User:Megan L. Channell/Notebook/Horseradish/2013/09/04: Difference between revisions
From OpenWetWare
(Autocreate 2013/09/04 Entry for User:Megan_L._Channell/Notebook/Horseradish) |
(fix raw html notebook nav) |
||
| (18 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
{|{{table}} width="800" | {|{{table}} width="800" | ||
|- | |- | ||
|style="background-color: #EEE"|[[Image: | |style="background-color: #EEE"|[[Image:BDLlogo_notext_lr.png|128px]]<span style="font-size:22px;"> Biomaterials Design Lab</span> | ||
|style="background-color: #F2F2F2" align="center"| | |style="background-color: #F2F2F2" align="center"|[[File:Report.png|frameless|link={{#sub:{{FULLPAGENAME}}|0|-11}}]][[{{#sub:{{FULLPAGENAME}}|0|-11}}|Main project page]]<br />{{#if:{{#lnpreventry:{{FULLPAGENAME}}}}|[[File:Resultset_previous.png|frameless|link={{#lnpreventry:{{FULLPAGENAME}}}}]][[{{#lnpreventry:{{FULLPAGENAME}}}}{{!}}Previous entry]] }}{{#if:{{#lnnextentry:{{FULLPAGENAME}}}}|[[{{#lnnextentry:{{FULLPAGENAME}}}}{{!}}Next entry]][[File:Resultset_next.png|frameless|link={{#lnnextentry:{{FULLPAGENAME}}}}]]}} | ||
|- | |- | ||
| colspan="2"| | | colspan="2"| | ||
<!-- ##### DO NOT edit above this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit above this line unless you know what you are doing. ##### -->==Adenosine and Inosine UV-Vis and Analysis== | ||
== | |||
==Objective== | |||
*Collect the data on the UV-Vis from the ionisine dilutions | |||
The inosine samples came from [[User:Megan_L._Channell/Notebook/Horseradish/2013/09/03 | yesterday's]] protocol | |||
*Calculate calibration curves for Adenosine and Inosine | |||
*Run a data regressions on the class' data | |||
==Protocol== | |||
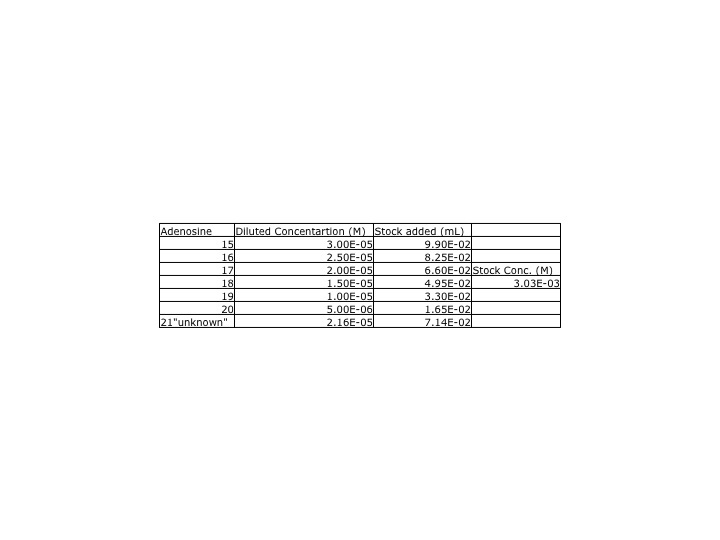
Yesterday, our adenosine dilutions were much lower than the class' average, so we redid the adenosine samples. A new stock solutions was made using [[User:Megan_L._Channell/Notebook/Horseradish/2013/09/03 | yesterday's]] protocol and the stock solutions was made from .0809g of adenosine. | |||
[[Image:Adenosine2 9 4 2013 cmj.jpg]] | |||
==Data== | |||
A UV-Vis spectra of the new adenosine samples and the inosine samples were collected. The parameters for the UV-Vis were measured between 450 and 200nm. The peak for the insonie spectra was at 249nm while the adenosine was 259nm. This wavelength was used when calculating the calibration curve. | |||
[[Image:Inosine 9 4 2013 cmj UVVIS.jpg]] | |||
[[Image:Inosine 9 4 2013 cmj cali.jpg]] | |||
[[Image:Adenonsinefixedtrial2 9 4 2013 cmj.png]] | |||
[[Image:Adenosinetrial2 9 4 2013 conc.jpg]] | |||
==Data== | |||
[[Image:Adenosine 9 4 2013 cmj trial2 grubbs.png]] | |||
[[Image:Inosine 9 4 2013 cmj grubbs.png|1050px]] | |||
*The Grubbs test was calculated using the formula | |||
G=|mean-x|÷std. dev. | |||
*The data that our group collected from 9/3/2013 was lower. We remade Adenosine concentrations from a new stock solution, but did not do that for inosine, thus our group (5) data is not recorded. | |||
<!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | ||
Latest revision as of 23:17, 26 September 2017
 Biomaterials Design Lab Biomaterials Design Lab
|
|
Adenosine and Inosine UV-Vis and AnalysisObjective
The inosine samples came from yesterday's protocol
ProtocolYesterday, our adenosine dilutions were much lower than the class' average, so we redid the adenosine samples. A new stock solutions was made using yesterday's protocol and the stock solutions was made from .0809g of adenosine. DataA UV-Vis spectra of the new adenosine samples and the inosine samples were collected. The parameters for the UV-Vis were measured between 450 and 200nm. The peak for the insonie spectra was at 249nm while the adenosine was 259nm. This wavelength was used when calculating the calibration curve.
Data
G=|mean-x|÷std. dev.
| |