BISC220/S10: Mod 1 RasMol Tutorial
Bio 220 RasMol Tutorial
What is RasMol and how do I get a copy for my computer?
RasMol is a computer program, developed by Roger Sayle, that allows the user to visualize, manipulate and investigate the structural characteristics of large variety of molecules, both large and small. The software runs on a variety of different types of operating systems and can be downloaded from the web. If you are interested in downloading a copy of RasMol for your computer (for Macintosh computers it is sometimes called RasMac), connect to the RasMol web site at http://www.openrasmol.org/ or http://www.bernstein-plus-sons.com/software/rasmol/INSTALL.html. You will find instructions for downloading a copy of RasMol to your personal computer. Be sure to choose the 2.7.1.1 version of Ras in the operating system used by your computer. Version 2.7.1.1 is the one that you will be working with in class today, so make sure you are downloading the same one to continue your work outside the lab. For today’s exercise there is no need to download the program since we will use the copy already found in the RasMol folder on the desktop of the lab computers.
How do I load/close a file using RasMol? Loading a file into RasMol:
Open the RasMol folder and click on the icon that consists of three connected spheres labeled raswin.exe. An empty window with a black background (called the Main Window) will appear on the screen. Under the File menu open the file in the RasMol folder labeled 1JYVsubA.pdb. This file contains the coordinates for the atoms in one subunit (the molecule is a homotetramer) of the enzyme β-galactosidase from E coli. The coordinates of the atoms were obtained by X-ray crystallography and are available to the public at the protein data bank (http://www.rcsb.org/pdb/). A wireframe 3D image of one subunit (A) of the enzyme β-galactosidase should now appear in the Main Window. By clicking, holding and then moving the mouse (with the crosshair within the margins of the Main RasMol Window) you will immediately see how easy it is to rotate the molecule in virtually any direction. We will use this protein to practice using the RasMol commands. Look for the Command Line in the tool bar at the bottom of the RasMol window. Open it by clicking on it. To close a file select Close from the File menu.
Please Note: In this tutorial the commands to be typed into the Command Line Window will be in bold and the options to be selected from the RasMol menu will be in italics. You will find it useful to adjust the size of the windows such that they do not overlap on the screen.
How can I use RasMol to investigate my 3D image?
Now let’s investigate RasMol’s menu and explore some of the options for manipulating the 3D model of β-galactosidase. The three most useful menu options for manipulating the image in the main window are Display, Colours and Options. However, before experimenting with these options try to manipulate the size and the position of the image in the Main Window using the zoom and translate options.
ZOOM--it is possible to enlarge or reduce the image on the screen by using the mouse in combination with the shift key. Hold the shift key down and left click and hold the mouse button. Moving the mouse up or down reduces or enlarges the image (crosshair must be within the margins of the Main Window).
TRANSLATION--you can position the image anywhere you want in the main window by moving the mouse after a right click and hold on the mouse (crosshair must be within the margins of the Main Window).
Now try clicking and dragging down to the various choices in the Display menu option. Other than Wireframe, which is the type of image that currently exists in the window, the various options dramatically change the image that you see. Below are some capsule definitions of the various display options.
Menu Items:
Display
- Wireframe: molecule shown as a series of intersecting lines which represent bonds-- reveals bond distances, angles and the overall geometry of the molecule; colors denote the atoms that form the covalent bonds denoted by the lines--red (oxygen); light blue (nitrogen); yellow (sulfur); light gray (carbon)
- Backbone: a gray rod showing geometry of the backbone or main chain of the protein--this cylindrical line or rod connects adjacent carbons.
- Sticks: similar to wireframe only the bonds are thicker lines--see #1 for color coding.
- Spacefill: represents atoms as spheres with van derWaals radii.
- Ball & Stick: atomic nuclei are shown as colored spheres of identical size, bonds as connecting sticks--see #1 for color coding
- Ribbons: main chain shown as a thick gray ribbon that traces the backbone of the molecule. Clearly reveals those regions of the molecule that form α helices and β sheets
- Strands: similar to ribbons but now a series of gray lines trace the main chain or backbone of the protein
- Cartoon: in general similar to strands and ribbons-- shows non- α helical and non-β sheet regions of the molecule as a gray rod (similar to #2 backbone in general appearance but is smoothed to form curves rather than straight lines between α carbons). Note arrowheads on the β sheets, allowing one to discern the parallel or antiparallel nature of the association of the strands.
