BME100 f2015:Group2 8amL4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction Sample List
HEATED LID: 100°C
FINAL STEP: 72°C for 2 minutes
Research and DevelopmentPCR - The Underlying Technology
Template DNA is the target strand of DNA where the disease in question exists as a mutated base pair.
Primers are short segments of DNA that are synthesized in the lab. Primers consist of forward primers and reverse primers, each made of about 20 nucleotides that match with a unique segment of the template DNA. Primers attach to either end of the template DNA strand. They allow DNA Polymerase to attach to the DNA to begin replication.
DNA Polymerase is the complex of proteins that is responsible for replicating a strand of DNA by matching base pairs of nucleotides with the template DNA. DNA Polymerase attaches to a segment of template DNA via primers, then runs along the DNA like a train track, pulling nucleotides from the surrounding environment to build a complimentary strand. The most frequently used polymerase is Taq Polymerase, which is taken from the Thermus aquaticus (thus, “T-aq”) bacteria, which lives in the hot springs of Yellowstone National Park. Taq polymerase is used because it can function at the high temperatures required for the PCR process without denaturing.
Deoxyribonucleotides, or simply “nucleotides,” are the A’s (Adenine), T’s (Thymine), C’s (Cytosine), and G’s (Guanine) that code for the amino acids that make up proteins. Adenine matches to Thymine and Cytosine matches to Guanine via hydrogen bonds. In PCR, nucleotides are pulled from the surrounding environment by the Taq Polymerase and are matched to the template DNA strand in order to create a replica strand.
This step heats the polymerase to prepare it for the PCR reaction.
During this step, the hydrogen bonds joining the base pairs of nucleotides together in the template strand of DNA are separated, or "denatured." This separation sets the stage for the primers to bind to their respective strands in order for the PCR reaction to begin.
Now that the template DNA is separated into two strands, the primers can now bind to their target sites during this step. This temperature is just hot enough for the primers to behave as they should and bind to the DNA strands. Once the primers bind to the DNA, the Taq Polymeras binds to the primers.
At this temperature, the PCR reaction begins. The Taq Polymerase runs along the single strands of DNA in the 5' to 3' direction and builds complimentary strands by pulling unbound nucleotides from the surrounding liquid environment. The Denature, Anneal, and Extend steps are then repeated, causing the target DNA strands to multiply exponentially.
This step insures that any remaining PCR reactions can be completed.
The replicated target DNA strands can now be stored for short-term use during this step.
SNP Information & Primer DesignBackground: About the Disease SNP What is a nucleotide? A nucleotide is the structural unit or building block of a DNA molecule. What is a polymorphism? A polymorphism refers to the occurrence of two or more phenotypes within the same species.
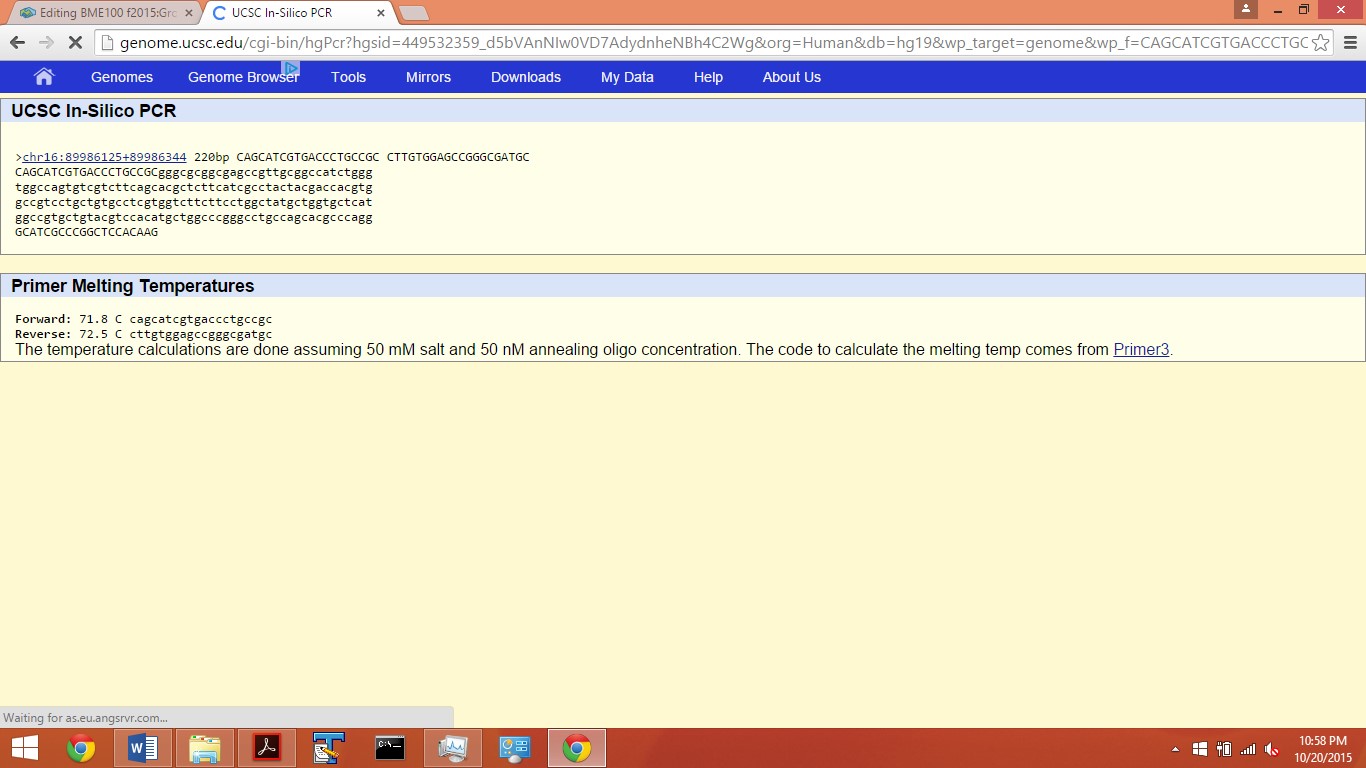
MC1R stands for melanocortin 1 receptor (alpha melanocyte stimulating hormone receptor). The MC1R gene provides instructions for making a protein called the melanocortin 1 receptor. The first three terms unique saw were G-protein coupled peptide receptor activity, melanocortin receptor activity, and melanocyte-stimulating hormone receptor activity. An allele is one of two or more alternative forms of a gene that arise by mutation and are found at the same place on a chromosome. The disease-associated allele contains the sequence TGG. The numerical sequence of the SNP is 89919736. The non-disease forward primer (20nt) is 5'-CAGCATCGTGACCCTGCCGC. The numerical position exactly 200 bases to the right of the disease SNP is 89919936. The non-disease reverse primer is 5'-CTTGTGGAGCCGGGCGATGC. The disease forward primer is 5'-CAGCATCGTGACCCTGCCGT and the disease reverse primer is 5'-CTTGTGGAGCCGGGCGATGC.
| |||||||||||||||||||||||||||||||||