Arking:JCAOligoTutoria20
Translational Fusions
One of the primary motivations in developing the BglBricks standard was to facilitate the construction of fusion proteins. The underlying principle here is that proteins tend to fold in modular units called "domains". Some proteins are composed of a single domain, while others contain multiple domains. These folding units are often also functional units, and it often it is possible to recombine two such functional units from different sources into a single molecule and preserve the activity of both functional units. To explain this further, let's take a look at an example. Shown below is the crystal structure of an IgG protein and below that is a simplified representation of the IgG protein. IgG proteins are one of several forms of antibodies that are made within your body as key components of the immune system.
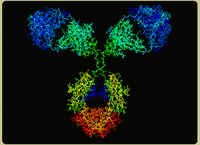
Image from www.physics.purdue.edu/people/faculty/nolte.shtml
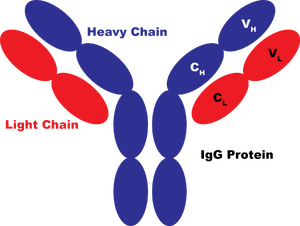
In the diagram above, the IgG protein is colored based on its polypeptide chains. There are 4 polypeptides in the single IgG protein: two heavy chains (in blue) and two light chains (in red). Each oval in the diagram represents a single folding domain. All the domains that make up antibody proteins have a type of fold called the immunoglobulin fold, and these domains are referred to as immunoglobulin domains. All the information to make a properly-folded polypeptide is encoded within each immunoglobulin domain. So, if you constructed a coding sequence (CDS) for a single immunoglobulin domain and provided an acceptable promoter, ribosome binding site, and terminator in the right places, you could express a properly-folded immunoglobulin domain within a cell. Now, that would be a properly-folded domain--not necessarily a fully-functional antibody. In fact, one immunoglobulin domain itself would be pretty useless. To understand this, let's first examine what IgG proteins do. Primarily, they bind to a particular "epitope" which is usually a short peptide often from a virus. Additional proteins within the immune system also interact with the IgG protein and induce responses that result in the destruction of the bound species.
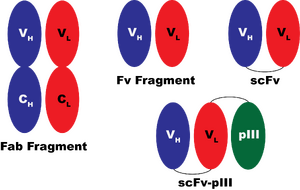
So, the IgG molecule is able to do lots of things in the context of your bloodstream. The most basic of these activities is its ability to bind to its epitope. The epitope is bound in the cleft between the VH and VL (V="variable") domains of the IgG. In practice, the minimal component of the IgG protein that retains the binding ability is called the Fab fragment which contains 2 polypeptides, each of 2 domains (two variable domains and two "constant" domains, CH and CL). One might expect that just the VH and VL domains together might be sufficient to get the binding activity. In practice, the affinity of VH and VL for one another is fairly low, and expressing these two domains as separate proteins is usually not sufficient to obtain a functional binding protein. However, it is possible to construct a fusion protein, referred to as a single chain antibody, or "scFv" and get a single polypeptide that retains all the binding activity of the original IgG molecule.
Even more interesting, one can fuse other polypeptides that have nothing whatsoever to do with the immune system to the scFv molecule and obtain a functional polypeptide that is the sum of the scFv protein's activity and the activity of the protein fused to it. So, for example, an enzyme can be fused to the scFv and the product would both bind to the epitope and also perform the chemical reaction of the enzyme. One of the most common things fused to scFv proteins is the pIII protein from M13 phage. The pIII protein lies at the tip of the M13 phage particle. By fusing the scFv to pIII, it is possible to generate M13 phage particles that display the scFv on their surface. This technology has been used extensively to engineer antibodies that bind to altered substrates and is called "phage display".
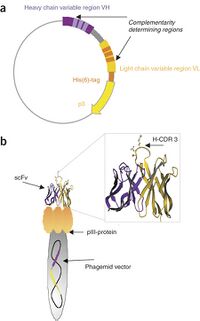
Now, constructing a fusion protein is not truly an act of "gluing two proteins together". In reality, what you are doing is constructing a DNA encoding the fusion protein. To explain this, check out the graphic at right that illustrates how phage display looks at the level of the DNA in (a) and on a phage particle in (b). In phage display, the fusion protein is encoded within a plasmid. On the 5' end of the fusion protein gene is the VH coding sequence, and then the VL coding sequence, and finally the pIII coding sequence. In between each domain is a sequence referred to as the "linker".
