BISC209/F13: SpectraMax 340PC instructions
SpectraMax 190:
Located in the back of lab 302
Molecular Devices SpectraMax 190 and SoftMax Pro6.3™ software instructions

View of back of Spectra Max 190
Turn on the Spectra MAX 190 spectrophotometer using the switch located next to the plug in the back on the right hand side (as you face the spec). The drawer will open. The drawer may, or may not, close on its own. If it doesn't close, please press the DRAWER button found on the Spectra MAX control panel (pictured below). It is important to keep the drawer closed as much as possible to prevent dust from entering the spectrophotometer.


The machine will automatically start a required calibration. Allow up to 10 minutes for the Specta MAX 190 to warm up
after the calibration and BEFORE you attempt to insert your plate.
The computer should be on. (If not, turn it on.)
When the warm up period is over, double click on the SOFTmaxPRO6.3 shortcut on the desktop of the computer. ![]()
The draw may open again, or you can open it manually by pushing the DRAWER button ONCE. (Drawer button is found on the Spectramax control panel)
Position your 96 well plate into the tray drawer so that it fits securely in the holder and the well A1 is to the top left. Close it using the DRAWER button .
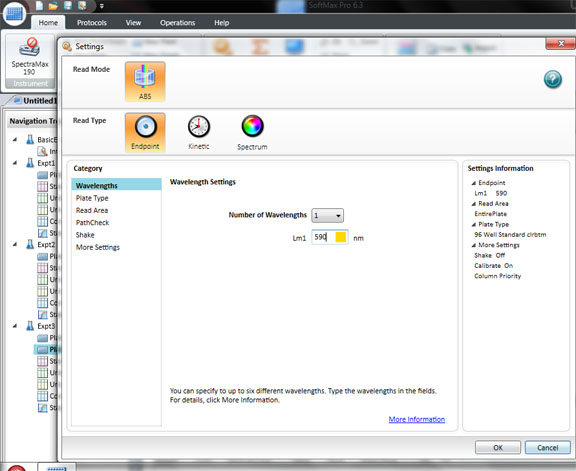
Click: SETTING
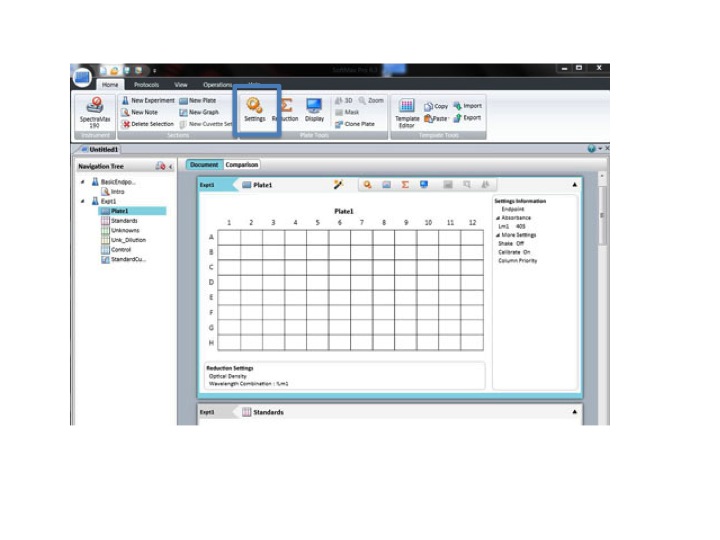
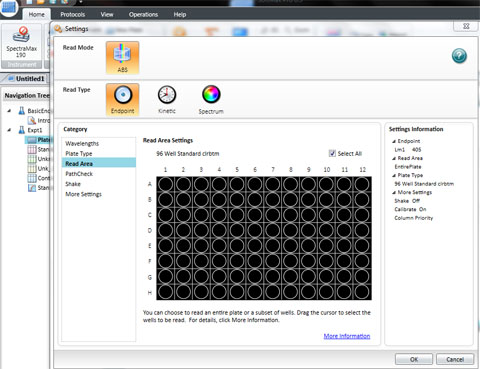
Set the wavelength to 590 for the BIOLOG Ecoplate. Check that Plate type is 96 well. Check that the Read area is the whole plate.
Click OK
The spec is ready to read all the wells in a 96 well plate. Click Read.
You will hear the spectrophotometer slit lamp passing over the plate to "read" it, meaning that it is measuring absorbance at the 590 nm wavelength best for detecting purple colored pigments. The purple color formation is a function of the total redox reactions that occurred in the well as the microbes use a carbon source metabolicaly causing the reduction of the colorless form of the dye, 5-cyano-2,3,-ditolyl tetrazolium chloride (CTC), to a purple- colored product.
The drawer will open when the readings are completed. Remove your plate and CLOSE the drawer using the DRAWER button on the face of the spectrophotometer. It is important to keep the drawer closed as much as possible to prevent dust from entering the internal parts of the machine.