Useful Colour Selections
- Monochrome: molecule becomes white
- CPK: see code given above in #1 under Display
- Temperature: assigns amino acids colors according to the degree of mobility/uncertainty in their position as given in the crystal coordinates--high values in red, lower in blue. Try Wireframe and note that the surface residues have the higher values. Does this make any sense?
- Structure: α-helices are colored magenta ; β-sheets are yellow; loops are blue and the remainder of the molecule is white with shades of gray--quite effective with the molecule in the Cartoon, Ribbons or Strands configuration
Options:
Note: turn off (clicking on the menu selection a second time) each option after you try it.
- Slab mode: allows you to slice your way through one plane of a molecule by rotating it.
- Hydrogens: reveals hydrogen atoms--none in the 1JYVsubA.pdb file
- Hetero atoms: shows non-protein atoms
- Specular: enhances 3D effects by providing highlights (use with Spacefill)
- Shadows: enhances 3D effects by providing shadows (use with Spacefill)
- Stereo: can give you a stereo image if you care to stare at the screen for long periods of time--not highly recommended.
- Labels: labels all of the amino acids (three letter abbreviations and numerical positions in the sequence. It can get very messy with all of the amino acids named. However, it is possible to restrict the command to certain regions or to enhance the resolution by zooming into region of the molecule (see below for zooming commands).
Next we’ll explore a few of the many commands that can be entered into the Command Line Window and practice using the menu items introduced above. Let’s try this in the context of reviewing some of the basics of protein structure using β-galactosidase as our model.
The amino acids and the primary structure of a protein:
- Type show sequence into the Command Line Window. This command reveals the amino acid sequence of β-galactosidase in the window. You will see the amino acids numbered 43-1023 with their names abbreviated by the standard three letter codes. Numbering always begins at the amino terminus of a chain.
- Type restrict Thr741,Thr742 or simply restrict 741,742 into the Command Line Window--the restrict command eliminates from the screen everything except that designated in the expression following the word restrict. In the Command Line type save followed by a name for your file that ends with .pdb. This file will be saved to the RasMol folder. You can now see how it’s possible to create a new file of any portion of the entire molecule just by using the restrict command and saving the file. Now Close the 1JYVsubA.pdb file and Open the one you just saved. Change the display to Ball & Stick and the Colour to CPK and activate Labels under the Options menu. Try clicking on some of the atoms and see what happens in the Command Line Window--Identify the two α-carbons--the abbreviation used is CA. (The picture below should remind you what we mean by α-carbons.) See if you can find the peptide bond and the side chains or R groups of the two threonines. As I’m sure you have already observed there are no hydrogens in this molecular model.
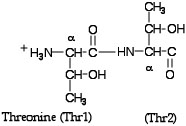
Before we leave our threonines, try the command dots on. This command generates a series of dots roughly corresponding to the van derWaals radii of the constituent atoms-sort of a transparent view through a Spacefill view (if you change to the spacefill representation you’ll see what I mean). Now try background white and color dots black --another way to play with the image. Close this file and Open the 1JYVsubA.pdb file.
Secondary Structure:
Select Ribbons from the Display menu and Structure from the Colours menu. You can now visualize the location of the secondary structural elements. For β-galactosidase we can see several α-helices (magenta), and β-sheets (yellow). Using the Cartoon mode under the Colour menu will generate arrow heads which allow you to discern if the β-sheets are parallel or anti-parallel. Loops are colored blue and regions of the sequence that do not possess any defined secondary features are shown in white.
Let’s look a bit more closely at some of the structural features of one of the α-helices present in β-galactosidase. Let’s restrict our view to just one of these helices: type restrict 466-478 and return to the Ball & Stick view and CPK colours. Try rotating the helix. You will probably find that it swings a bit wildly because the center of rotation is still the center of gravity of the complete β-galactosidase molecule. This will become even more evident if you zoom in on the helix. If you want to have it rotate about a particular atom, type set picking center, and use the mouse to click on an atom that you would like to be the new center of rotation. The command line will read; Rotating about ....... Now type set picking ident--this returns the mouse to its function of identifying atoms that are clicked on. If you have not already done so increase the image size and center it using the shift and option keys. Now type hbonds on and then hbonds 0.1 (adjusts the thickness of the bonds) and color hbonds green --you should now see a series of green lines representing the hydrogen bonds connecting the main chain nitrogens and carbonyl oxygens. It is these hydrogen bonds that stabilize the α-helical architecture. How about the side chains? How are they positioned in an α-helix? Type in the Command Line Window (not selected from the menu) ribbon and then color ribbon red.
