BME103:T930 Group 17 l2
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||
OUR TEAM
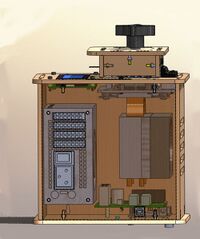
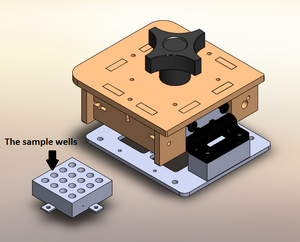
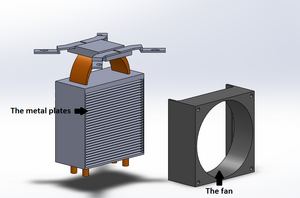
LAB 2 WRITE-UPThermal Cycler EngineeringOur re-design is based upon the Open PCR system originally designed by Josh Perfetto and Tito Jankowski. This is the original Open PCR Machine The originally designed Open PCR Machine can rapidly duplicate DNA, or in other terms amplify it, as well as attach marker to make traits such as cancer visible. PCR stands for polymerase chain reaction. It works by heating up samples to first denature DNA and create single stranded DNA. Then it cools to allow the primer to attach and replicate the DNA. The open PCR machine starts with an initialization step where the temperature rapidly increases to 95 degrees Celsius to create a hot-start for DNA polymerization that requires heat activation. The second step is to denature the protein, where the first cycling event heats the DNA strands at a temperature of 95 degrees Celsius for 30 seconds to melt the DNA template through the disruption of hydrogen bonding between paired bases, effectively splitting the double stranded helix into two single strands of DNA. The third step is the annealing step where the temperature is rapidly lowered to around 50 degrees Celsius to allow for the annealing of primers. The polymerase then binds to the hybrid of primers with the template to begin DNA formation. Then begins the elongation step, which differs depending on the polymerase used; typically the optimum temperature is around 75 degrees Celsius. During the elongation process, DNA polymerase synthesizes a complementary new strand of anti-parallel DNA. The amount of time required for elongation differs depending on the DNA polymerase used as well as the length of the amplified DNA fragments being used. On average, DNA polymerase amplifies at a rate of one thousand bases per minute. Next is the final elongation step where the temperature is held around 75 degrees Celsius to ensure that the DNA strand is fully elongated and will generally hold for around five minutes. Finally there is an end hold temperature that keeps the reaction at a steady temperature (between four and fifteen degrees) until the amplified DNA is ready to be utilized and further studied. The Open PCR machine has several pros and cons. In hopes to improve efficacy and speed, our lab group has suggested several ways that the Open PCR machine can be modified to speed up the process, save energy, and provide extra insulation to prevent heat loss during the initial heating period. In addition, we also increased the size of the sample wells so that insulation, which in turn increases efficacy by distributing the heat applied to the samples in a way that requires less heat. Also, the heat sink and fan are among the least efficient components of the Open PCR machine as a whole. Thus, we intend to update both our fan and heat sink. The cooling components in between heating cycles take up the most time and doesn't cool evenly. With an updated and more advanced fan will hopefully shorten the time needed for the Open PCR system to operate. Finally, the heat sink is smaller than it needs to be. By increasing the number of plates in the heat sink and decreasing the size of each plate, there will be smaller gaps between plates, allowing the heat projected to dramatically increase. With the addition of a more efficient fan, the additional heat shouldn't cause any problems, and the summation of the changes should cut out a percentage of time required for the Open PCR machine to run. This is the Sample Holder and Heated Lid In this image the top has been moved aside so that the sample holder is visible separately from the heated lid itself. The sample holder keeps the samples in place while the PCR machine cycles. The heated lid tightens so that the inside of the lid will press against the samples without crushing them to ensure that they are heated quickly and evenly. To improve this part and increase the heating rate we would widen the sample wells slightly so that insulation would be able to be placed inside them, then the whole sample block would be able to be heated without melting the plastic. This is an improvement over the current design due to the fact that it currently only heats through the lid. Heating solely through the lid is inefficient because the heated lid only comes in contact with the caps of the samples, not only is this the thickest point of the sample container, but it is also the furthest from the liquid contained in the bottom of the sample containers. By adding insulation to the sample wells we increase the heating capacity of the samples themselves as well as improving efficiency, if heat is distributed more evenly, less heat is required to allow the samples to reach the optimal temperature for each step. In addition, we wanted to add more insulation to the lid surrounding the sample holder to keep in the heat to reduce the amount of energy required to heat the samples to the correct temperature. This is the Heat Sink and Fan In this image the heat sink and fan have been isolated from the machine. The fan is used to keep the machine cool and running, whereas the heat sink is used to maintain the temperature of the samples during the PCR reactions. During the cooling cycles, between each temperature change, the fan will increase speed to gradually decrease the temperature of the sample holder and as a result the samples themselves. Our goal was to decrease the temperature by using a more advanced heat sink and more powerful fan. The number of plates in the heat sink is abysmally small, on top of this the fan does not move air very efficiently. If the plates were both slightly smaller and the number increased with a slightly smaller gap in between the plates the amount of heat being projected into the air between the plates would be dramatically increased. With this increase of ambient heat in the machine a more powerful fan would allow for the heat to be more quickly expelled. Therefore, investing more money in a more powerful fan and a better heat sink would allow the machine to cool much more quickly and shorten the time required to run each cycle.
