BME100 f2018:Group16 T1030 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||
OUR TEAM OF NINJASLAB 4 WRITE-UP GROUP 16ProtocolMaterials
PCR Reaction Sample List
DNA Sample Set-up Procedure
OpenPCR program
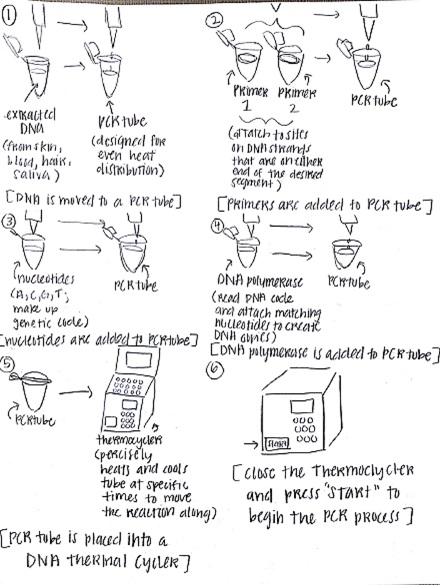
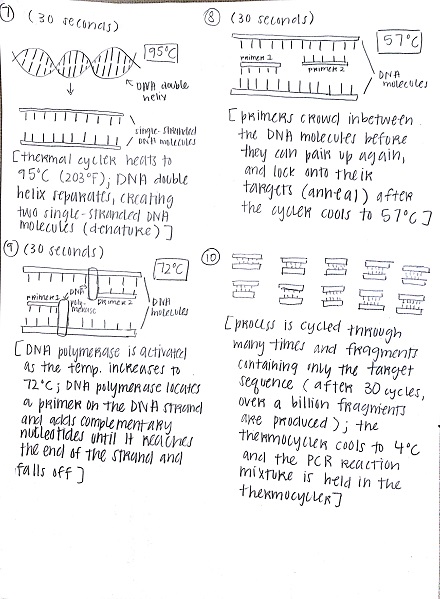
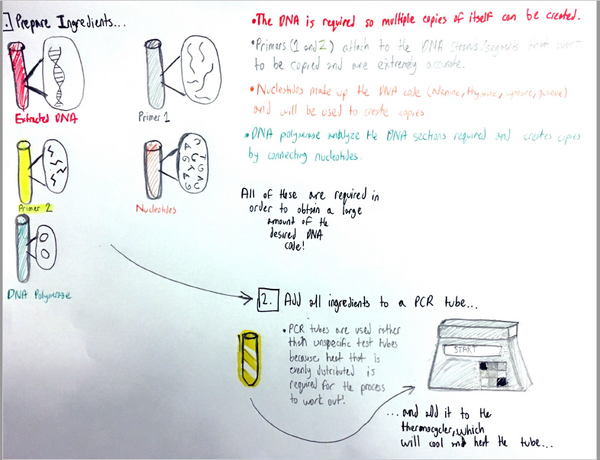
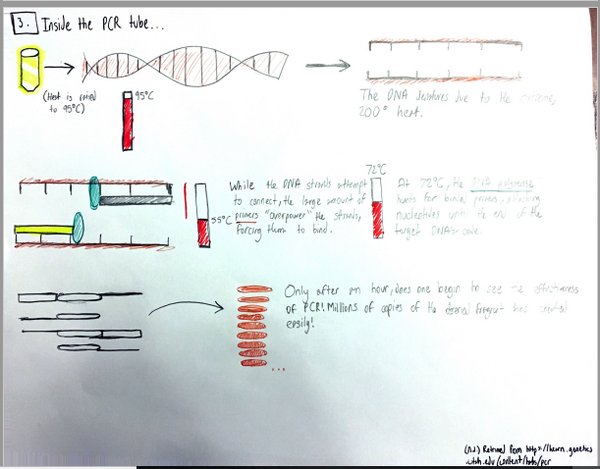
Research and DevelopmentPCR - The Underlying TechnologyA PCR, or a polymerase chain reaction, is a commonly used procedure in order to replicate a desired section of a DNA fragment. It is a relatively simple and cost efficient way to obtain said fragments in extremely large amounts; up to billions of copies can be made. The uses of PCRs vary widely, from its most common uses in biology and chemistry, to forensics and pathology. In PCR, there are four components vital to the reaction: the DNA template, primers, "taq" polymerase, and deoxyribonucleotides (dNTPs). The template DNA acts a guide for the PCR reaction to make more copies of that particular strand of DNA. Forward and reverse primers attach to each end of the strand and isolate the part of the DNA that will be replicated. The "taq" polymerase locates the primer and reads the template strand to produce a complementary strand from dNTPs, the building blocks of DNA (adenine, thymine, guanine, and cytosine). Once the tube with the reaction mixture with all the listed components is placed into a thermocycler, a series of steps ensue to amplify a particular strand of DNA. The initial step is heating the mixture to 95°C for 2 minutes, so that the DNA can denature at this temperature in 30 seconds from double stranded DNA to single stranded DNA. The next step is called annealing, where the mixture is cooled down to 57°C so that the primers can attach to the DNA template and prevent the single stranded DNA from adhering to the other. Next, the mixture is heated to 72°C for 30 seconds, called extending, and then held for 2 minutes. At this point, the "taq" polymerase activates, locates the primers, and starts attaching the complementary dNTPs to the single stranded DNA. Lastly, the thermocycler cools down to 4°C, and the PCR reaction mixture is held in the thermocycler. The four nucleotides adenine, guanine, cytosine, and thymine engage in base-pairing through hydrogen bonds to form DNA. The bases adenine and thymine anneal to each other, while guanine and cytosine anneal to each other. During the annealing and extending steps of thermal cycling, base pairing occurs. This happens first when primers locate and attach to the single stranded DNA through base pairing (annealing), and subsequently in extending, when the "taq" polymerase utilizes base pairing to attach dNTPs to the single stranded DNA to create a double helix once again.
