BME103:T930 Group 5 l2
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||
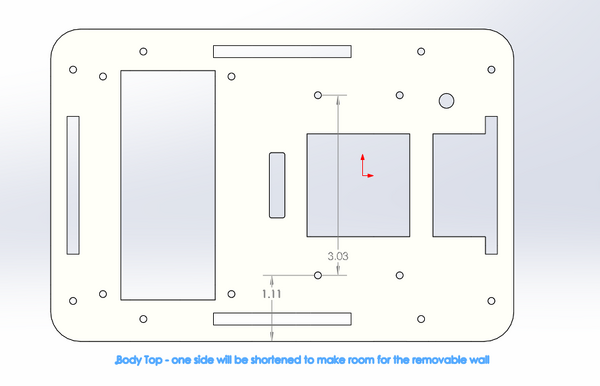
OUR TEAMLAB 2 WRITE-UPThermal Cycler EngineeringOur re-design is based upon the Open PCR system originally designed by Josh Perfetto and Tito Jankowski.
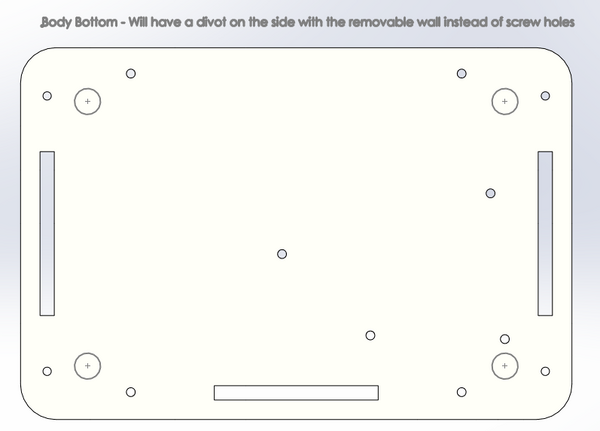
2. Screw the body top onto the body sides that are attached to the body bottom. Make sure the side of the body top that is shortened is above the side of the body bottom with the divot. 3. Then slide the removable wall into the sides of the machine with the divots in it. This wall can then be easily removed and put back in place.
ProtocolsMaterials
PCR Protocol How Polymerase Chain Reaction Works Utilizing DNA polymerase enzyme, PCR synthesizes complementary strands from a targeted segment of DNA. Test tubes within the PCR machine work need ample mixtures of the four DNA bases (Adenine, Thymine, Guanine, and Cytosine) to recreate the desired DNA sequence. The test tubes also need primers to kick-start the DNA replication process. Due to this, people who want to amplify DNA need to know the region of DNA they want to copy.
1. Extract DNA from the patient you are running tests on 2. Place your extracted DNA in a PCR tube (suited for rapid heating and cooling) 3. Remember if you’re using a micro-pipetter, you need to replace the tips before pipetting new chemicals (to avoid cross-contamination) 4. Add primer 1 to the PCR tube – specialized to cut the DNA segment at the front of the desired part 5. Add primer 2 to the PCR tube– specialized to cut the DNA segment at the back of the desired part 6. Add nucleotides (A, T, C, and G) to the PCR tube- to aid in the synthesis of replicated DNA 7. Finally, add DNA polymerase to the PCR tube- the enzyme responsible for DNA replication (or here amplification). 8. Place the PCR tube into a PCR machine or DNA Thermal Cycler – this expedites the DNA replication process. 9. The settings for your PCR machine should be as follows: Cycle 1 A. heat to 95°C to denature the double helix of DNA then B. cool to 55°C to activate primers then C. heat up to 72°C to activate DNA polymerase 10. Cycles repeat to amplify DNA
Instructions for proper set-up of the fluorimeter 1. Take all the parts out of the box and unbutton one of the sides and flip it upside down. 2. Take the stand slide the plastic white slide into it. 3. Take the pipette and place a drop of water onto one of the black circles. Then place another drop on a black dot right next to the previous one. Follow this by placing another water drop in the middle of the two previous drops to combine the agglomeration of water. 4. Turn on the switch to shine the blue light through the middle of the drop. (You may need to adjust the assembly to get the blue light to pass through it.) 5. Now place the assembly under the upside down black box. 6. Take a picture by putting a phone in the phone stand and closing the lid of the black box. (The picture will need to most likely be timed, no flash.)
