BME103:T930 Group 5
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingExperimenting With the Connections When we unplugged the mounting plate from the Open PCR circuit board, the machine's display shut off and we could no longer see any information. When we unplugged the white wire that connects the Open PCR circuit Board to main heating block the machine was no longer able to be measured accurately.
We first tested open PCR on 25 October 2012. The first experiment went well despite som edifficulty opening the lid and plugging the display in. Our program ran smoothly and we got the data we needed.
ProtocolsPolymerase Chain Reaction How Polymerase Chain Reaction Works Utilizing DNA polymerase enzyme, PCR synthesizes complementary strands from a targeted segment of DNA. Test tubes within the PCR machine work need ample mixtures of the four DNA bases (Adenine, Thymine, Guanine, and Cytosine) to recreate the desired DNA sequence. The test tubes also need primers to kick-start the DNA replication process. Due to this, people who want to amplify DNA need to know the region of DNA they want to copy. How to Amplify a Patient’s DNA Sample 1. Extract DNA from the patient you are running tests on 2. Place your extracted DNA in a PCR tube (suited for rapid heating and cooling) 3. Remember if you’re using a pipeter, you need to replace the tips before pipetting new chemicals (to avoid cross-contamination) 4. Add primer 1 to the PCR tube – specialized to cut the DNA segment at the front of the desired part 5. Add primer 2 to the PCR tube– specialized to cut the DNA segment at the back of the desired part 6. Add nucleotides (A, T, C, and G) to the PCR tube- to aid in the synthesis of replicated DNA 7. Finally, add DNA polymerase to the PCR tube- the enzyme responsible for DNA replication (or here amplification). 8. Place the PCR tube into a PCR machine or DNA Thermal Cycler – this expedites the DNA replication process. 9. The settings for your PCR machine should be as follows: Cycle 1 a. heat to 95°C to denature the double helix of DNA then b. cool to 55°C to activate primers then c. heat up to 72°C to activate DNA polymerase 10. Cycles repeat to amplify DNA
Promega Product Description of PCR master mix Product Contents GoTaq® Colorless Master Mix Cat.# Size M7141 10 reactions M7142 100 reactions M7143 1,000 reactions Includes GoTaq® Colorless Master Mix, 2X, and Nuclease-Free Water. Description: GoTaq® Colorless Master Mix (a,b) is a premixed ready-to-use solution containing a nonrecombinant modified form of Taq DNA polymerase that lacks 5´→3´ exonuclease activity. The master mix contains dNTPs, MgCl2 and reaction buffers at optimal concentrations for efficient amplification of DNA templates by PCR. GoTaq® Colorless Master Mix, 2X: GoTaq® DNA Polymerase is supplied in 2X Colorless GoTaq® Reaction Buffer : (pH 8.5), 400µM dATP, 400µM dGTP, 400µM dCTP, 400µM dTTP and 3mM MgCl2. Storage Conditions: See the Product Information Label for storage recommendations. Minimize the number of freezethaw cycles by storing in working aliquots. Product may be stored at 4°C for up to 6 weeks. Mix well prior to use. Quality Control Assays Functional Assay: GoTaq® Colorless Master Mix is tested for performance in the polymerase chain reaction (PCR). GoTaq® Colorless Master Mix, 1X, is used to amplify a 360bp region of the α-1-antitrypsin gene from 100 molecules of human genomic DNA. The resulting PCR product is visualized on an ethidium bromide-stained agarose gel. Nuclease Assays: No contaminating endonuclease or exonuclease activity detected.
Samples Explained: There were four heterogeneous solutions used in the PCR lab. The first of which was a POSITIVE CONTROL SOLUTION, allowing my lab partners and I to see what DNA with cancer would come up with after PCR and the fluorimeter analysis. The second was a NEGATIVE CONTROL SOLUTION to aid in seeing what a sample without cancer would come up with after PCR and the fluorimeter analysis. These two samples were crucial for comparison to our patient trial solutions. Two patients donated a set of three solutions each (for validity and reliability in testing). Their information follows: Patient 1: Sex=Female, Age=46, Code#=51000 Patient 2: Sex=Male, Age=56, Code#=92369 These were our EXPERIMENTAL SOLUTIONS to draw comparison to the negative and positive control solutions.
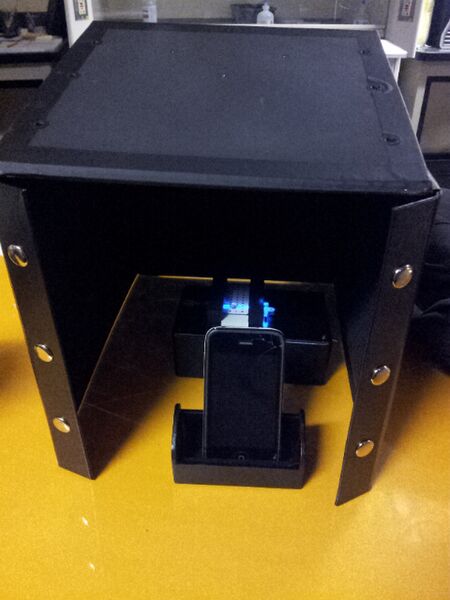
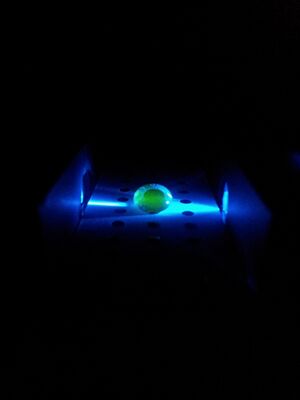
1 - Take all the parts out of the box and unbutton one of the sides and flip it upside down. 2 - Take the stand slide the plastic white slide into it. 3 - Take the pipet and drop a drop of water onto one of the black circles. Then place another drop on a black dot right next to the previous one. Follow this by placing another water drop in the middle of the two previous drops to combine the agglomeration of water. 4 - Turn on the switch to shine the blue light through the middle of the drop. (You may need to adjust the assembly to get the blue light to pass through it.) 5 - Now place the assembly under the upside down black box. 6 - Take a picture by putting a phone in the phone stand and closing the lid of the black box. (The picture will need to most likely be timed, no flash.) From an Image to an Image J 1 - Take a picture. 2 - Download file to computer. 3 - Right click file and open with Image J. WITHIN IMAGE J: 4 - Set measurement to integrate density and linear grey value. 5 - Isolate green. 6 - Measure INTDEN in drop. 7 - Subtract background.
