BISC110: Lab 6 Taster SNP2
Lab 6 -Taster SNP Week 2
Today you will finish determining your genotype at the PTC gene locus by testing for one of the SNPs in the PTC gene. Remember that last week you began the process by isolating your own DNA and setting up a PCR reaction to amplify a region of the PTC gene which contains the SNP of interest. Today you will digest your PCR product with the restriction enzyme Fnu4H1 and analyze the products by agarose gel electrophoresis.
PART I: Digestion of your PCR products with the restriction enzyme Fnu4H1
Now that you have plenty of copies of the region of DNA we’re interested in, you will set up a restriction enzyme digest. How will this tell you what SNP you have? Restriction enzymes are quite specific about what DNA sequences they will cut. If the sequence is different, even by one base, then they will not cut. We have made use of this fact by choosing a restriction enzyme, Fnu4H1, which will cuts one variant of the PTC gene SNP, but not the other. Because the restriction digest will take an hour, you will set it up at the beginning of the lab period.
1. Locate your PCR tube from last week, and set it out at your bench to thaw.
2. Each person at your lab bench will set up a restriction enzyme digest with their own PCR product.
3. Get a clean microfuge tube and label it with your name or initials, and some indication that you are digesting the DNA in the tube.
4. Pipette 10 μl of your PCR product into your microfuge tube. Be sure to save remainder of your PCR product to run uncut on your gel.
5. Add 10 μl of the restriction enzyme “cocktail” which contains Fnu4H1 and the appropriate buffer. Mix well by pipetting up and down.
6. Incubate your tubes in the 37C incubator for 60 minutes.
DO NOT THROW AWAY YOUR PCR TUBE!
You will run uncut DNA on your gel along with the product of your restriction enzyme digest.
PART II: Agarose gel electrophoresis.
An animation of how to pour and run an agarose gel is available at: [| http://www.wellesley.edu/Biology/Concepts/Html/gel.html]
You can do steps 1 -4 while you are waiting on your restriction digests.
1. Make an agarose solution:
Each pair of students should make a 30 ml solution of 1.5% (w/v) agarose in 1X TBE (89 mM Tris-Borate, 2 mM EDTA) buffer.
a. Calculate the amount of agarose you should weigh out to make your solution.
for 30 ml 1.5% agarose:
weigh out _____ g of agarose
b. You will be provided with a 10X TBE stock solution. You will need 300ml of buffer for preparing your agarose solution and running your gel. Calculate the amount of 10X TBE you will need to prepare 300ml of 1X TBE.
measure out _____ ml of 10X TBE
add_____ml of water
c. Check your calculations with your lab instructor before proceeding.
d. Use a stir bar to mix your 1X TBE solution in a 500ml flask.
e. Using one of the balances in lab, carefully weigh out the appropriate amount of agarose. Transfer the agarose to a 125 ml Erlenmeyer Flask and add the proper amount of 1X TBE buffer.
It will not go into solution until the next step.
f. Microwave your agarose solution, checking the flask often to prevent boiling over. Use the “hot hands” to handle the flasks, since the glass will get very hot.
The agarose should be completely melted and the solution should be uniformly clear when you swirl it around.
g. Take your flask back to your bench and let it cool until you can comfortably touch the glass flask.
If you pour the agarose while it is too hot, you can crack or warp the casting tray, which may cause leaking.
2. Add SYBR Safe DNA stain to your agarose:
a. After your flask of agarose has cooled enough that you can touch it near the top comfortably, it is time to PUT ON GLOVES.
Any time you work with your agarose or gel for the remainder of the lab, you will need to be wearing gloves.
b. Ask your lab instructor to add 3 μl of SYBR Safe stain to your flask. Wearing gloves, swirl the flask gently to mix, taking care to avoid the formation of bubbles. Then proceed to the next step.
3. Cast a gel:
We will be using a Biorad gel system. Each pair of students will use one of these systems.
a. Place the gel-casting tray in the gel box on the raised platform, positioning the comb slots nearest to the negative electrode (Black).
b. Place the metal wedges into the gel box in the slots on either side of the gel-casting tray.
c. Place the comb into the slots closest to the edge of the gel-casting tray.
d. Using a Pasteur pipette, run a thin line of molten agarose along the seam between the metal wedges and the gel-casting tray.
Allow 1 min. for the agarose to harden. This will seal the seam and prevent leakage.
e. Carefully pour the remaining agarose into the gel-casting tray, trying to avoid the formation of air bubbles.
If bubbles form, remove them with a Pasteur pipette.
f. Allow the agarose to set completely. When it has solidified, it will appear relatively opaque.
g. Carefully remove the metal wedges from the gel box.
h. Fill both sides of the gel box with 1xTBE running buffer, until the buffer is about 3mm above the gel surface.
i. To remove the comb out of the gel, pull it slowly, and straight up, so that the wells are not damaged.
4. While you are waiting for your restriction digests to finish and your gel to harden:
a. Prepare your undigested samples.
i. Label a microfuge tube with your initials and indicate that it contains undigested DNA.
ii. Pipette 10 μl of your PCR product from your PCR tube into the microfuge tube.
If you do not have 10 μl left, add as much of your PCR product as you have.
iii. Add 10 μl of water to the microfuge tube. Mix thoroughly by pipetting up and down.
iv. Add 2.2 μl of loading dye to the microfuge tube. Mix thoroughly by pipetting up and down.
Don’t forget to use a clean, new tip for each tube.
Loading dye contains the following components:
Bromophenol blue
This is a negatively charged dye that move towards the positive pole..
Ficoll
Ficoll is added to increase the density of the sample.
This causes the sample to sink into the well, since it is heavier than the liquid in the well.
