3D Paper Microfluidics - Adaline Nuesse
Introduction
Paper-based microfluidics are an attractive technology due to their affordability and feasibility to be implemented in remote locations. In places where medical testing materials are limited and funds are low, a lightweight and inexpensive device is necessary. [1] Since 3D paper-based microfluidic analysis devices (µPADs) use simple fabrication techniques with everyday materials, medical technologies can be produced quickly, used easily without requiring intense training, and disposed of without much effort.[1,2]
The invention of 3D paper-based microfluidics can be attributed to the Whitesides group. The first design was created by attaching layers of paper together with double sided tape. The hydrophilic and hydrophobic nature of the channels and walls respectively, allowed for flow in the vertical and horizontal direction through a series of channels. The goal of the Whitesides group was to enhance the capability of paper- based microflidics while keeping the systems low cost, lightweight, and easy to use. [3]
Fabrication
3D µPADs are superior to 2D µPADs due to their ability to include multiple reactions or process steps in a compact space. Since there is now the ability to flow in the z direction, the process speed and efficiency can be increased in a single chip by increasing the number of flow paths and increasing the functions within the layers. Since 3D µPADs build upon the concepts of 2D µPADs, the fabrication techniques for 2D are enhanced for 3D purposes. The two main methods for 3D µPAD fabrication are stacking and origami. [4]
Stacking
The process of stacking can be implemented by creating multiple paper layers and then connecting them together through the use of double sided adhesive tape. The adhesive tape is lined up between each layer to connect the paper without blocking the passages for fluid flow. [4] The connecting tape is cut using a laser cutter to create channels that would be filled with cellulose powder to link the fluid routes creating multiple hydrophilic paths. The cellulose powder is necessary to fill the gap that was created by the tape to induce flow between the paper layers. [4,5] In this method of adhesive stacking it is found that the paper and tape can be difficult to align and therefore other stacking methods have been investigated. [4]
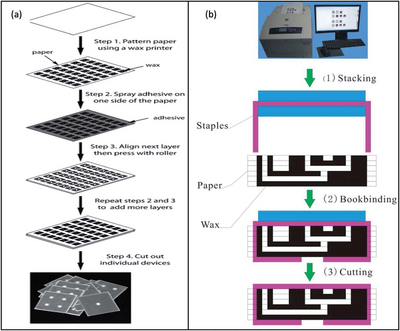
An alternative stacking method combined the technique of wax printing and an adhesive spray. In this method 2D µPADs are created using wax printing, but instead of aligning a tape to the material, an adhesive spray is layered onto the individual paper layers and pressed together. This removed the challenge of lining up the paper layers to the adhesive tape perfectly, since the spray was applied evenly and completely to the surface. [4]
Additionally, a method of stapling wax printed layers together was used to form 3D µPADs. This alternative, also called bookbinding, removed the need for an adhesive substance altogether. It was found that although this method did not align the layers perfectly, it was useful for mass fabrication of 3D µPADs. This is due to the quick mechanical process of connecting the paper layers. [4]
Origami
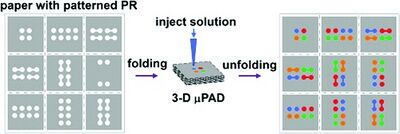
While stacking was a reasonable method for creating the 3D µPADs, origami method is beneficial because it does not need any adhesive at all. This removes any chance of additional contamination and need for laser cutting. [5] The layers have the desired patterns printed into the paper using photolithography with indicators for where the paper should be folded. The patterns control the flow of the fluid to avoid leaking, without additional energy needed. [6] The folds were held into place using a clamp or weight to keep the material in place. [4]
This technique is beneficial because it does not need additional materials like adhesive tape or cellulose powder. The contact between the channels is made only through folding and pressing of the paper. The origami folding methods can be completed in a short time span without needing extensive tools to hold it in place. Additionally, after the origami paper configuration has been used, it can be deconstructed and analyzed. The paths the different fluids traveled can be identified and analyzed for proper mixing and flow travel. [5]
Applications
The need for complex 3D microenvironments is growing rapidly. 3D µPADs allow for more assays to be analyzed within the same given area than would be on 2D µPADs. Additionally, the movement within three dimensions opens up more options for steps within a given device. There are more dimensions that occupy a smaller volume, allowing for less fluid in contact with the paper channels and therefore less material absorbed. This is an important advantage when dealing with biological samples that might be limiting in sample size. [2]
