BME103:T130 Group 17
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||||||||
GROUP17LAB 1 WRITE-UPInitial Machine TestingThe Original Design Experimenting With the Connections When the PCB board of the LCD screen was disconnected from the PCB circuit board the display output was turned off. When the white wire connecting the 16 tube PCR block to the PCB circuit board ability to regulate the temperature of the PCR machine was lost.
October 25, 2012 After finishing the diagnostic analysis, the PCR machine was tested by setting the thermal cycler program to three stages. Stage one was one cycle of 95 °C for 3 minutes, the second stage was 35 cycles of 95 °C for 30 seconds, 50 °C for 30 seconds, 72 °C for 30 seconds, and stage three was one cycle of 72 °C for 3 minutes. The test run lasted for about an hour and thirty minutes and confirmed that the temperature readings on the LED screen of the PCR machine and the computer matched.
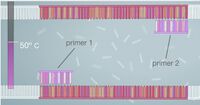
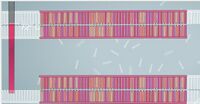
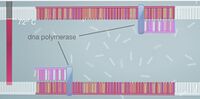
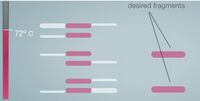
ProtocolsPolymerase Chain Reaction The PCR replicated the wanted DNA fragments from the patient. The PCR will heat up to 95°C and cool down to 50°C. Then heat back up to 72°C within one cycle. Over all there will be 30 cycles. At the end of the 30 cycles we have over a billion of the wanted fragments, and 60 unwanted DNA molecule strands in the solution. Step by Step instruction to amplify the Patient's DNA Sample: 1. Need to extract the DNA from the patient. 2. Put the DNA into a special PCR tube. 3. Add primer #1 to the PCR tube with the DNA. 4. Add primer #2 to the PCR tube with DNA. 5. Add Nucleotides (the A,C,T,and G). 6. Add the DNA polymerase to the PCR tube. 7. Place the PCR tube into the thermal cycler. 8. Set the temperature of the thermal cycler to 95°C and set the machine to run 30 cycles. 9. Now the thermal cycler cools down to 50°C and primer #1 and #2 attach to the single strands of DNA. 10. Now the thermal cycler temperature changes to 72°C. This begins the DNA polymerase. This pairs the DNA with its complimentary nucleotide through to the end of the DNA strand. 11. Repeat step 8-10 29 more times. 12. During cycle #3 the wanted DNA begins to appear. 13. The wanted piece of the DNA begins to double. 14. After 30 cycles are complete over a billion wanted DNA fragments will show in the DNA solution and there will be 60 copies of unwanted DNA molecules in the solution. Components: 1. MgCl_2 2. Taq DNA polymerase that lacks 5'--> 3' 3. dNTPs 4. Reaction buffers Reagent and Volume Table
Description of samples: Image 1: 3 drops of sybrgreen, 2 drops of calibrator solution. Dot was all blue not green. This implies the solution is negative for cancer. Image 2: 4 drops of sybrgreen, 2 drops of water solution. Dot was all blue no green. This implies the solution is negative for cancer. Image 3: 3 drops of sybrgreen, 2 drops of patient 1 solution A. Dot has some slight blurrs of gree. This implies the DNA solution is positive for cancer. Image 4: 2 drops of sybrgreen, 2 drops patient 1 solution B. Dot was all blue with no sight of green. This implies the DNA solution is negative for cancer. Image 5: 3 drops of sybrgreen, 2 drops of patient 1 solution C. Dot was all blue with no sight of green. This implies the DNA solution is negative for cancer. Image 6: 3 drops of sybrgreen, 2 drops of patient 1 solution D. Dot was all blue with no sight of green. This implies the DNA solution is negative for cancer. Image 7: 3 drops of sybrgreen, 2 drops of patient 2 solution A2. Dot was all blue with no sight of green. This implies the DNA solution is negative for cancer. Image 8: 3 drops of sybrgreen, 2 drops of patient 2 solution B2. Dot has a spec of bright green. This implies the DNA solution is positive for cancer. Image 9: 4 drops of sybrgreen, 2 drops of patient 2 solution C2. Dot had some spattered green. This implies the DNA solution is positive for cancer. Image 10: 5 drops of sybrgreen, 2 drops of patient 2 solution D2. Dot had a really green center. This implies the DNA solution is positive for cancer. Patient ID: Patient 1: 74065, Male, Age:63 Patient 2: 64835, Male, Age:46
