BME103:T930 Group 4 l2
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||
OUR TEAMLAB 2 WRITE-UPThermal Cycler EngineeringOur re-design is based upon the Open PCR system originally designed by Josh Perfetto and Tito Jankowski.
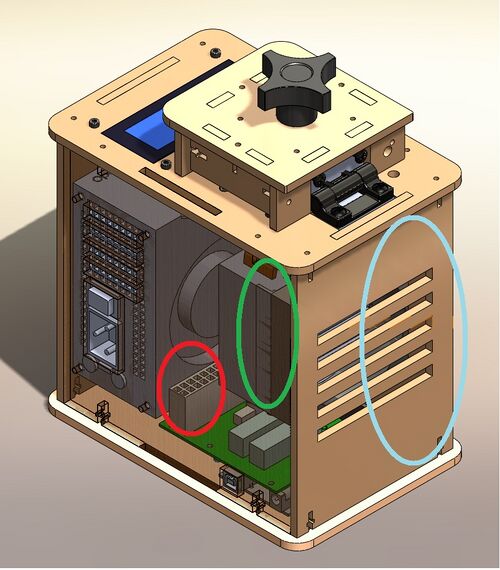
The image above depicts the original PCR design we used as the basis for our new design, with a few minor highlighted changes. In red, we have addressed the wells, which we intend to add another row too. We would also redesign the well plate and its encasing to allow another row and column of wells. Two tiers of wells could also be another design for an improved well plate. In the blue, we highlighted our interest to change what the machine's external casing was made of, which could easily be a better insulated material. Our final change is addressed by the green, which highlights the internal heating mechanism, which should be made of more conductive material to maximize heating capabilities.
The key two features we decided to address were simple, achievable changes that we thought would improve the system of PCR the most. The first feature we thought was necessary was to add another row of wells to the machine. We all agreed that another row of wells would be a very simple addition to the machine, and it would also maximize the number of reactions we were able to perform per session. Also, we figured that a single row of wells would not be excessive to make the heating process less effective. The function of the wells is simply to hold the samples during the reaction process. However, by adding the wells, we do not improve their function, but it does help us achieve our second major goal of our redesign of the PCR machine. The second things we decided we wanted to improve upon was making the machine "harder, better, faster, stronger." In order to do that, we came up with a few minor changes to help us achieve this goal. We would like to make the temperature changes faster, and this is easily achievable, assuming we know how rapid temperature changes effect DNA. We would also achieve this goal by using the most effective insulating material as possible for the outside of the machine, and the most conductive materials for the inside. Perhaps the greatest change in design we could make would be to find primers and polymerases that can be activated at less extreme temperatures so that there would be no need for extreme heating and cooling. We also thought to address a few problems with the flourimeter set up. First, we decided that the measurement process needed to be more uniform, so we thought something easy like adding Velcro to the phone stand and flourimeter would help keep everything uniform because distances would always be constant. However, since it is just Velcro, it is just as easily movable and accessible. We also decided to include an ImageJ app for iPhones, Androids, and other smart phones which could be downloaded, included with settings such as no flash and a ten second timer to allow for complete closing of the lid to keep the light out of the image. This app would allow direct uploading to the software used on the computer. With the way technology is moving, this seems to be a very simple but effective way to improve flourimeter measurements.
Instructions Heating Well: 1. Replace the well plate and well lid. 2. Add another row (or column) of wells. OR 2. Incorporate two tier of wells. 3. Increase the well plate size.
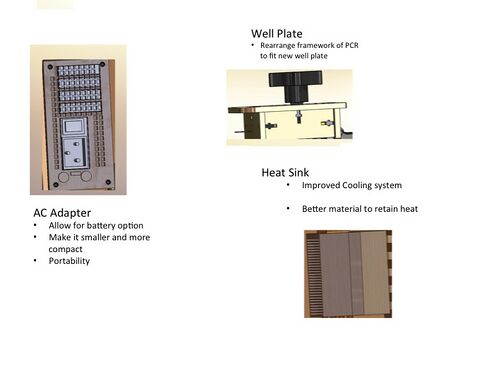
1. Remove the AC adapter. 2. Replace it with a rechargeable battery that allows the PCR's to be portable. The rechargeable battery will allow the computer to alternative between AC and DC. 3. Redesign the adapter to allow it to still fit into the outer shell.