Samples: Begin with a POSITIVE CONTROL SOLUTION, the calf thymus DNA, and a NEGATIVE CONTROL SOLUTION, water. Follow with three repetitions for each of the DNA samples of your patient(s). Now: 1. Collect samples produced from the PCR machine. 2. Using the micro-pipetter for each individual sample, transferring the 150μL into the larger test tubes containing 400 mL of the (provided) buffer solution. 3. Set up the fluorimeter machinery as instructed above. 4. A “blank" sample of distilled water should be utilized as a negative control. 5. With the provided equipment, add two drops of each PCR sample onto the glass plate, followed by two drops of SYBR green. Be careful when adding the drops ensuring that they are initially spaced out, otherwise they will combine when more reactants are added. 6. Insulate the system as the instructions below say, setting up the camera as well. Set the ISO at 800, remembering to turn off the flash. 7. Now, clear the sample from the glass tray. 8. Repeat the same process with the next sample utilizing a different space on the removable tray. Only three sets of samples should be used per tray. 9. Repeat this process until finished
1. Take a picture. 2. Open Software Package (IJ) (that needs to be downloaded to camera/device) 3. Click “Run”.
*The software will run the performance required, however for your information this is how the application processes the information.* 1. Set measurement to integrate density and linear grey value. 2. Isolate green. 3. Measure INTDEN in drop. 4. Subtract background.
1. Collect the INTDEN for your positive and negative controls and your patient samples. 2. Calculate the DNA μg/mL with the following equation: 2*INTDEN of sample/INTDEN of DNA Calf Thymus Research and DevelopmentBackground on Disease Markers A very large disease that is wide spread all around the world is Alzheimer’s. Nearly one in every 85 people around the world has this disease. On average, one’s life will end seven years after the diagnosis. Fewer than three percent of patients live longer than fourteen years after the determination of the presence of Alzheimer’s. Of the type of Alzheimer’s that is autosomal, the DNA produces a protein that is different than the expected one. This misfolded protein causes the disease. The DNA sequence that codes for the specific amino acids that form these mutated proteins are called SNPs. An Alzheimer’s related SNP is called rs429358. It exists on the 19th chromosome of the human genome. More information about this SNP can be found by looking up the reference number, rs429358, on OMIM. The normal sequence appears TGC while the mutated Alzheimer’s sequence will show CGC. The respective amino acid produced changes from Cysteine to Arginine. This change in amino acid, causes the proteins to fold differently.
The reverse primer for this gene would be: GGCGGCCGCACACGTCCTCCCA. The critical sequence begins at 45,411,950 on the 19th chromosome. Now looking 200 base pairs to the left, a forward primer can be created. In this case it would go as follows: CATCCCAGCCCTTCTCCCGC.
This image displays the location of the DNA in the nucleus and the exponential rate of amplification, 2 raised to the n. 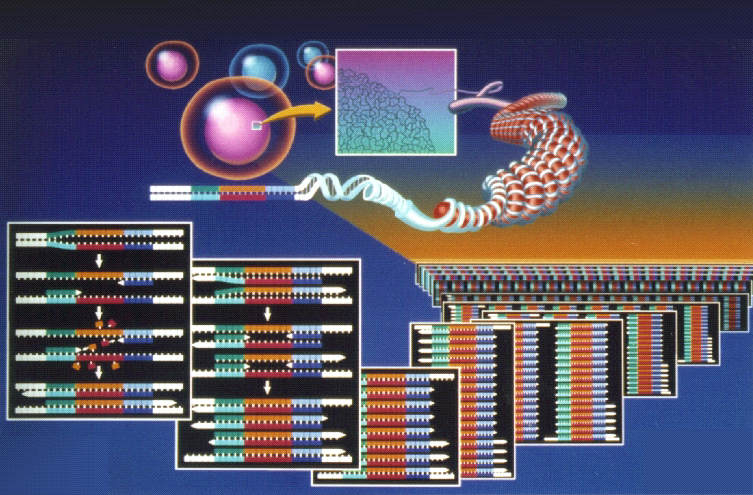
Bayesian Statistics Bayes' Law can calculate the probability that something is true, given a particular set of circumstances. Essentially the probability of A if event B occurred. From a medical perspective, Bayes' Law can be applied to determine the probability of the accuracy of a result. It takes into account the statistics of true positive results, true negative results, false positives, and false negatives. An article published by the American Journal of Epidemiology in 1996, displays the results of a study regarding late-onset Alzheimer’s disease. Below are these results. “A total of 1,454 subjects who had been recruited into the Medical Research Council Elderly Hypertension Trial between 1983 and 1985 completed cognitive tests at entry to the trial (when they were without signs of dementia) and 1 month later. Their dementia status was ascertained in 1990-1991. The test package identified 52% of Alzheimer's disease cases with a 9% false-positive rate or 90% of Alzheimer's disease cases with a 29% false-positive rate” 1
|
|||||||||||||||||||||||||||||