1. Turn on the blue light in the Flourimeter using the switch for the Blue LED. 2. Place the smart phone accordingly so that the super-hydrophobic slide is in front of the smart phone. 3. Turn on the camera on the smart phone. **TURN OFF THE FLASH** Set the ISO to 800 or higher. Increase the exposure to maximum. 4. Set the distance between the smart phone and the machine so that the smart phone can take a clear picture of the droplet. 5. First label the blank pipettes according to the patients (A,B,C,D... all eight of them). The pipettes given by the instructor are color coded. The white coded pipette is used for water. the red coded pipette is used for the calibrator (the tube with the red dot). The blue coded pipette is used for the sybrgreen (the tube with the blue dot). The black coded pipette is used to pick up the waste and put it in the cup that collects the waste droplets. 6. First calibrate the machine (to make sure the machine works). Put two droplets of the cyber green on the first two dots in the middle. If the droplets are not connected then add a third droplet that combines the two droplets. 7. Then put two drops of calibrator solution with the calibrator solution. Then set the smart phone accordingly to take a clear picture of the droplet. Then put the black box on top of the phone and the machine so that when the the light is completely blocked and the shade of blue of green is shown when the picture is taken. 8. Record observations. 9. Remove the solution from the glass dish using the black pipette and discard it in the plastic cup given for the waste droplets. 10.Repeat step 6-9, but instead of adding calibrator solution, add 2 droplets of water. 11.Repeat steps 6-9 for patient 1 solutions (A,B,C,D) and patient 2 solutions (A2.B2,C2,D2).
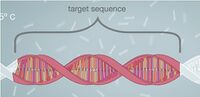
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology The r17879961 sequence will produce a cancer mutation at Chromosomes 22 of the gene sequence. The normal sequence has a T (Thymine) nucleotide at chromosome 22, while the mutation sequence has an associated C (cytosine) nucleotide. With the replication that occurs in the Open PCR machine, determining whether or not the r17879961 sample is cancerous is possible for there will be an exponential growth of the DNA sample if positive. A negative result will show a significantly smaller amount of replicated DNA; graphically this appears as a linear growth. Positive and negative strands are inserted into the PCR with a certain primer. The primer in the reaction will attach to the C nucleotide, which signifies that it is cancerous. Open PCR will attempt to replicate both strands, however, since only one of the DNA samples will be able to fully replicate with the primer, that is the one in which exponential growth can be noted. The negative strand will grow in a linear fashion, for it is only replicating one strand instead of both. The PCR process goes through 30 cycles; after the PCR process is complete, fluorescent dye is added to the solutions. The fluorescent dye causes the DNA to glow. Since the PCR has grown the double stranded positive DNA exponentially the fluorescent dye will be more noticeable in the cancerous strand in comparison to the negative strand which will have strands glowing just not as much as the positive sample. Therefore the cancerous DNA is in the sample that has most glow.
Source: Genetic Science Learning Center (2012, August 6) PCR Virtual Lab. Learn.Genetics. Retrieved November 8, 2012, from http://learn.genetics.utah.edu/content/labs/pcr/
The r17879961 sequence: GGAAGTGGGTCCTAAAAACTCTTACA[T/C]TGCATACATAGAAGATCACAGTGGC Normal has T at Chromosome 22 Mutation has C at Chromosome 22 Forward: CCTTCACCCAGGATTTTTGAG Backward: ATGTATCTTCTAGTGTCACCG
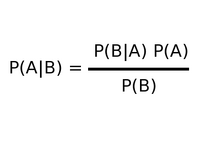
p(CIT) = (p(TIC)*p(C))/((P(TIC)*p(C)) +((p(TI~C)*p(~C))) Bayesian Theorem Results
| |||||||||||||||||||||||||||||||||||||||||||||||||||