BME103:T930 Group 16 l2
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||
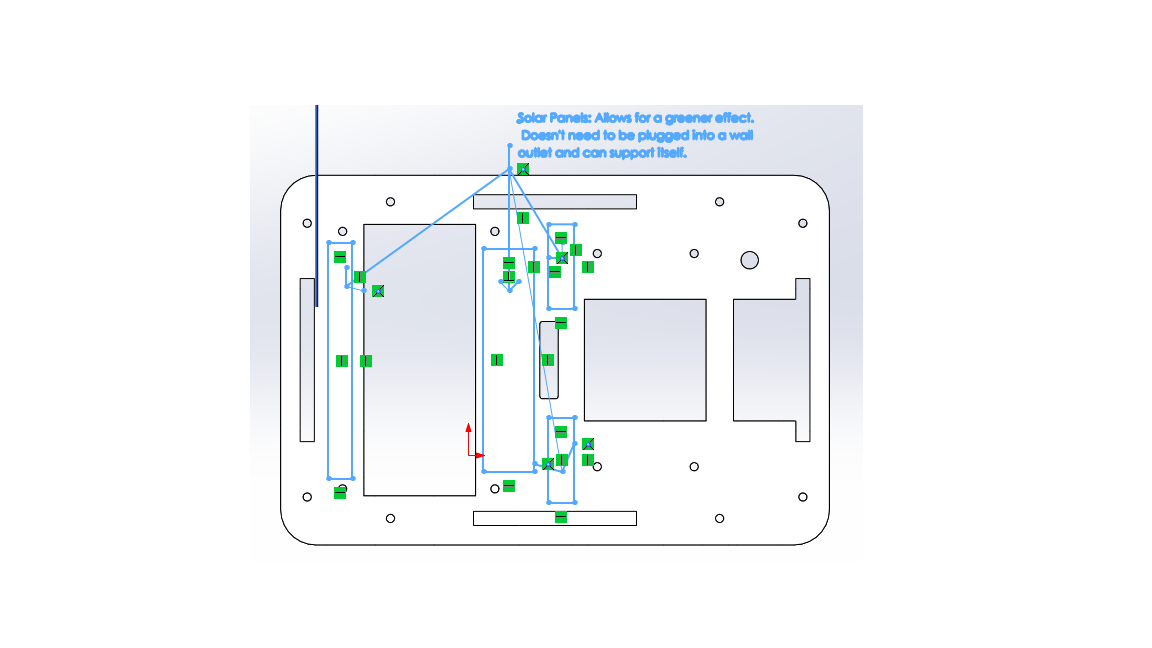
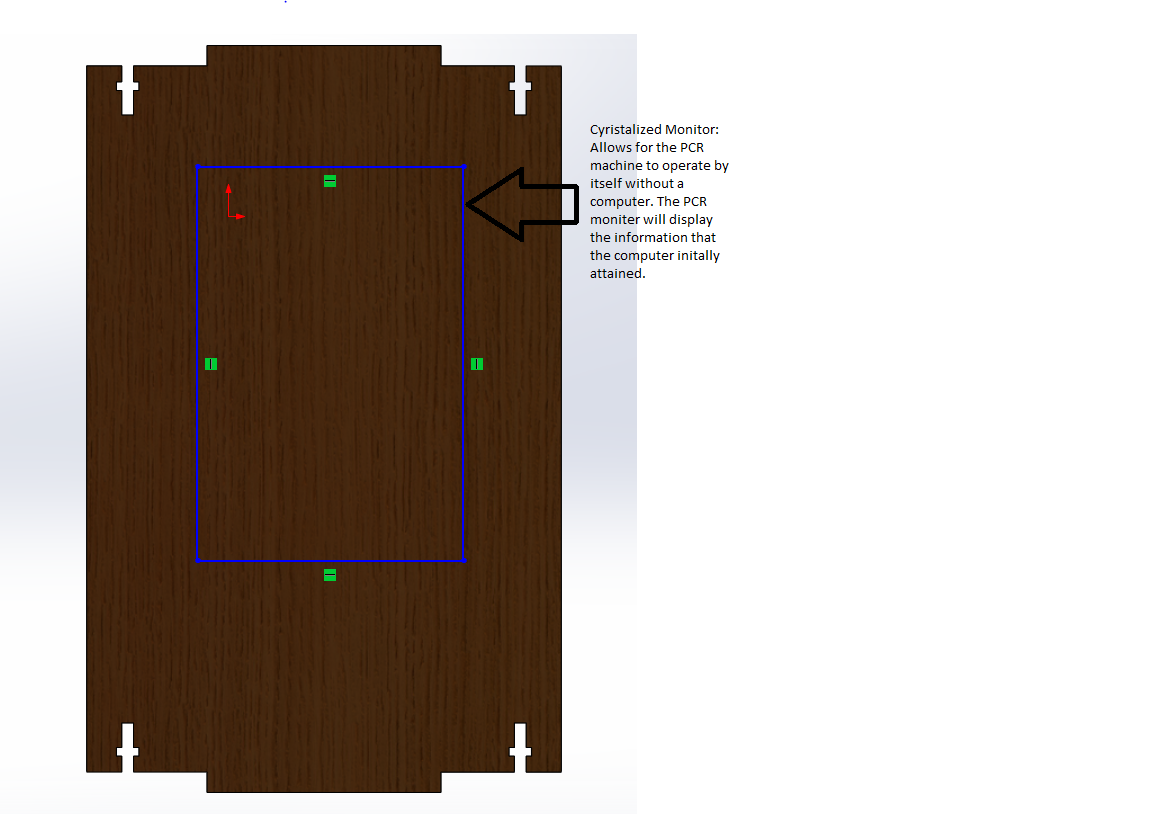
OUR TEAMDear Dr. Haynes, In this group, we all have done equal work on the wiki, but because it took longer for some people to get approval to edit the wiki, and because we worked on each other's computers while still being logged onto each other's accounts, not everyone has an equal amount of entries. However, we all did the same amount of work. From, Rachel, Swetha, Marianna, Omar, Mary, and Muawiya LAB 2 WRITE-UPThermal Cycler EngineeringOur re-design is based upon the Open PCR system originally designed by Josh Perfetto and Tito Jankowski.
ProtocolsMaterials
PCR Protocol
2. To prepare the DNA samples for the PCR, label the DNA test tubes with the patients name and replication number or label it in some way so that you will be able to distinguish between the patient sample and which replication it is (for example P1 R1 for patient one replication one). 3.Put a corresponding label on the tubes containing the reaction mix that will be placed into the actual PCR. After you have labeled all of your patient samples,tubes containing the reaction mix and your positive and negative control, you are ready to add the samples to the PCR reaction mix. The reaction mix includes: Taq DNA polymerase, forward primer, reverse primer, MgCl2 and dNTP's. 4. Each DNA sample should be added to 100 μL of premixed reagent. Using a different transfer pipette each time, transfer your DNA samples, and positive and negative controls to separate tubes of the reaction mix. 5. The samples are now prepared for the PCR machine. 6. Open the PCR, place sample tubes and controls in the machine. 7. Set the Thermal Cycler Program. To set the cycle use the touch screen located on the side of the PCR. 8. After the cycle is set, press start for the PCR to begin the replication process.