Rotate the image using the mouse so that you are looking directly down through the cylinder of the helix. You should be able to now clearly see how the amino acid side chains project out from the main chain. We can also play around a bit more with the side chains. Type select sidechain then color blue . The select command is similar to the restrict command in that after you select a give atom, chain, set etc., subsequent commands are performed only on the selected region. It differs from the restrict command in that the remainder of the molecule stays on the screen. The side chains should now all appear blue. Are any of these side chains hydrophobic? Type select hydrophobic and then color green and you’ll see those side chains that fall in RasMol’s hydrophobic set of amino acids. If you click on them in the Main Window you’ll be able to identify them by the output in the Command Window. Consult the help manual available in the RasMol folder for a complete list of “sets”. Now select Select all and Ribbons, Structure from the menu and continue on. You should also be able to see where the helix that we just viewed fits into the entire molecule.
Let’s try a similar approach to that used on the α helix to look at β-galactosidase’s β-sheets.. Return to the Wireframe view and CPK colours and type restrict 627-697. Adjust the center of rotation by using the commands set picking center and set picking ident. Increase the image size and center it. Now type hbonds on and then hbonds 0.1 (adjusts the thickness of the bonds) and color hbonds green. You should now see a series of green lines representing the hydrogen bonds connecting the main chain nitrogens and carbonyl oxygens. It is these hydrogen bonds which stabilize the β-sheet’s architecture. Select Cartoons from the Display menu and Structure from the Colour menu. Are these parallel or anti-parallel β-sheets? How about the side chains? How are they positioned in a β-sheet?
To answer this question return to the Ball and Stick display type in the Command Line Window (not selected from the menu) ribbon and then color ribbon red. Rotate the image using the mouse so that you are looking directly down at the edge of a sheet. You should be able to now clearly see how the amino acid side chains project from the upper and lower surfaces of the sheet. We can also play around a bit more with the side chains. Type select sidechain then color blue. Are any of these side chains acidic? Type select acidic and then color green and you’ll see those side chains that fall in RasMol’s acidic set of amino acids.
Tertiary Structure:
The overall three dimensional geometry of a protein is called the tertiary structure. The unique structural architecture of a given protein is determined by a variety of interactions between the side chains of the constituent amino acids. We have already seen examples hydrogen bonding. However, the hbonds that you viewed earlier were not between the side chains of amino acids but were main chain hbonds involved in stabilizing secondary structural elements of the protein. Remember that hbonds between side chains and between side chains and water can play an important roles in stabilizing protein architecture. The following are other forces that are also thought to contribute to tertiary structure:
- sulfur bridges
- van derWaals forces
- hydrophobic interactions
- ionic interactions
The fundamental unit of tertiary structure is called a domain. Each β-galactosidase subunit has 5 domains. The third domain contains the majority of the residues that comprise the active site. Bring your image back to its original configuration (Select all; Wireframe; CPK colours) land type select 334-627. Choose Ribbons and Structure from the Colour menu. You should now be able to see the location of the third domain in the protein. Information on the other domains can be found in Jacobsen et al. (1994). Now let’s eliminate the remainder of the molecule from the viewing window by typing restrict 334-627.
One last bit of terminology we need to introduce here is the fold. A fold is a frequently encountered arrangement of secondary structural elements. For example the TIM barrel fold is one in which a series of α-helices and β-sheets are organized into the shape of a barrel. This particular arrangement, which was first seen in triose phosphate isomerase (TIM), has subsequently been found in a number of different proteins whose three dimensional structures have been solved. The reason we are introducing this is that β-galactosidase has a distorted TIM barrel fold in its third domain. You should be able to visualize this barrel fold by rotating the third domain in the Main Window. As noted above, this barrel fold forms part of the active site. The inside of the barrel is a β-sheet with the outer surface defined by a series of α-helices. Some authors (Branden and Tooze, 1999) break this down further and visualize the barrel fold as composed of a series of β/α structures (the building blocks) called supersecondary structures or motifs.