DiagramsSNP Information & Primer DesignBackground: About the Disease SNPNucleotides are units that comprise of nucleic acids, which are composed of a five sugar carbon, at least one phosphate group, and a nitrogenous base. Polymorphism is a type of genetic variation where nucleotide differences exist between individuals in a population. The species this variation is found in is Homo sapiens, otherwise known as humans, on chromosome 12 of 46. The listed clinical significance of this SNP is with uncertain significance allele. The outlying condition linked to the disease SNP is Parkinson's disease, which is associated with mutations in the LRRK2 gene. LRRK2 stands for leucine-rich repeat kinase 2 gene, and some of its functions include GTP-dependent protein kinase activity, MAP kinase kinase activity, and actin binding. GTP dependent protein kinase activity refers to the reaction to break down ATP and a protein serine/threonine to ADP and a protein serine/threonine being catalyzed in the presence of GTP. MAP kinase kinase activity (MAP2K) is involved in the mitogen activated protein (MAP) kinase signaling pathway, in that it is activated by MAP3K and phosphorylates to activate MAPK to perform various cellular functions, such as differentiation and survival. Lastly, actin binding affects the movements of actin filaments within the cytoskeleton. Genes have variant forms known as alleles. The disease-associated allele in this SNP contains the codon GAG. The numerical position of the SNP is 40315266. The non-disease forward primer sequence is TTAAGTGACTTGTACTTTGT. The numerical position 200 bases to the right of the rs721710 SNP is 40315466. The non disease reverse primer sequence therefore is TGAAGCTCTTCAAGTAGTCT. The resulting disease forward primer according to the disease SNP nucleotide is TTAAGTGACTTGTACTTTGA. The disease reverse primer is the same as the non-disease reverse primer. Primer Design and TestingThe non-disease forward and reverse primers are tested using the UCSC In-Silico PCR non-disease human genome sequence database. The non-disease forward and reverse primers (that center around the SNP rs721710) have a match in the genome sequence and result in a 220 bp sequence. However, the disease forward and reverse primer does not have a match in the database because the test only validates non-disease primers rather than diseased ones, and the one missense mutation that alters the codon from GTG to GAG causes the mismatch to the human genome sequence. 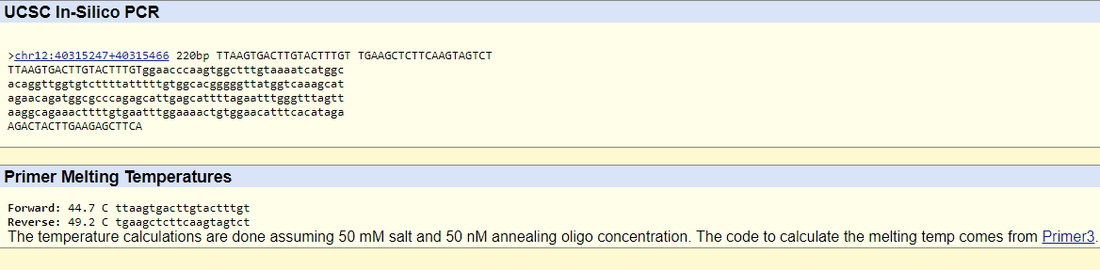 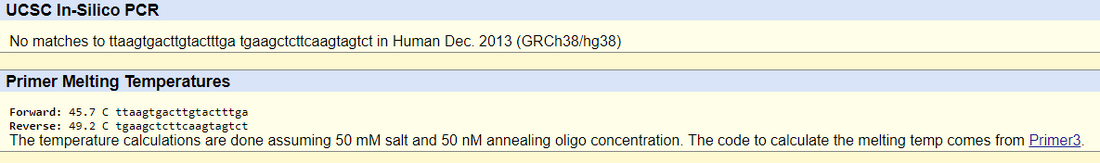 |
||||||||||||||||||||||||||||