BME103:T930 Group 10
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
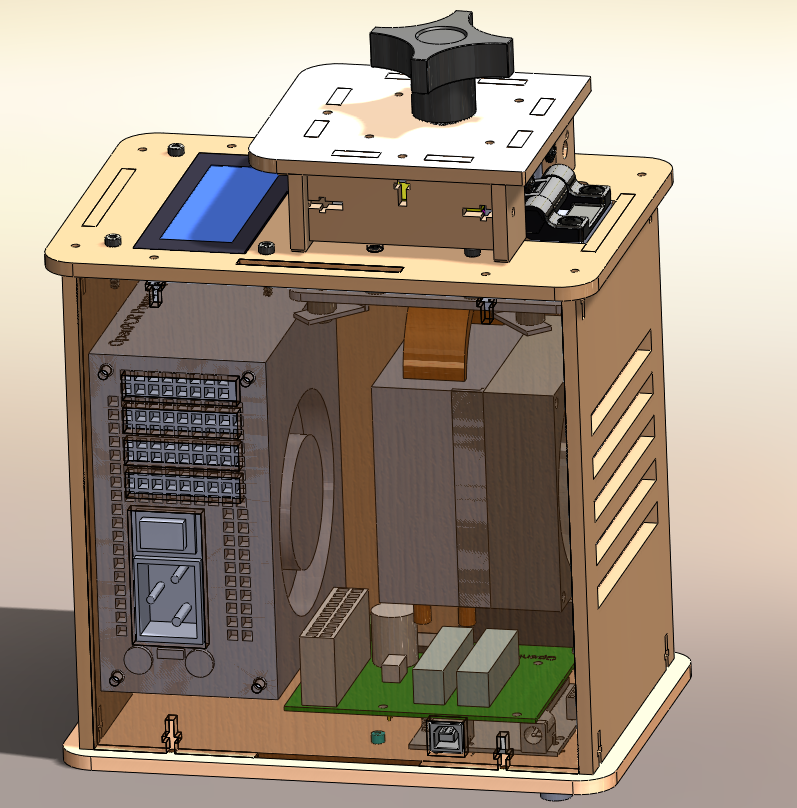
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingA PCR machine is a device that replicates DNA segments by performing polymerase chain reactions. This is useful for purposes such as ours of cancer marker detection. Our PCR machine is a relatively simple and inexpensive machine that is portable and inexpensive. It can hold 16 DNA samples and is compatible with most computers and operating systems. As indicated by the cross section, our PCR machine is composed of few components and is designed for ease of use.
When we unplug the circuit board from the mounting plate, the LED display on the PCR machine stopped functioning. When we unplugged the white wire that connects the Open PCR circuit board to the heating plate, the temperature recordings that were displayed on the LED display stopped functioning.
We first tested our PCR machine on October 23, 2012. Before testing, we attempted to take the PCR machine apart to gain a better understanding of how it works. Following the instructions, we examined individual components of the machine and unplugged special components to examine the effect. After unplugging the wire that connects the circuit board to the LED display, we were unable to plug the wire back in as some of the prongs to the metal adapter had bent. Thus, upon our initial test, we were able to test the machine and observe its changes through the computer display but we could not compare the changes to the LED Display on the PCR machine.
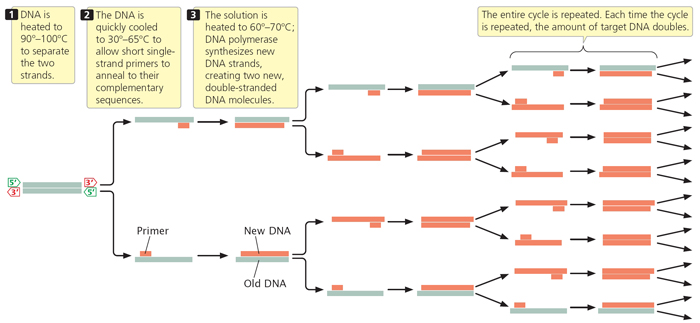
ProtocolsPolymerase Chain Reaction Polymerase Chain Reaction (PCR) is a technique used to amplify fragments of DNA. This allows researchers to see the base sequence of the DNA. It works by the DNA polymerase enzyme synthesizes a complementary strand of the fragmented DNA when mixed with primers that signal where the DNA sequencing should begin. When the DNA polymerase enzyme, MgCL2, dNTP’s, forward primer, and reverse primers are all added to the test tubes and placed in the PCR machine, the mixture is first heated to separate the double helix, then cooled to allow the primers to bind. After the primers bind, the polymerase completes the new complementary strands. The PCR machine then repeats heating and cooling cycles to multiply the fragmented DNA. After a couple hours, the now amplified segments of DNA can be analyzed to test for a cancer marker. Procedure: The PCR (GoTaq) Master Mix is advertised as a “ready-to-use solution” and it contains the Taq DNA polymerase, dNTPs, MgCl2 and reaction buffers. These substances are mixed at proper concentrations so the user can achieve a useable amplification of DNA segments by PCR.