EDTA
EDTA helps keep the nucleic acid in the sample stable by inhibiting nucleases that might be present.
It does this by binding divalent cations, such as Mg2+, that are necessary for nuclease activity.
Do not throw the stock tube of loading dye away when you clean up after this lab!
b.Your lab group should acquire a sample of marker lane DNA (ready to load) from your lab instructor .
This marker lane DNA contains DNA fragments of the following sizes: 766 bp, 500 bp, 300 bp, 150 bp and 50 bp. Using what you know about the SNP region we amplified with PCR and the location of the Fnu4H1 II cutting site, you should be able to predict where you might see bands with respect to the marker lane bands.
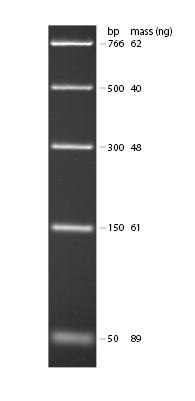
DNA Ladders: 0.3µg of “PCR Ladder” and visualized by ethidium bromide staining on 1.8% TBE agarose gel.
c.Your lab group should acquire a sample of control DNA (ready to load) from your lab instructor.
This is a sample of heterozygote DNA that has been digested with Fnu4H1 and mixed with loading dye."
d. Practice gel loading.
Loading an agarose gel can be tricky. Your lab instructor will demonstrate how to use the practice dishes to learn to load a gel properly. Practice until you are comfortable and proficient.
5. Prepare your restriction digest for loading:
a. After your restriction enzyme digest is complete, microfuge your tubes briefly (15 seconds) to get any condensation from the lid into the bottom of the tube.
b. Add 2.2 μl of loading dye to each tube of digested DNA (one tube for each member of your lab group, plus the tube containing your lab instructor’s sample).
Each pair of students should now have 6 tubes which contain DNA and loading dye: one digested tube for each group member, one undigested tube for each group member, one control DNA sample, and one tube of marker lane DNA.
6. Loading and Running the Gel
a. Load the contents of each tube into the gel. Be careful not to let any bubbles into the pipette tip when you are drawing the sample into the tip.
Check your tubes before loading: microfuge briefly (around 15 seconds) if necessary. In your notebook, make a record of which sample is loaded into which well.
b. When all of the samples have been loaded, check the orientation of the gel with respect to the electrical leads.
Remember “RED AHEAD”; DNA, having a net negative charge, will migrate toward the positive (red) electrode.
c. Placing the lid on the box connects the leads to the gel box. Connect the other ends of the leads to the power supply. Check that the power supply is set at 100 volts, and then turn on the "juice". Observe bubbles rising from the electrodes at each end of the tank (more from the negative electrode than the positive electrode). After 1–2min, observe the dye leaving the wells. Run the gels until the Bromophenol blue tracking dye is ? cm from the bottom.
7. Gel photography
a. When the gel finishes running, disconnect the leads from the power supply and remove the lid from the gel rig.
b. Wearing gloves, carefully remove your casting tray (still containing the gel) from the gel rig and transfer it to a labeled plastic sandwich box at your bench.
c. After the electrophoresis is complete, your instructor will take you to the instrument room next door, so that you may take pictures of your gels.
SYBR SafeTM stain is a fluorescent dye that binds to nucleic acids. It has excitation maxima at 280nm and 502nm, and an emission maximum at 530nm. You will place your gel on a UV transilluminator, a device that shines ultraviolet light through the gel. Where nucleic acids have been stained with the SYBR SafeTM stain, a fluorescent green glow will be detected by the digital camera connected to the transilluminator.
d. Once you have captured a satisfactory image of your gel, send it to your FirstClass® account, as well as posting it on your lab conference.
8. Analyzing the Gel
a. Ascertain whether your enzymes cut your DNA.
How can you tell? How many fragments do you see for each person in your lab group?
b. Use the DNA ladder to estimate the size of the fragments you see.
The migration of linear DNA molecules on the gel is proportional to the inverse of the log of the molecular weight.
c. You will now perform a quantitative assessment of the size of each fragment.
You will determine the size of your DNA fragments using a standard curve generated from the migration pattern of your DNA ladder fragments.
i. Measure the distance in cm each DNA fragment in the DNA ladder migrated on the gel. Record your data in your notebook.
ii. Graph the size of each fragment (bp) vs. the distance migrated (cm). Perform a linear regression of your graph. Appendix E has directions for making a linear regression line in Excel.
iii. Measure the distance each of your experimental fragments migrated on the gel.
iv. Plug in the values into your linear regression equation.
Do the values agree with your predicted values?
9. Recording Your Data
Determine what genotype you are: TT, Tt, or tt. Be sure to check your interpretation of the gel with your lab instructor. Once you are sure of your genotype, record your data in the class excel spread sheet where you have already recorded your PTC sensitivity phenotype.
Class results will be compiled and presented to protect anonymity. You are not required to share your personal data with anyone else in lab (though you are free to do so if you wish).
Part III. DTC Testing Group Work
While your gel is running your lab group will discuss the following questions and prepare a brief Powerpoint presentation for next weeks lab.
1. What service does your DTC company provide?
2. Does your DTC company meet the AJHG and ACMG recommendations for DTC testing? For a list of these recommendations consult the following:
3. Do you think your company provides a valuable service?
Assignment
1. Prepare a formal figure of the gel picture you obtained today, complete with title, legend and labels. For help, see the Resources section of this wiki: Guidelines to Scientific Writing.
2. Each lab group will prepare an Oral Presentation on their DTC company that addresses the questions your discussed in Part III. For help, see the Resources section of this wiki: Tips for Oral Preparations