BME100 f2018:Group6 T1030 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||
OUR TEAM
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In BME 100 lab, 17 teams took 34 patients' DNA samples and tested them for a single nucleotide polymorphism associated with Parkinson's Disease. Each group recieved 2 patient samples, a positive sample, and a negative sample. They ran a total of 8 samples: 3 runs of the first patient, 3 runs of the second patient, and a positive and negative control. We ran patient samples 3 times to make sure the error in our results were minimal. We also compared our samples to the positive and negative control to verify the presence or no presence of the SYBR green color. We viewed our samples in a fluorimeter with SYBR Green I added to the sample so we could watch the sample's fluorescent color if it was positive or no color if it was negative. We took pictures of our samples and processed them in ImageJ. We calibrated ImageJ using Calf Thymus DNA solution and SYBR green I and measured the different ranges of green color with different DNA concentrations. The classes' final data was kept in a spreadsheet with the doctor's conclusion and our conclusion. We analyzed the accuracy of our results using Bayesian statistics.
The probability that the patient will develop the disease, given a positive final test conclusion, was less than 0.5, meaning they are likely to not develop the disease. The probability that the patient will not develop the disease, given a negative final test conclusion, was very close to 1.00. This means that the patient will not develop Parkinson's disease given a negative test result.
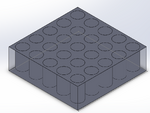
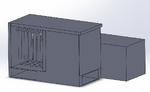
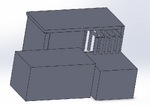
Intro to Computer-Aided Design3D Modeling Our group worked with Solidworks to create a model of our device. Though slightly more complicated than the other recommended software, Solidworks allows for more precision (and more options) for 3d modeling. Two out of our group had previous experience with the software. Our device is largely a modification on existing PCR systems, so our team had a reference for the general shape of the 3d model. We also used files found on the provided website for existing models with accurate dimensions of PCR tubes. Our Design
Feature 1: ConsumablesThe following items will be included in the kit provided by the manufacturer: -Primer mix and acting reagents within the PCR reaction: To catalyze the reactions and allow for measurement. -25 Stainless Steel micropipette tips: Reusable micropipette tips that can be sterilized in an autoclave, also very eco-friendly for our environment. -Stainless steel PCR tray: Will include 25 spaces for samples that are set in rows of 5 that can be sterilized as well as be very eco-friendly for our environment. These allow for the mixtures of DNA and reaction fluids to be used well, acting as indicators for specific samples. Feature 2: Hardware - PCR Machine & FluorimeterThe AZS100 PCR machine that we have implemented into our idea or new concept would be a stainless steel, eco-friendly, robotic machine. This particular way of robotics will lessen the human mistakes made in the wet lab. Also, this machine will self-clean all stainless steel pipette tips used to transfer the gene mix used inside the PCR machine into the stainless steel tubes with fitted caps or lids. At the bottom of the AZS100 PCR machine will have two openings, one in the bottom front layer of the machine for the insertion of the PCR samples tray. Once the tray is locked into place the robotic steel pipettes when then dispense the already mixed gene samples into each of the 25 spaces, 5 at a time in a matter of five seconds. Also, the machine will have another opening in the side bottom going from the tray loader to the actual PCR machine whereas it slides slowly from it's loading position, the scientist may see inside to reach in to make sure the sample tray was capped correctly before going into the actual PCR machine itself.
| |||||||||