Cell Culture Analysis
3D µPADs are extremely useful when creating microdivices to simulate different situations for cell cultures. Due to paper-based devices being biocompatible, the cell cultures can thrive in these conditions, allowing for different small molecules to be targeted and analyzed. For example, small molecules like glucose and lactate can be characterized by their concentrations over varying conditions. [7]
The 3D µPADs allow multiple layers to be utilized. For cell cultures analyzing concentrations of glucose and lactate, three unique layers can be identified which each layer being separated by adhesive tape, a hydrophobic layer, and a channel. Alternative cell culturing techniques analyze the lung tumor cells to notice the metabolic activity that occurs during radiation treatment. By creating 3D µPADs to analyze the tumor cells in a micro simulation, the growth of these cells after radiation can be characterized over different time spans and conditions allowing for greater understanding of the cells behavior. [7]
Cell Migration
A 3D environment is useful for analyzing how cells would behave when disturbed. 3D µPADs are used to monitor the movement of cells within the three spatial dimensions, collecting more applicable findings than if analyzed in a 2D setting. This simulation is observed when analyzing breast carcinoma cells through the addition of epidermal growth factors (EGFs) and cancer- associated fibroblasts (CAFs). Through altering EGFs and CAFs in a varying chemical gradient the behavior of the original cells is motivated into observable directions. These gradients in conjunction with a microenvironment allows for a controlled area to observe how cells would potentially react in a human body. By allowing a 3D environment, the variables can be precisely controlled with known values to allow for confident results to be determined and applied in a larger scale situation.[8]
Malaria Diagnosis
In countries where malaria is a prominent cause of death, immediate diagnosis is necessary to receive treatment. Ordinarily, to identify whether a patient has malaria, a blood sample must go through a series of laboratory examinations and tests to obtain working results. Due to the locations where malaria is prominent, typical laboratory equipment and trained technicians are not available, so an affordable diagnosis alternative is desired. A multiple layered 3D device was created by Deraney to detect any combinations of malaria. This paper-based device was created for the sample to be inputed, stored, tested, separated, and analyzed. Once the samples qualities were determined, the device would display specific colors to inform the technician of the negative or positive results. This is a easy to use and cost effective method of malaria testing in developing countries. [9]
References
[1] de Oliveira, R. A., Camargo, F., Pesquero, N. C., & Faria, R. C. A simple method to produce 2D and 3D microfluidic paper-based analytical devices for clinical analysis. Anal. Chim. Acta, 2017, 957, 40-46. https://doi.org/10.1016/j.aca.2017.01.002
[2] Yetisen, A. K., Akram, M. S., & Lowe, C. R.Paper-based microfluidic point-of-care diagnostic devices. Lab Chip, 2013, 13(12), 2210-2251. https://doi.org/10.1039/C3LC50169H
[3] Martinez, A. W., Phillips, S. T., & Whitesides, G. M. Three-dimensional microfluidic devices fabricated in layered paper and tape. Proceedings of the National Academy of Sciences, 2008, 105(50), 19606-19611. DOI: 10.1073/pnas.0810903105
[4] He, Y., Wu, Y., Fu, J. Z., & Wu, W. B. Fabrication of paper-based microfluidic analysis devices: A review. Rsc Advances, 2015, 5(95), 78109-78127. https://doi.org/10.1039/C5RA09188H
[5] Liu, H., & Crooks, R. M. Three-dimensional paper microfluidic devices assembled using the principles of origami. J. Am. Chem. Soc., 2011, 133(44), 17564-17566. https://doi.org/10.1021/ja2071779
[6] Sechi, D., Greer, B., Johnson, J., & Hashemi, N. Three-dimensional paper-based microfluidic device for assays of protein and glucose in urine. Anal. Chem., 2013, 85(22), 10733-10737. https://doi.org/10.1021/ja2071779
[7] Tribhuwan Singh, A., Lantigua, D., Meka, A., Taing, S., Pandher, M., & Camci-Unal, G. Paper-based sensors: Emerging themes and applications. Sensors, 2018, 18, 2838. https://doi.org/10.3390/s18092838
[8] Campbell, J. M., Balhoff, J. B., Landwehr, G. M., Rahman, S. M., Vaithiyanathan, M., & Melvin, A. T. Microfluidic and paper-based devices for disease detection and diagnostic research. Int. J. Mol. Sci., 2018, 19(9), 2731. DOI: 10.3390/ijms19092731
[9] Li, H., & Steckl, A. J. Paper microfluidics for point-of-care blood-based analysis and diagnostics. Anal. Chem., 2018, 91(1), 352-371. https://doi.org/10.1021/acs.analchem.8b03636
