Arking:JCAOligoTutorial7
Common Pitfalls, Tricks and Tips
Using contaminated media
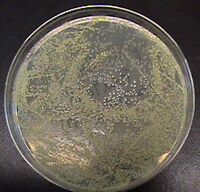
A common error is rescuing transformations in contaminated media. The plate at right shows two colony morphologies. The minor population are white E. coli colonies, the majority are yellowish. They came from a bottle of 2YT media that had visible cloudy streaks. Before you use growth media for picking colonies or rescuing a transformation, check to make sure the media has no cloudiness or floating debris. You can't rescue contaminated media, so throw it out. Along these lines, it is important when making media that you autoclave it without significant delay. The powder in the media bottles is not sterile, and will readily grow as soon as you add water. This growth will alter the composition of the media. So, it is important that you kill that stuff by autoclaving before it gets a chance to amplify. Waiting 1 or 2 hours to autoclave is ok. Waiting overnight is fatal.
Storage methods for E. coli
There are many ways to store E. coli including liquid cultures, agar plates, agar stabs, agar slants, and -80 cultures. It is important to know how long these things stay viable. Liquid cultures growing at 37 degrees are viable for approximately 48 hours. After this time (at 37), the bacteria will be mostly dead, and even if they aren't dead will likely have lost plasmids. If you take liquid cultures out of the incubator and place them on the bench at room temp, they will be reasonably stable for a few more days. If you put a saturated liquid culture in the refrigerator, it will last at least a week. Never directly freeze media at -20 or -80 without adding a cryoprotectant such as glycerol or DMSO. They are not stable without it. However, if you do add DMSO (~10%) or glycerol (8 to 25%), cultures of E. coli in pretty much any media (M9, LB, SOB, PBS buffer, etc.) coming from any type of growth (any growth phase of liquid growth, scraped off a plate, pelleted from liquid cultures and resuspended in media) are stable for years stored at -80. Many people will flash freeze these stocks before placing at the -80, but it is perfectly ok to simply drop the samples into the -80 and let them slow-freeze. Agar plates are generally stable stored at 4 degrees as long as they are preventing from drying and contamination by wrapping in parafilm. They can last as long as 2 months if you're lucky. Agar stabs and agar slants involve storing bacteria on the surface or embedded within agar in sealed tubes. They are stored at room temperature and are stable as long the agar does not dry out. They can last years but are generally not as stable as -80 stocks.
Finding oligos from your oligo inventory
Let's say you need a sequencing oligo that anneals within your gene of interest, and you know you've done this before, but you don't remember the oligo name. Assuming you've maintained an "all oligo" style spreadsheet, you're in luck. *Go to the pairwise alignment tool at [1].
- paste the gene of interest sequence into the first box
- Highlight the column of sequences in the all oligo spreadsheet, copy it, and paste in the second box
- hit submit
This will only give you one good forward and one good reverse oligo at a time, but you can iteratively go through the list by deleting each one you find.
Analysis of microarray data
So, say you want to find a promoter from E. coli that turns on under some set of conditions. Microarrays are a great way to do that. You should be aware that there are multiple types of microarray, but the most common two are spotted arrays and affymetrix arrays. For all types, you prepare 2 RNA samples: one of the experimental condition, and one of a reference condition. You then do some in vitro enzymatic manipulations to convert the RNA into DNA and add dyes. For spotted arrays, you mix the two samples and do a hybridization to the chip. For Affymetrix arrays, you run two chips. In either case, you'll get lots of data out about how much RNA was present for each gene in E. coli. If you rank the ratios of the two values, you'll hopefully have some genes that induced strongly and others that perhaps repressed strongly under your experimental conditions. You will then want to read up on those genes. Pubmed papers about lacZ or GFP reporters are often available. There are also some good reference database:
Ecocyc provides an encylopedia-like database about the function of E. coli genes and their regulation. If search for the gene names, it will tell you about what the gene does, if it is metabolic it can tell you what pathways its involved in, it shows known operator sites in its promoter, and it links to many papers.
The University of Oklahoma's E. coli Gene Expression Database is my personal favorite. It has a database of microarray experiments, and you can type in several gene names and it compares whether they are induced or repressed under a wide set of conditions. From this, you can tell if a gene is responding to many environmental stimuli, or if the response you are observing is pretty specific.