DNA Measurement Protocol 1. When the PCR has finished replicating the DNA samples, remove the tubes from the machine. 2. Label remaining pipettes and tubes to correlate with the samples, and label another tube and pipette for the provided Calf-Thymus (which acts as another positive control.) One pipette should be set aside for the transfer of SYBR Green and another should be labeled for waste. 3. Once everything is labeled, transfer the DNA samples using the specifically labeled pipette (to avoid cross contamination do not use a pipette that has touched a DNA sample to transfer a different sample). The DNA should be released into an eppendorf tube containing 400 mL of buffer. Repeat this procedure for all samples and controls including the Calf Thymus. 4. On the rough side of the provided glass slides place two drops of SYBR Green solution with the SYBR green marked pipette over two of the holes in the slide. 5. Place two drops of one of your sample (using that samples specifically labeled pipette) on top of the SYBR green. 6. Carefully move the slide in place so that the light is on it. 7. Place the slide and phone under the black box and take a picture. 8. Using the pipette marked for 'waste,' remove the solution from the slide. 9. Repeat steps four through 8 for all DNA samples and controls including water. To more thoroughly analyze the results upload the images onto imageJ software program. Generally the sample that glow are positive. Research and DevelopmentBackground on Disease Markers Group 16 decided to investigate cystic fibrosis a disease that is caused by mutations in a specific gene on the seventh chromosome, and is then passed down through families as a recessive trait. In this disease, mucus accumulates inside the lungs, digestive tract, and other cavities in the body.[1] This life-threatening disease is the most common chronic lung disease to affect children and is often diagnosable by the age of two. However, weaker strains can go undetected until early adulthood. The effects of this disease, however, have been mediated through the use of treatments that can postpone some of the changes that occur in the lungs. Half of the patients with cystic fibrosis live past 28 years, and patients who only have mucus buildup in the digestive tract are even better off.[3] Gene therapy holds great promise for treating cystic fibrosis. The marker associated with cystic fibrosis is a two nucleotide deletion and has identity rs200007348.[4] This two nucleotide deletion is due to an adenine-guanine swap that occurs in the codon TGG. A codon is a genetic code involved in the RNA process that impacts protein translation. When the TGG undergoes the adenine guanine swap, it becomes TGA which codes for the stop codon, ending protein translation.[2]This SNP heightens susceptibility to cystic fibrosis, a disease with the frequency 1 out of 2000 in Europe and 1 out of 3500 in the United States
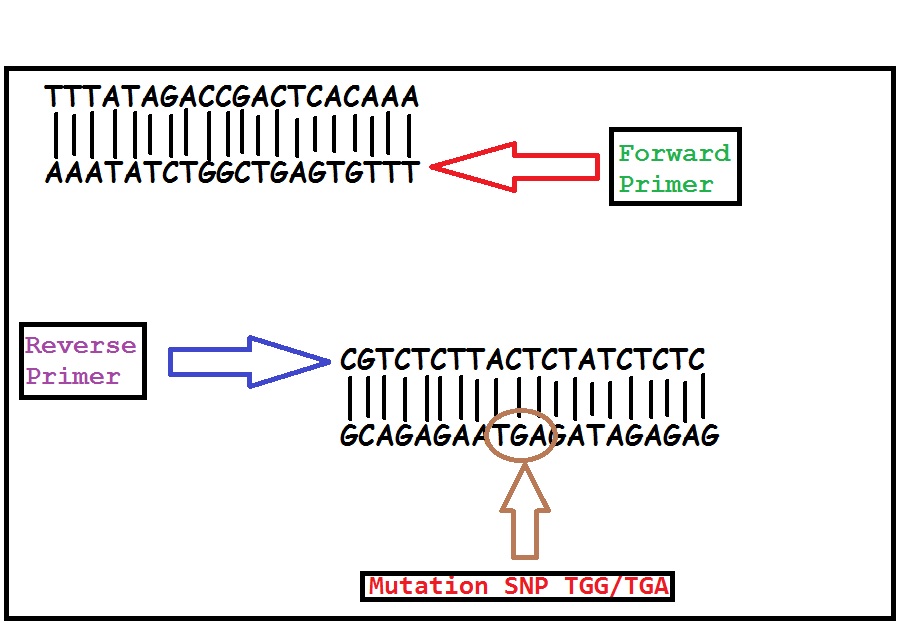
Primer Design Reverse primer: 3' CGTCTCTTACTCTATCTCTC 5' Forward primer: 5' AAATATCTGGCTGAGTGTTT 3' Cystic fibrosis is caused by a 3 bp deletion that leads to a protein which lacks a critical phenylalanine amino acid in the protein. PCR primers have been developed that can distinguish a normal gene from a mutant gene. With these primers a 154 bp product is produced from a normal individual and a 151 bp product is amplified from DNA of an individual with the disease. The disease allele is complementary which will result in a positive result in Open PCR. However, a regular allele cannot give a positive result because the lost nucleotides will be added back into the primer causing a frameshift mutation of three. Thus, these primers are built 151 bp apart and this shortens the temperature cycles from 30 seconds to 10 seconds.
Illustration
| |||||||||||||||||||||||||||||||||||||