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology The function of this Polymerase Chain Reaction experiment is to determine whether or not there is cancer in a patient. In order to fully explain the processes of determining a prognosis, a basic understanding behind the science is necessary. Deoxyribonucleic acid acts as the blueprint for who we are. It consists of two anti-parallel strands of nucleotides that form a double-helical structure. A nucleotide consists of two bases that form a pair. There are four different types of bases: Guanine (G), Cytosine (C), Adenine (A), and Thymine (T). When base pairs are formed Adenine bonds with Thymine and Guanine forms with Cytosine. A set of three base pairs creates an amino acid. The twenty three amino acids form in different ways to create different proteins. When a base pair changes the corresponding amino acid will change. In the case of colorectal cancer a Cytosine change to a Thymine. This would create an Threonine acid instead of Isoleucine acid. This would completely change the structure of the protein that was formed, causing cancer. Therefore cancer in the human body can be traced back to a certain sequence of DNA nucleotides. The sequence of DNA that is critical to determining the presence of cancer is called the template DNA. When a strand of DNA is copied, it is split into two single strands. While separated, new nucleotides bond with the exposed side of the strand. However, a primer is required in order begin the replication. Each different type of primer is specific to a certain strand of DNA. A protein called Taq Polymerase catalyzes the separation and bonding of the nucleotides to the DNA strand. The binding nucleotides are called dNTPS. Deoxy Nucleotide TriPhosphates are free unpaired nucleotide bases that bind to exposed DNA the separated strand of DNA. The main function of the PCR machine is to replicate the DNA sample that is placed inside of it. This machine uses three steps to complete the replication process. First, it heats up the samples up to 95 degrees Celsius. This process splits open the DNA strand. When the strands are separated, the nucleotide bases are exposed. Next, the machine then allows the samples to cool down to 57 degrees Celsius. This allows the primers bind to the specific bonding site. Once the primers have bonded, the machine heats the samples up again. Finally, the machine heats it up until it reaches 75 degrees Celsius. The DNA strands are then extended and a new DNA strand is formed. The r17879961 cancer-associated sequence will produce a DNA signal because the reverse primer used, AACTCTTACACTCGATACAT will only attach if the DNA has the same coding with the cancer-associated sequence “ACT”. A positive result will be evident by the large quantity of replicated DNA. This is because the specified primer will find and bind onto the sequence of nucleotides that signal for the cancerous genes. Once they bind the process will continue and they will be replicated over and over. However, a negative result also has some unique characteristics. The process, when applied to a negative sample, will yield very few strands of DNA as it does not have a beginning place to attach because the sequence is AACTCTTACACTTCGATACAT. The sequence that is needed for the primer to bind is ACTC or in reverse CTCA. Therefore the dNTPs will not be able to bind to the exposed nucleotides and a new DNA strand cannot be formed.
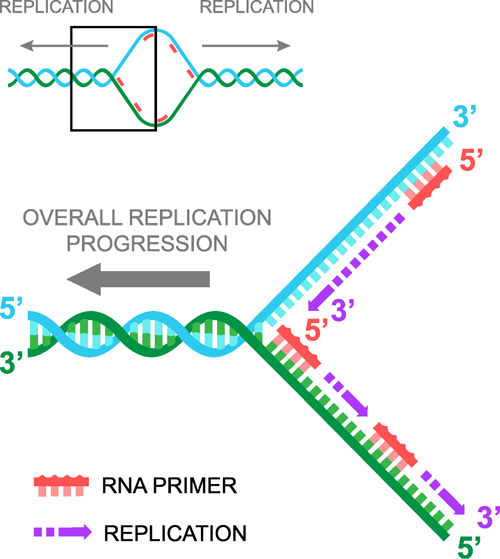
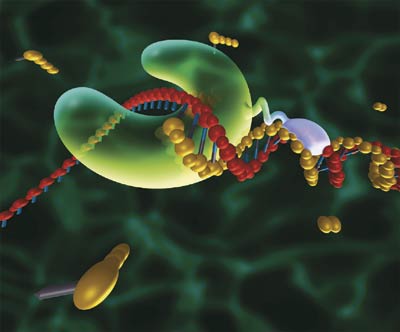
A very helpful, interactive explanation of the process can be found on the PCR Virtual Lab website.
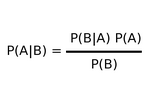
Baye's Rule
Results
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||