, under the "analyze" tool bar select "Set Measurements" 4. Make sure the following boxes are checked, and no other boxes other than the ones listed: -Area -Mean Grey Value -Integrated Density 5. From the download folder drag the image file into ImageJ 6. When the image is opened in ImageJ from the top drop down menu select "Image" and then "Color" and then "Split Channels" 7. Close out the red and blue channel, only use the green channel 8. With the Oval tool select only the entire drop of liquid, avoiding the background but encompassing every bit of the droplet 9. After the droplet part of the image is selected, go to "Analyze" then "Measure" or press Ctrl + M 10. Now move the circle to the background of the image by clicking and dragging on the circle, be careful not to change its size 11. Finally, save the results as an excel spreadsheet (set by default)
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology Polymerase chain reaction (PCR) is a biochemical technology that is used to amplify a specific piece of DNA by generating thousands to millions of copies of a particular DNA sequence. The method relies on thermal cycling by heating and cooling the DNA so that the double strands break apart and can be replicated. What are the component of a PCR reaction? Template DNA: A specific DNA sequence that is trying to be detected. Primers: A short segment of DNA sequences that is complementary to the target DNA sequence and binds to the targeted nucleotides. Taq Polymerase: An enzyme that is active at high temperature that begins at the primer and takes complementary nucleotides from the solution and binds them to the "unzipped" strands. Magnesium Chloride: A cofactor that attaches to the Taq Polymerase to affect the speed of the reaction. dNTP's: Deoxynucleotide triphosphates; individual nucleotide bases in that solution that are attached to the replicated DNA strands by Taq Polymerase. What happens during each step of the thermal cycle?
The r17879961 cancer-associated sequence that is being tested for is AAACTCTTACACTGCATACA. The bolded C represents the cancer gene that appears because the C took the place of the normal T base that should be present. The one nucleotide base that was changed results in producing the amino acid of threonine instead of isoleucine. Because of the change in the amino acid being produced, a study in Finland has found that this sequence has been link to susceptibility to colorectal cancer. Of the patients tested, 7.8% of the patients with colorectal cancer had the allele while 5.3% of patients without colorectal cancer had the same allele. Why does a cancer gene produce a positive result while a normal gene produces a negative?
Relation to Bayes' Rule: Bayers' rule is used to determine the probability of true positives, false positives, and false negatives to predict the reliability of the process in detecting the cancer sequence in patients.<br< The equation for Bayes' Rule is P(A|B) = P(B|A)P(A) / P(B), wherein P(A) represents the initial belief in the occurrence of A, P(A|B) represents the degree of belief in A with regards to B, an known factor, and P(B|A) / P(B) represents the support B has for A.
Results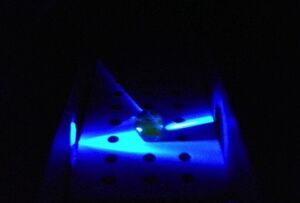 
KEY
| |||||||||||||||||||||||||||||||||||||||||||||||||||




