BE.180:Exam2reviewQ
Here are a few simple questions that should remind you of the four sections of our course:
- Python programming
- DNA programming
- DNA->mRNA->Protein coding
- Spatial programming
Q1. Consider the following two simple Python code examples:
Case A.
def multiply(x,y):
___prod=x*y
___return prod
print multiply(2,3.14159265)
print multiply(4,'MIT')
Case B.
y=2*3.14159265
print y
z=4*'MIT'
print z
Which is better? Why?
A1. Case A is better because a reusable function, multiply, has been defined that can be called many times other as needed (i.e., in different programs).
Q2. What is the difference between an engineered biological part and an engineered biological device?
A2. A part is a basic biological function that can be encoded as genetic material (e.g., a open reading frame encoding a specific protein). A device is a combination of one or more biological parts that performs a human-defined function (e.g., a genetically encoded inverter). Devices are made from parts. Functions encoded by parts are defined by the underlying biology. Functions encoded by devices are defined by the people making the devices.
Q3. How many different codons encode the amino acid methionine? What about leucine?
[please note, I'm not expecting that you have memorized the standard genetic code, rather that you appreciate and could exploit the facts of the answer to this question].
A3. In the standard genetic code, six different codons can specify the amino acid leucine (UUA, UUG, CUU, CUC, CUA, CUG) whereas only a single AUG codon specifies methionine.
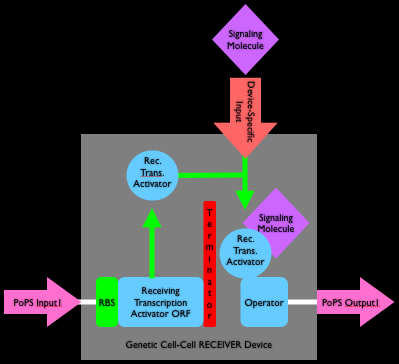
Q4. The luxR open reading frame encodes a transcription factor that activates gene expression in the presence of the cell-cell signaling molecule, AHL. Using other biological parts of your choosing (e.g,. ribosome binding sites (RBS), transcription terminators, promoters, et cetera), draw a parts-level device diagram for a genetically-encoded receiver device that accepts AHL as an input signal and produces a generic PoPS signal as output. On your sketch, be sure to include the device boundaries and label the input and output signals. Finally, how would cells containing this device respond across a spatial gradient of AHL?
A4.