Linkers are the unpredictable component of fusion proteins. Sometimes they are arbitrary, sometimes they are really important. When constructing fusion proteins, it is often necessary to make multiple variants with different linker lengths and compositions to get the desired properly-folded and functional product. If it is too short you can get interference between the folding of the two domains, or steric clashes between them. If it is too long you can get proteolytic cleave of the domains. So, linkers can get tricky.
The other bit of uncertainly is whether a polypeptide will permit fusions to its N or C terminus. Often the ability of a protein to accept fusions is predicatable from the crystal structure. Solvent-exposed and flexible termini generally permit fusion. Termini that a recessed or components of active sites generally will be disrupted upon fusion. Those principles are by no means universal, and the best evidence is always empirical evidence--an example of someone fusing another sequence to the terminus of your domain of interest.
BglBrick Assembly of Fusion Protein Parts
One of the key motivations in developing the BglBricks standard for assembly was to enable the encoding of fusion proteins as composite parts of individual protein domain basic parts. The product of BglBrick assembly is the sequence GGATCT which encodes Gly-Ser which is a very commonly-used minimal linker sequence for constructing translational fusions.
The key difference between fusion protein coding sequences and regular CDS basic parts is the lack of start or stop codons in these parts. Their existence complicates the repertoire of parts in our collection, so they require some care in bookkeeping and design. To facilitate this, we have defined a nomenclature for the different types of parts. Here is the full spectrum of different types of coding sequences. Greater-than and less-than signs represent parts that allow C-terminal or N-terminal fusions (or in essence, the coding sequence is in-frame with the BglBricks scar and lacks the start codon and/or stop codon).
{<part>} No start, no stop, in frame with BglII and BamHI sites
{<part!} No start, has stop, in frame with BglII site
{part>} Start codon directly after BglII site, no stop, in frame with BamHI site
{part!} Start codon directly after BglII site, and stop codon
As an example of this, let's consider the His-tag family of parts. The peptide sequence HHHHHH, or His6, is often fused to the N or C terminus of a protein to allow for its rapid purification on Ni-NTA resin. Here is what such parts would like in DNA:
{<his6>} agatctCATCATCATCATCATCATggatcc
{<his6!} agatctCATCATCATCATCATCATTAAggatcc
{his6>} agatctATGCATCATCATCATCATCATggatcc
{his6!} agatctATGCATCATCATCATCATCATTAAggatcc
Protein Fusion Quiz
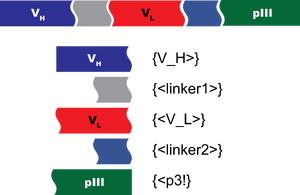
For your quiz, let's make BglBricks basic parts from which you can assemble a composite part encoding an scFv-pIII fusion protein. In the image above, the curvy lines represent the junctions between two domains, and the straight lines are the start and stop of the coding sequence. So, to make the scFv, you'll have a total of 5 basic parts as illustrated.
You are given some M13 phage genomic DNA and a plasmid encoding the heavy light chains for expressing a Fab fragment of an antibody in E. coli. You'll make the parts not encoded within these source DNAs using the methods you've learned for short parts.
Design oligos and write up 5 construction files to make the 5 basic parts you'll need for this project. You can put all 5 construction files into a single text file. Use EcoRI and BamHI to construct your parts in plasmid pBca9145.
For the first linker, make the part:
tacttcgatgtctggggcgcagggaccacggtcaccgtctcctcaggcggaggcggttcaggtggaggtggttctggaggagga
For the second linker, make the part:
tggacgttcggtggaggcaccaagctggaaatcaaacgtgcggccgctcaccaccatcaccatcacggcattgaaggccgt
You can find the sequence of the pIII protein at pubmed LOCUS AB158263. You can find the sequences of VH and VL in this file: Media:FileAJ852004.seq. I've annotated the VH and VL regions of the heavy and light chains in yellow.
...and as always, go through your construction files in ApE to check your work! In checking your answers, make sure your start codons, stop codons, frames, and silent mutations are all designed properly.