AMPure Mods
Our group uses AMPure beads to clean Illumina DNA-seq and RNA-seq libraries, as there are many advantages to using SPRI beads over other products (see Quail et al., 2009). The biggest advantage is that conditions can be modified to selectively 'lose' small DNAs in the size range of 50 - 200 bp, as is shown in this image
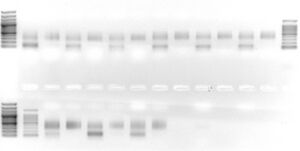
. This can be advantageous if Illumina libraries contain adapter dimers or other contaminating oligonucleotides. While we use AMPure beads in our laboratory, this is not an endorsement of AMPure per se; similar results can be obtained using other paramagnetic SPRI beads.
General Guidelines for AMPure-bead based isolation
• Illumina DNA-Seq libraries: Use 0.7 volumes AMPure to 1.0 volume sample. Eliminates 100bp and 200bp bands, retains > 300bp bands
• Illumina RNA-Seq libraries: Use 0.9 volumes AMPure to 1.0 volume sample. Eliminates 100bp band, retains > 200bp bands
• Routine DNA and PCR cleaning: Use 1.5 volumes AMPure to 1.0 volume sample. Eliminates primers and small oligos
Example volumes for DNA-seq and RNA-seq libraries
• for DNA seq
AMPure : Sample+Water = 45 µl AMPure : 65 µl Sample+Water
• for RNA seq
AMPure : Sample+Water = 45 µl AMPure : 55 µl Sample+Water
Abbreviated protocol for AMPure bead purification
• Prior to use, remove the beads from storage and let them stand for 30 minutes to warm to room temperature.
• Just before use, vortex or shake the AMPure bottle to resuspend magnetic particles that have settled. The suspension should appear completely homogenous.
• Add the desired volume of Agencourt AMPure beads. Mix the solution ten times by pipetting up and down.
• Incubate samples + water + AMPure at room temperature for 15 minutes for maximum recovery (5 minutes for decent yield).
• Magnetize and then incubate for at least 5 minutes (more time may be required for viscous solutions).
• Aspirate the cleared supernatant SLOWLY and discard. You can leave up to 5 ul of supernatant remaining in the tube at this step.
• Dispense 200 µl of 80% EtOH (freshly prepared every time) into each tube and incubate for 30 seconds. Aspirate the EtOH wash and discard. Repeat for a total of two washes.
• Remove all traces of EtOH with a fine pipette tip, and place on bench until completely air dried. We recommend at least 10 minutes.
• Add the desired volume of elution buffer (10 mM Tris-HCl, pH 8.0 or water) and vortex for 15-30 seconds.
• Incubate the samples at room temperature for at least 2 minutes.
• Magnetize the sample one last time for 5 minutes.
• Aspirate the supernatant (the ELUTED DNA) and transfer to a new tube.
TIPS!!! Don’t let the beads go dry between wash steps. Don’t dry beads in the spin-vac or PCR machine; air-temp drying is preferred.
return to Cronn_Lab:Protocols page