BIOL368/F14:Chloe Jones Week 4
Exploring HIV Evolution In-Class Activity
Methods
Activity 1/Part 2: GenBank
Using the Genbank database, sequences were obtained from one of the fifteen subjects from the previously studied Markham et al. (1998) HIV-1 article .The sequence was chosen from a particular subject at a recorded visit, with a code for example that would read S4V2-4, signaling subject 4, visit 2. After a subject was selected the Genbank database allowed the viewing of the full record and the FASTA formatted sequence, which would be saved to a word document for further analysis in the proceeding activities.
Questions
- What was the accession number of the sequence you chose?
- ACCESSION: AF089142
- Which subject of the study was that HIV sequence from? Which section of the record contains information about who the HIV was collected from?
- From subject 3, in the definition portion
Activity 1 // Part 3: Introduction to Biology Workbench
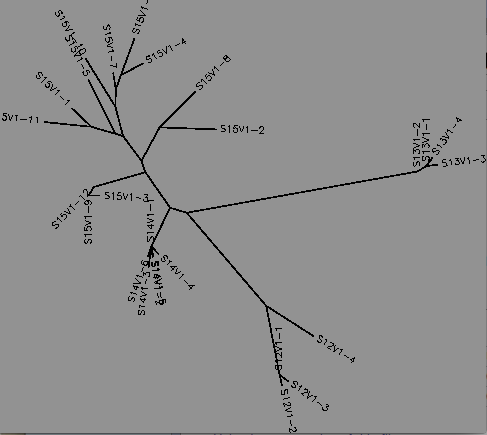
Based on random selection of 5 subjects sequence information obtained from activity 1/part 2, the FASTA formatted sequence would be be analyzed using the ClustalW tool which runs a multiple sequence alignment. The output is an unrooted tree the allows for pairwise similarity scores to be observed.
Activity 2 // Part 1: Quantifying Diversity Within and Between Subjects
Files labeled “Visit_1_Subjects_1_thru_9_HIV.txt” and “Visit_1_Subjects_10_thru_15_HIV.txt” were downloaded and uploaded into Biology Workbench. The files contained 97 sequences, and the program only allows 64, so they had to be uploaded separately. Multiple sequence alignment and the tree distance relationships were analyzed using the ClustalW tool, based on 12 sequences (3 clones from each of the 4 subjects),
Activity 2 // Part 2: Quantifying Diversity Within and Between Subjects
In the sections sequence similarity and differences were quantified using all the clones from one subject and aligning them. A table was then constructed that took into account the S’ values, theta, min difference, and max difference for each subject. The s’ value is determined by counting the number of positions where there is at least on nucleotide difference. The maximum and minimum values were calculated by generating a matrix using the Clustdist tool. The highest and lowest pairwise scores were chosen and then multiplied by the length of the sequence. Next, the subjects were compared using the pairwise distance matrix, which allowed for the calculation of the minimum and maximum differences across the subjects.
Table 1. Clustadist analysis of the sequences from Visit 1 of subjects 12,13, and 14, (S12V1, S13V1, S14V1) and the difference amongst their clones.
| Subject | Number of Clones | S | Theta | Min Difference | Max Difference |
|---|---|---|---|---|---|
| 14 | 6 | 37 | 16.2 | 86.4 | 3225.6 |
| 13 | 4 | 3 | .61 | 114 | 199.5 |
| 12 | 4 | 13 | .14 | 114 | 1311 |
Table 2.Clustadist Analysis comparing distance matrix for the alignment between subject pairs.
| Pairs of subjects being compared | Min Difference | Max Difference |
|---|---|---|
| 14 & 12 | 87.3 | 5063.4 |
| 14 & 13 | 86.4 | 4435.2 |
| 13 & 12 | 114 | 4389 |
Conclusion
In this analysis data was taken from the Markham et. al paper “Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. ”. From the fifteen subjects, four were studied closely based on their nucleic acid data and the relatedness of their clones. Using the Genbank database it was able to generate full records and the FASTA formatted sequences for the different subjects. Biology Workbench had various tools to create unrooted trees in order to study diversity amongst the different subjects clones. The subjects I chose to study were Subject 12, 13, 14 and 15. Subject 15 had double the amount of clones in comparison to the other subjects. The unrooted tree for subject 15 showed a lot of diversity based on unrooted tree having multiple nodes and branches. Also, subject 15 displayed variable distance among the clones that were present, signifying that their clones were not as closely related as in comparison to the other subjects. Subjects 12, 13 and 14 had clones that were similar to one another, and thus were more clustered. From observing this data and taking into consideration the Markham article conclusion that diversity of clones signifies a more active form of the HIV virus, I would consider subject 15 to be a representation of a rapid progressor, which correlated to the group placement in the article.. Subjects 12, 13, and 14 were further analyzed by using the Clustadist Tool to specifically measure data values that would give more insight to the similarities and differences of the clones. Subject 14 had the highest theta and s values which would show there was more of a difference amongst the clones; essentially greater diversity. The minimum and maximum differences look at extremes in terms of similarity and difference. Again , subject 14 had the highest maximum difference, and the branches were not as clustered in comparison to subject 12 & 13. Subject 14 according to the data I analyzed and that of the article falls into the category of a moderate progressor. Subject 12 & 13 were highly clustered with little distance between the pairs of sequence, suggesting that they are non-progressors which again was simultaneous to the Markam et al article. When comparing the subjects 12, 13, and 14 to one another the subject were the most part all similar to one another, with the lowest minimum difference being between subject 14 & 13 and the lowest maximum difference between subject 12 & 13.
Defining Your Research Project
- What is your question?
- Do subjects with similar clones that share the same place on the phylogenic tree such as in subject 1 & 2, at what rate (what visit do they become) start to have more dramatic affect in terms of diversity and divergence. Particularly, want to study Subjects 1 &2 because overlap on phylogenetic tree, but one subject is a rapid progressor and one is a non-progressor. I want to analyze as the years past did the clones progress at the same rate since they started off at visit one at the same position
- Make a prediction (hypothesis) about the answer to your question before you begin your analysis.
- I don’t think they will be as similar as time passes and data is collected from the following visits because other clones in the data pool form each subject will have an effect on the progression of the clone.
- Which subjects, visits, and clones will you use to answer your question?
- I chose subject 1 & 2 to analyze because of how phylogentically closely they are related based on the first visit. I will be studying all of the visits that were gathered to see at what point did the clones start to diverge or if they remain at the same location or they diverge seperatedly or together. I am specifically going to keep a close eye on clone 1 & 2 for subject 2, and clone 3 for subject 1 because they essentially are the same thing by sitting at the same place on the phylogenetic tree.