A new screen will appear with your recorded data in a 96 well template.
The data should be saved automatically in softmax pro, but please also save to our BISC209 folder on the desktop, labeling clearly with your group letter and the date. Because the instrument computer is not networked, you will have to Save the data to a FLASH DRIVE. Export the labeled data to the thumb drive provided by your lab instructor. If you can export as an excel doc, do so.
Close the Spectromax software. DON'T SAVE! (You have already saved and exported adequately.) Eject the Thumb drive hardware properly. Shut off the SpectraMax spectrophotometer unless you know another group is waiting. Take the Flash drive over to the Instructor’s Mac computer and insert the flashdrive. Process the data. Save the daily data file to a folder for your soil sampling site on the Instructor’s computer desktop AND then EDIT: PASTE SPECIAL: TEXT the data carefully to the Google spreadsheet for your group found in SAKAI. The file is called: BIOLOG-CMD_calc&GraphGroup__2013fa. Each group has its own google doc linked in the navigation bar in SAKAI. Each day that you collect data has a separate page in the google doc. Notice that you will copy and paste one column in the spectrophotometer 96 well format at time into the google spreadsheet. After pasting column 1, 2 and 3 you will begin the next set of replicates (columns 4-6 then 7-9).
This is a very important step because it aligns the replicate cells properly.
When you have finished, drag the FLASHDRIVE icon to the TRASH and wait for it to disappear from the desktop. Please store the flash drive in the USB port in the computer on the instructor desk for all BISC209 students. If it is not there, send a message to the class to see if someone took it home by mistake and email your instructor.
PRECAUTION: If you haven’t saved your data and another group reads their plate, your data will be overwritten and lost.
DATA COLLECTION
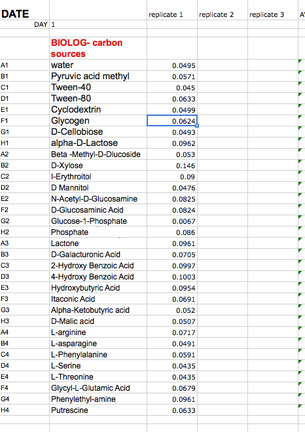
We have provided you with a spreadsheet called BIOLOG-CMD_calc&GraphTEMPLATE_2013_Tues_siteA (or B, or C, and _Wed_siteD, or E, or F)
You will find this spreadsheet on a thumb drive. Please follow your instructor's adivice about how to insert your daily data after each reading into this template.
You will have to carefully copy and paste the day's absorbance readings to the template. Be sure to use the correct one for your team! It is crucially important that
you align the right carbon source to its value (don't forget water) and get the right well numbers transferred to the appropriate position on the
Worksheet DAY (1, 2,etc) in the template workbook. Note that the Template contains a separate worksheet (tabs on the bottom) for each day within the full workbook.
Make sure that you fill out the appropriate day (DAY 0, 1, 2 etc). If you miss a day, leave that page blank.
The Google Doc is pre-formatted with the calculations for these data. It includes the formulas to average replicate measurements each day and it will automatically subtract the background (readings in the water wells). There is a normalization for background that will be subtracted automatically (this threshold absorbance is 0.25 for each carbon source). We hope that including these pre-made calculations in the template will make your calculation of community metabolic diversity (CMD) and the data analysis of carbon source utilization patterns relatively uncomplicated. Please make sure you understand the EXCEL performed calculations. When you are sure that your data has been copied correctly into the appropriate template day, you will notice that the built in calculations will provide a final average absorbance reading for each substrate.
Fill in the last column labled: SCORE 1 for wells with OD above 0.0. Once you fill in this final column, the template will calculate the total number of positive wells out of 31 for you and the SUM will appear in a cell below your data. This is the CMD for that day. The total will also appear on the last tab of the workbook (CMDtable&graph).
GRAPHING THE CMD DATA:
The graph of our community metabolic diversity(CMD) is built into the last page of the Workbook template: You will observe this figure for the calculated soil sample's
CMD values on the y axis versus time on the x. It provide a sense of the time frame for the community to begin using the carbon sources and by day 7 how many
different sources out of 31 were used: The community functional metabolic richness in carbon source utilization. What is important about this graph in providing
evidence for our hypothesis that a soil microbial community must be able to utilize a wide variety of carbon sources to support and maintain its abundance?
What else can we learn from assessing Community Metabolic diversity (CMD)
CMD is calculated by summing the number of positive responses (wells with a positive A595nm value after all the corrections) at each incubation time.
CMD is a simple way to represent the total number of substrates able to be effectively metabolized by the microbial community. It's a measure of diversity in use of
carbon sources but it does not identify the carbon substrates or help us find a pattern of preferred substrates.
Carbon source utilization pattern:
Since CMD analysis does not provide specific information about the pattern of carbon substrates used in your soil community, how could you examine the pattern of use of carbon sources?