1. Replace the fan with a new fan to maintain certain temperatures. 2. Add titanium to the cooling system would to greatly improve the heat retention of the PCR. This would lead to faster cycles and faster results.
ProtocolsMaterials
Research and DevelopmentBackground on Disease Markers Much like the cancer-linked genes we observed in our first lab write up, we were able to pinpoint several other diseases that can be linked to genetic variation. The first of these to be discussed will be Alzheimer's disease. Alzheimer's is a degenerative brain disease that results primary in memory loss, as well as loss of other cognitive brain functions. The Reference SNP number for this genetic variation is 63750082, and similar to the mutation that causes cancer because it is a missense mutation. However, rather than a single-nucleotide missense mutation, there are a few different mutations that can occur. Basically, in the sequence GGT, the second G can become an A, C, or T (GAT, GCT, GTT). For more information about the gene alteration, visit here! Another genetic variation we chose to study was muscular dystrophy (MD), which can be identified by the Reference SNP number 5030730. Much like the other two studied, it is the result of a change to the sequence "CGA" to "TGA." Muscular dystrophy is a disease where muscle tissue is weak and often degenerates. Those who suffer from the disease likely do not have normal muscle function, and often need a wheelchair or other mobile device to travel. In some cases, mental retardation may also occur. More information about the genetic variation behind muscular dystrophy can be be found here.
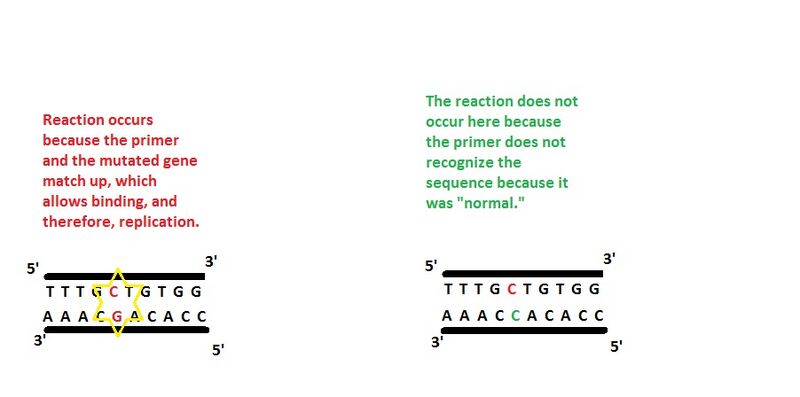
Primer Design The forward primers for Alzheimer's and muscular dystrophy are 5' TTTGCTGTGG 3' AND 5' ATGTCGCATC 3' respectively. Similarly, the reverse primers are the complements of each sequence, because when DNA is paired in strands, each C (cytosine) is paired with a G (guanine), and each A (adenine)is paired with a T (thymine). Therefore, the reverse primers for Alzheimer's and MD respectively are 3' AAACGACACC 5' AND 3' TACAGCGTAG 5'.The way PCR (polymerase chain reactions) are designed is so that they replicate DNA when they come across a certain sequence. Therefore, it is very beneficial to those trying to discover if someone caries the mutated gene sequence for a disease like Alzheimer's or MD. For instance, if the gene were discovered in a person, they could likely expect an early onset of Alzheimer's and prepare for such. Similarly, if a child was found to have the mutated gene for MD, they could also prepare to keep the atrophy at bay for as long as possible. As far as detection goes, PCR primers are specialized so that they have the mutated gene on them. By having the mutated gene as the primer, it will only bind to DNA that contains the mutated gene. Therefore, only those who have the gene for the disease will have DNA replication occur during PCR operation.
Illustration
| |||||||||||||||||||||||||||||||