BME100 f2018:Group11 T1030 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
B.B. BioMed
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System
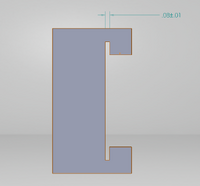
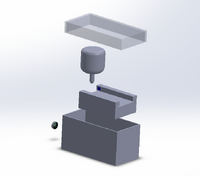
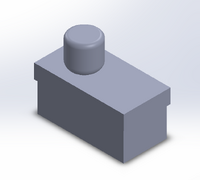
The results from calculations 1 and 2 imply that using individual PCR replicates to conclude that a person has a disease SNP is reliable. This is seen by both of the calculations being so close to one. The first shows that if a person has a positive result they have a high probability of having the SNP. The second calculation shows that having a negative result will have a high probability of not having the SNP. The results from calculations 3 and 4 imply that PCR is pretty reliable. For calculation 3, the reliability was really low, closer to 50%, when the PCR test was used for positive specimens. When PCR is used to test for a negative result, then the results are very reliable and can generally be used to correctly predict the results. Possible sources of error could have occured in transferring the liquid from the PCR tubes into the big Buffer tubes. Remaining liquid could have been dicarded along with the used micropipette tip. Human error that could have occured in the detection steps would be placing the SYBR Green 1 drop on the wrong side of the glass slide. The drop of SYBR Green 1 would disperse if placed on the wrong side. This would affect the analysis using ImageJ because an innacurate area recording would be taken since the shape of the drop would be changed. A third possible source of error could have occured in aligning the drop and the blue LED light. Failure to do this would not ensure a high emission of light when a high concentration of DNA was added to the SYBR Green 1. Intro to Computer-Aided Design3D Modeling Our Design This design was chosen as we believed that the main issue with the original product was the use of the phone camera as well as the intrusion of light into the system due to the design of the storage unit. We virtually eliminated this by using a camera that can be connected to a computer through a wireless connection. This along with the seal from the lid makes it so that there is no extra light when taking the picture. This leads to a more accurate result. Another way to help improve the design is by making a container that stores the SYBR Green when in use. This container is solid so that no light can come through and will automatically pipette the appropriate amount of solution when it is told to do so. This process is another step to prevent light from affecting the data.
Feature 1: Consumables
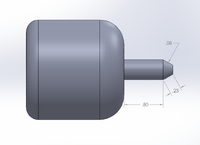
Feature 2: Hardware - PCR Machine & FluorimeterIn our system, both the Open PCR machine and the fluorimeter will be used and are extremely important and relevant to the accuracy of the test. Both of these individual part have a particular job and help develop the test results. The Open PCR machine and the fluorimeter have very specific jobs that are necessary to complete the system. Our system does have several changes that were made to the fluorimeter to improve the process and system. This being said, it is important to mentions the weaknesses of the original fluorimeter and the improvement made.
| ||||||