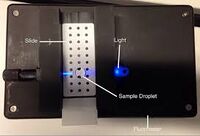
Fluorimeter Measurements Fluorimeter set up Procedure:
Image J Procedure: Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology The DNA sequence r17879961 is the cancer-associated sequence for Colon Rectal Cancer. The goal of this experiment was to detect a cancer-associated sequence in a PCR machine and through the detection method. PCR stands for polymerase chain reaction, a method that uses DNA polymerase and primers, small sets of DNA, to amplify a sample of DNA to study and see specific sequences in it. Through thermal cycling, DNA sequences are melted apart from their complimentary base pairs and then cooled to allow primers to connect to the open DNA sequences. This process is repeated again and again to get many copies of the specific DNA strand. In this experiment, the reverse primers used will be in the sequence of AAACTCTTACACTGCATACA and that will accompany to the TTTGAGAATGTGACGTATGT which is the sequence of Colon rectal cancer that is being studied. This occurs on the 22nd chromosome and the misspent of the disease comes when the sequence AATGT has the T in the middle is changed to a C which is cancer associated and creates a Protein Change to occur. In this experiment, this sequence will be put in with the reverse primer sequence listed above. Through the PCR process, the primer will attach to the sequence and replicate the DNA sequence. If the cancer sequence is present, the primers will attach and replicate until there are numerous samples of the cancer sequence. If there is no cancer sequence present, the primers will not bond since the sequence will be different and the ending DNA sequence will be the same as the original sequence put in. This will provide a correct detection for the r17879961 SNP. To test the reliability of the PCR machine, the experiment will employ Baye's Rules for probability and his equation. The equation is P(A/B)= (P(B/A)P(A))/P(B). To test the efficiency of this test, the experiment will use PPV (Positive Predictive Value) and the NPV (Negative Predictive Value) to determine if someone who tests positive has the disease and if someone who tests negative does not have the disease. These two values can be found using PPV=TP/(TP+FP) and NPV=TN/(TN+FN) with TP being true positive, FP being false positive, TN being true negative, and FN being false negative. The non-mutated form of the gene is found in about 98.9% of the population while the mutated form is found in 1.1% of the population. Furthermore, the mutated gene is found in 7.8% of patients with colorectal cancer and 5.3% of the rest of the population without colorectal cancer. These will help to determine the quality and accuracy of the aforementioned tests. This process above highlights the PCR technique and shows how scientists are able to figure out the sequence for a specific disease or cancer. This process is important because it can produce several strands of the DNA that allow for research. Pray, Â Leslie A.. "The Biotechnology Revolution: PCR and Cloning Expressed Genes | Learn Science at Scitable." Nature Publishing Group : science journals, jobs, and information. N.p., n.d. Web. 13 Nov. 2012. <http://www.nature.com/scitable/topicpage/the-biotechnology-revolution-pcr-and-the-use-553>.
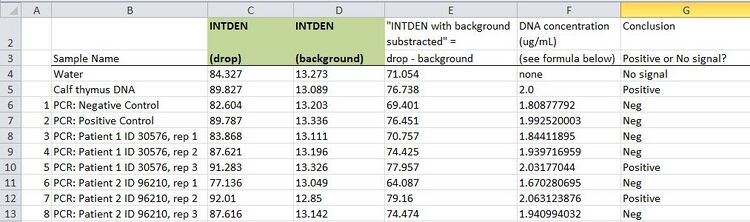
Results
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||