Quaternary Structure:
Quaternary structure describes the 3D structure of a multisubunit protein, particularly how subunits interact and fit together. β-galactosidase consists of four identical subunits—a homotetramer. Close the 1JYVsubA.pdb file and open 1JYV.pdb file. This is the file for the complete or holoenzyme. You can assign a different color to each subunit by typing in the command line select :A. and then color blue . If you want to you can repeat this for subunits B C and D. As noted previously the third domain contains the active site so it is an area of particular interest. Type select 334-627 (the included in the third domain) and then color red. You should now be able to see how the four active sites are arranged in the molecule.
Studying Heteroatoms:
RasMol calls all non-protein molecules heteroatoms. This generally includes the water molecules seen in the crystal (if the structure of the molecule was solved by X-ray crystallography), metal ions and/or a variety of organic ligands. Using a single subunit file (1JYVsubA.pdb) in the Wireframe form and CPK colours type select hetero and then select Spacefill from the menu. Identify these heteroatoms by clicking on them and monitoring the output in the Command Line Window. DMS is dimethyl sulfoxide (a cryoprotectant used in some of the experiments- not biologically relevant to the function of the enzyme). You should also find sodium and magnesium ions and two larger organic molecules called 145 2001 and 145 2002. These are the codes in this coordinate file for the synthetic substrate (ortho-nitro-phenyl-galactoside (ONPG)) that we will be using in our lab experiments to measure enzyme activity. Remember the natural substrate is lactose. It is often of interest to researchers to study the binding sites for ligands. For example, questions such as:
- Where is the active site or binding site for a given ligand?
- What types of amino acids form the geometry of a binding site?
- Does the chemistry of the amino acids (i.e. acidic) give you some clue as to the forces involved in binding?
- If the molecule is an enzyme can one identify the amino acids that might play an important role in mediating the catalysis? A very useful command for this type of analysis is the within command. With the subunit in the Wireframe mode with CPK colors type restrict within (14.0,2001). This restricts the view to the atoms within 14.0 units (this can be varied) of the ligand 2001 (remember this is one of the numerical codes for the two synthetic substrate molecules) This should allow you to identify the various amino acids and other hetero atoms in the immediate vicinity of one of the two bound ONPG molecules in each active site.
Note: This is one possibility for the image you might generate as part of your homework assignment, but it is probably not the best you can do. You will want to learn more about the structure and crucial amino acids in the active site by reading the Jacobson et al (1994) article and other references before you create an image to turn in. To save a favorite image refer to the next section.
Can I save images that I’ve generated using RasMol?
It’s possible to save images in a number of fashions. In the exercise immediately above you saved files generated by the restrict command. In this case the files saved contain only the coordinates for the restricted portion of the molecule. This method of saving a file does not preserve any changes you might have made to the image (i.e. color, type of representation etc.) before you saved it (other than restrict). When such a file is re-opened in RasMol it comes up as a standard CPK Wireframe. Perhaps your best bet to save your favorite molecular images is as static images or “snap shots” of your molecules using the Export command. Save your images as GIF files and you can then import them into Microsoft Word or PowerPoint. When printing it is best to change the background of the image to white (background white in the Command Window).
After you have saved your image you are ready to work with ClustalW.
References:
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular Biology of the Cell, 4th ed.
Branden C, Tooze J (1999) Introduction to Protein Structure. 2nd ed. Garland Press, New York.
Jacobson RH, Zhang X-J, DuBose RF, Matthews BW (1994) Three-dimensional Structure of β-galactosidase from E. Coli . Nature 369; 761-766.
Juers DH, Heightman TD, Vasella A, McCarter JD, Mackenzie L, Withers SG, Matthews BW (2001) A structural view of the action of Escherichia coli(lacZ) β-galactosidase. Biochemistry. 40: 14781-94.
Martz, Eric, 2000 August 28. Molecular Visualization Freeware: RasMol Home page <http://www.umass.edu/microbio/rasmol> Accessed 2004 Jan. 17.
Roth NJ, Rob B, Huber RE (1998) His–357 of β-galactosidase (Escherichia coli) Interacts with the C3 Hydroxyl in the Transition State and Helps To Mediate
Catalysis Biochemistry 37: 10099-10107.
RSCB Protein Data Bank. 2004 February 24. RSCB-PDB Home page <http://www.rcsb.org/pdb/>. Accessed 2004 Jan. 17.