BME103:T130 Group 1
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
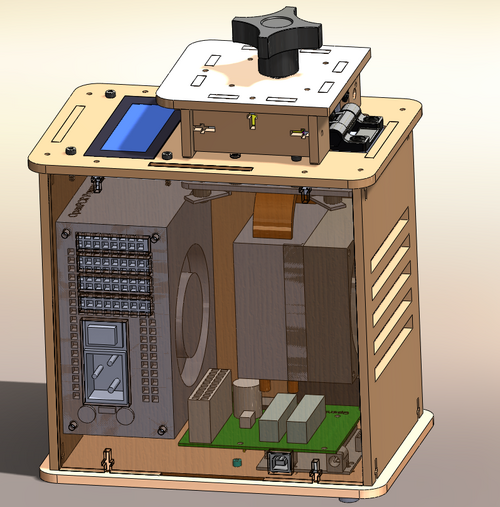
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingThe OpenPCR runs from your computer and connects through your USB port. You can add and delete steps, edit temperatures, and create thermocycler protocols from your computer.
- When the PCB board of LCD is unlplugged from the Circuit Board, the LCD goes off. - When the white wire connecting the circuit board to the heat plate, the LCD showed an incorrect reading of the temperature.
Our First practice run was conducted on October 25th, 2012. Our experience with the open PCR went smoothly. The test run took approximately one hour and forty five minutes. It was easy to set up and begin a test run. The only trouble we ran into was when adjusting the heat lid it was hard to tell when the lid was tightened down enough and it if very easy to tighten it too much and squish the test tubes in the PCR.
ProtocolsPolymerase Chain Reaction Polymerase Chain Reaction is the process of rapid duplication of a strand of DNA. The process first requires a DNA template, which contains the strand of DNA that is intended to be copied multiple times. The template strand is then been heated in order to separate the double stranded DNA and DNA Polymerase is added in order to create a strand of DNA that is complimentary to the original intended DNA strand, which creates a primer. Then the temperature is lowered so that the nucleotides will bind together in complementary to the primers in order to create a copy of the targeted DNA strand. The heating and cooling process will be repeated over a period of approximately 2 hours in order to create a large amount of DNA strand copies. Steps to amplify DNA 1.) Collect a blood sample from the patient.
2.) Use chemicals and centrifuge to separate the DNA materials from the rest of the blood components.
3.) Apply heat (95 degree Celsius) for 3 minutes to the DNA strand to separate the double stranded DNA into 2 single strands.
4.) In stage two, there are 35 cycles which consists of 95 degree Celsius for 30 seconds.
5.) Add primers and cool (57 degree Celsius, then 72 degree Celsius) the sample to allow the primers to bind to the separated original DNA strands.
6.) Then reheat the samples again to separated the resulting replicated strands.
7.) Add primers and cool the samples again.
8.) Repeat the process of heating and cooling in order for the primers to bind to all the
separated DNA strands.
9.) Over the course of 2 hours or so, there will be a large amount of duplicated DNA strands.
10.) Scientists can replicate DNA strands to study.
For a 25μL reaction volume
For a 50μL reaction volume
For a 100μL reaction volume
all credit goes to: http://www.promega.com/resources/protocols/product-information-sheets/g/gotaq-colorless-master-mix-m714-protocol/
Flourimeter Measurements Procedures for Fluorimeter Assembly: 1.)Unbutton the file box and flip over like shown below. 2.)Place glass slide down on fluorimeter. 3.)Place a drop of the sample on the slide then another close by to form one large drop like show below. 4.)Turn on light so that it may pass through the drop into the opposite hole. 5.)Place smart phone in stand. 6.)Move the system into the file box. 7.)Take picture and record image. (*close the lid for more accurate results)
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology There are many factors that contribute to a PCR reaction. First, a sample of DNA from a patient must be extracted. This sample is called template DNA. During the reaction, primers, or artifically synthesized bits of DNA, bind to the target sequence if it is present in the DNA. The enzyme taq polymerase's job is to regenerate the DNA strand that was melted away. Magnesium chloride (MgCl2) binds to the taq polymerase protein as a cofactor to help it function properly. In addition, there are dNTP's floating around in the solution of DNA. These deoxynucleotidetriphosphates are the bases (A,T,C,G) ready to be bound by the taq polymerase enzyme. During a PCR reaction, the solution of DNA is first heated to a temperature of 95 degrees Celsius to essentially melt the hydrogen bonds between the bases on the double strand of DNA. Next, the solution is cooled down to 57 degrees Celsius so the primers can bind to the target sequence of the DNA if it is present. Finally, the solution is heated up to 72 degrees Celsius, where polymerization occurs. This process of heating and cooling is repeated for a total of 34 cycles. This will ensure that the DNA has been amplified enough so it is able to be seen when a flourescent solution is added. Since the primers will not bind to the DNA if the target sequence is not present, a non-cancer patient will produce a negative result. This is because instead of having millions of double stranded DNA in the solution, the solution will only contain single strands. The specific cancer-associated gene sequence being analyzed is the r17879961 SNP. The mutation is present at position number 29,121,087 in the DNA. The "ATT" sequence in a non-cancer human is replaced by the "ACT" sequence. Therefore, during a PCR reaction, primers will bind to the sequence, "ACT", since it is the cancer gene.
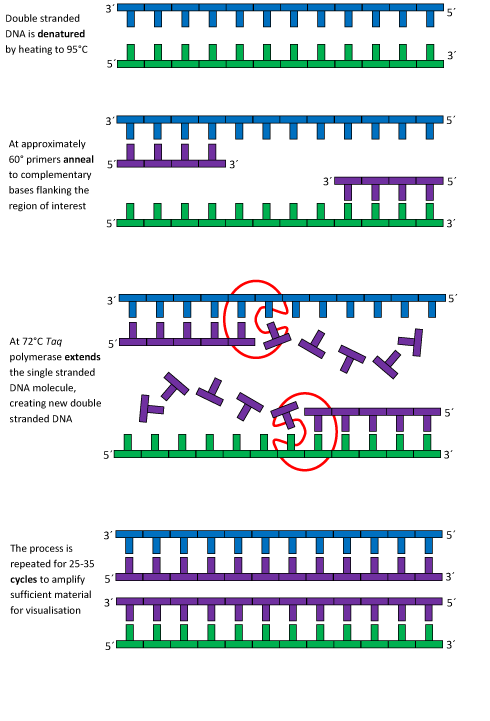
This image shows what happens to the DNA strand each time it replicates. First, the bonds are melted and separated. Then, primers attach themselves to each end of the DNA strand. Next, taq polymerase adds the appropriate bases to create a new piece of double-stranded DNA. Finally, this process is repeated for approximately 30 cycles. Image borrowed from: [1]
ResultsImageJ Software Processing
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||