ProtocolsMaterials
1. Gather all components for PCR reaction (all located in the Pre-mixed solution provided).
4. Download the Open PCR software onto the computer from online access.
9. Start the new program.
This is because this "Base Stacking Tm" Calculation-- which calculates the temperature that a primer works best in-- is 65 °C ( see website: http://www.promega.com/techserv/tools/biomath/calc11.htm#melt_results and select PCR Master Mix for the salt concentration adjustment; leave the primer concentration as standard; select "lambda gt11 forward" to see all calculations).
Research and DevelopmentBackground on Disease Markers Alzheimer’s disease is the slow deterioration of the brain. There is no known cure for this disease and it eventually results in death. This disease usually begins with the inability to remember things that have recently happened and in the late stages patients will have much difficulty in remembering basic cognitive functions. The specific missense mutation that I am examining changes a Thymine to a Guanine on the ninth chromosome. This mutation has a 3.8 times increased risk for an early onset of Alzheimer’s. The data reference number is Rs908832. http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=908832 Cystic Fibrosis is caused by a mutation of the protein cystic fibrosis transmembrane conductance regulator. This protein regulates the movement of chloride and sodium ions. Symptoms include slow growth, accumulation of mucus, chest infections, coughing, and shortness of breath. The average patient will be able to live 37 years. The specific mutation I am looking at is a deletion of three nucleotides on the seventh chromosome and causes the deletion of phenylalanine from the polypeptide. In 1979 about 70% of all cystic fibrosis patients carried this mutation. The data reference number is rs113993960. http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=113993960
When testing the Alzheimer’s mutation the forward primer will be TGGCTCCACCACCTCGTGCCC and the reverse primer will be TTTGTGGGGCACGAGGTGGTG. When testing for the cystic fibrosis mutation the forward primer will be TCTTTTATAGTAACCACAAA and the reverse primer will be AACACCAAAGATATTTTCTT.
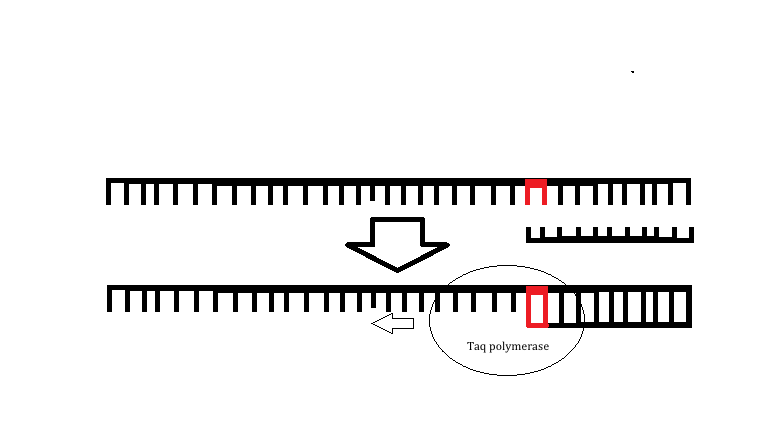
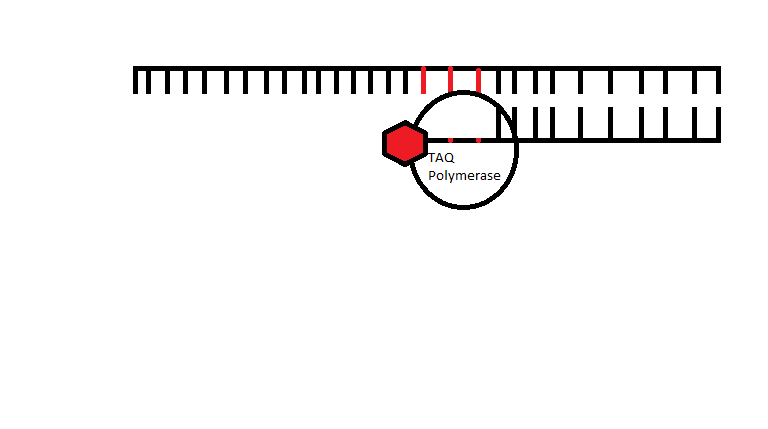
Illustration The following illustration is an example of how the primers for Alzheimer's disease would be able to bind to the template strand. Then the TAQ Polymerase would be able to amplify the DNA and result in a positive PCR reaction. The red shows the location of the SNP. The following illustration shows how a primer would not be able to bind to the mutation resulting in Cystic Fibrosis as it still contains the three nucleotides that would be deleted if their was a mutation present. Since the primer is designed for the mutated DNA the primer is not able to bond to the normal DNA, which results in zero amplification and a negative PCR result. The red parts show which three DNA nucleotides should be deleted due to the SNP. | |||||||||||||||||||||||||||||||||||||||||||||