20.181/Lecture9
From OpenWetWare
Jump to navigationJump to search
Placing atoms a la HW6
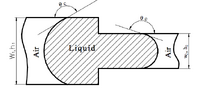
- We can put the first atom (green) anywhere; put it at the origin
- The second atom (blue) is placed a fixed distance away. It can be anywhere on the circle centered at the green dot, so we can choose to put it on the x-axis.
- The third atom (red) defines the angle a123 and so it must be placed on the cone. But we also know it is a certain distance from the blue atom, so that constrains it to the "base" of the cone, the cirle perpendicular to the xy plane. We can choose to put it at the intersection with the xy plane.
Protein design problem
- Given: protein backbone (a 3D structure)
- Find: amino acid sequence (1D)
Build 3D protein structure incorporating known chemistry, rotation angles, torsion
- the chemistry of an amino acid is the same no matter what protein its in
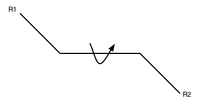
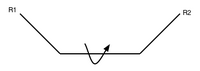
- so what's free to change is the stereochemistry
- Recall trans and cis conformations from ochem. (In these picture the middle bond is a double bond.)
- proteins are packed together like a jigsaw puzzle - you can rotate about that central bond
- protein design problem is NP-hard
Approach
At each residue:
- find a 'rotamer' (rotamer has two peices of information: what amino acid is there, and the conformation (if a residue has rotatable bonds, we'll b
- choose the set of rotamers at all positions that give the global energy minimum
- Why are we looking for the minimum? Recall low energy = stable.
- Call energy "E" but its actually a free energy
What makes proteins fold?
What are the interactions that make proteins fold ?
Entropy
- hydrophobic effect:
- primary interaction driving proteins to fold
- largely an entropic effect (delta S)
- entropy is also the largest barrier to protein folding
- Comformational entropy loss upon folding, an estimate:
"3 conformations/bond" estimate below comes from considering ethane:
- how many conformations does the bond have?
- basically 3 energy minima: the three different staggered orientations
- So an rough estimate for the loss of entropy for a folding protein:
100 residues * 2 backbone rotatable bonds per residnue * ~2 sidechain rotatable bonds ( per residue, on average) * 3 conformations/bond = 1200 conformations --> 1 conformation when folded entropy loss ~= 1200kT ~= 700 kcal/mol !!
- Most stable (known) protein on the planet: 13 kcal/mol
- Designed on a computer and built in David Baker's Lab.
- A stable protein in nature might be ~5kcal/mol.... Nature doesn't have any reason to make proteins any more stable than 5 kcal/mol (already 99.98% folded... good enough, don't have to be 99.99999995% folded)...
- If the loss of entropy is on the order of 700 kcal/mol and the stabilizing interactions are on the order of 705 kcal/mol ... we have to do some very careful bookkeeping to get those numbers right!
Algorithm
- build a lot of structures
- calculate energy of each
- pick the best one
Stunning Outdoor Interpretative bonding dance by TAs
you really had to be there