User:Alison D. Bunnen/Notebook/Biology 210 at AU
2/11/15 and 2/13/15 and 2/16/15 and 2/18/15 and 2/20/15 and 2/23/ and 2/25/1515 Lab 6: Embryology & Zebrafish Development
I. Purpose: The purpose of this lab is to learn about the development of zebrafish by setting up an experiment to study how the environment affects embryonic development. I hypothesize that the zebrafish embryos exposed to nicotine will result in morphogenesis abnormalities in eye size and notochord length. If the control group’s morphology represents normality, then the experimental zebrafish exposed to nicotine will vary.
II. Materials and Methods: Start by labeling two covered petri dishes with initials, date, and egg fertilization date. Label one petri dish “control” and label the other petri dish “nicotine”. Next, fill one petri dish with 20mL of Deerpark Water and add 10 translucent zebrafish embryos. Then, fill the other petri dish with 20mL of 25mg/L nicotine water and add 10 translucent zebrafish embryos. Make observations and record data every Monday, Wednesday, and Friday. Every observation day, remove 10mLs of water from the control group and replace with 15mLs of deerpark water and repeat for the nicotine group with nicotine water. Remove and dispose of empty egg casings every observation day. Save any dead embryos and dead hatchlings in paraformaldehyde. On day 7, feed both the control group and the nicotine group 1 drop of paramecium. End experiment on day 14 and release the surviving hatchlings into the lab’s labeled zebrafish bowl.
III. Data and Observations:
IV. Conclusions: My hypothesis was too difficult to measure given the laboratory settings and materials. Unfortunately, the lab forgot to preserve our formaldehyde preservation hatchlings, and both specimen were compromised, yielding inadequate results due to decomposition. However, it is likely that the nicotine contributed to premature mortality and decreased motor ability.
--Alison D. Bunnen 21:21, 9 March 2015 (EDT)
1/28/15 and 2/4/15 and 2/25/15 Lab:16S Sequencing Analysis
I. Purpose: The purpose of the 16S sequencing was to identify an isolated bacterial colony from our transect.
II. Materials and Methods:
· Run PCR products on agarose gel
· If there is PCR product, purify for DNA sequencing
· BLAST at http://blast.ncbi.nlm.nih.gov/Blast.cgi
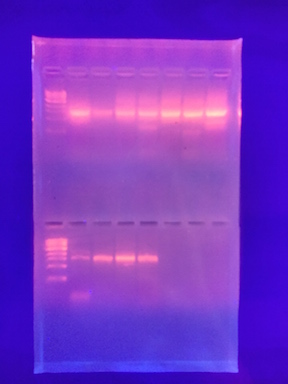
III. Data and Observations: The PCR product was identified from the bacteria in my transect, see Figure 1. The sequence of my 16S PCR product was: 16S sequencing MB 15= >MB84-Rev_16S_G02.ab1 NNNNNNNNNNNNNNNNNNNNNNCATNNNNNCCNNGGTCACANTACNANNNGGGTTCCGGAATTCCTGGCTTGACGGGCGG TGTGTGCAAGGAACGGGAACGTATTCACCGCGCCATGGGTGATGCGCAATTACGATCGATTCCGACTTCAAAGAGTCGAG TTGCAGATTCCNATCGCAACTGAGACCGGCATTGGAGATATGCATCACATCTCCTNTGTANCTGCCCTCTGTACCGGCCA TTGTATTACGTGTGAGGNCCGAAGCCCAAGGGCCGTGATGATTTGACATCTTCCCCACCTTCCTCCCTCCTTGCGTATGC AGTCTCACTACACTCTACANCTTAAAGGTAGCAGCTAGTGACAGGGGTTGCGCTCGTGGCCGGACTTAACCTAACACCTC CAGGCACGAACTGACGACGACCATGCAGCACCTTGAAAATTGTCCGAATGAAGGTACAATTCCCAACCGATCATTTCCCA TTTAATCCTTGGTAAAGTGCNGCTCGNATCATTCAAGTAAACCCNATAANCCACCGCTTGTGCGGGCCTNCTTCAATTCC CTTGAATTTCATTCNNGCTAANGTACTCCCCACGTGGTTAACTTATCANTTTCTTTAAGTCTCTGATTCNNAANACCCAA AANNNATNNNNAACCNNTANACTGANTGNACTACCAGNNTNNCTNATCCTGTTCGCTCCCCACGCTNNCNNCCATCACCG TCANTTAAGACATACTANCCTGCCTTCGNNNATGGTGTTCTAANTAATATCTATGCATTTCACCGCTACACTACTTAGTC CNGCTACNTCTACCTTACTCAAGACCTGCANTATCAATGGGCAGTTTCACAGTTCAGCTGTGANATTTCACCACTGANTT ACANATCCGCCTACNGACCCNTTAAACNCAATATATCCNGANAACNCNTNNNNNNNCCGTATTACCNCNNCTGCTGNGCA CGGANTTNGNNNGTGCTTATTCNTATAGTACNTTCANCTTTCCNCANNTGGANCGGTTNATCCCTNATACAAANANNTTN ANNNCNNNTANNGCNGTCNNCCNTTCACGNNGGCATGGNTNNNTCAGGCTCTCATNCCATTGNTCCANNNTTCNTNNNTG NTGCCNNNNNNNNNAANNNNGGNNNNNNN
mb16= >MB85-Rev_16S_H02.ab1 NNNNNNNNNNNNNNNNNNNNANNAGCTCCTGTTACGGTCACCGACTTCAGGTACCCCAGACTTCCATGGCTTGACGGGCG GTGTGTACAAGGCCCGGGAACGTATTCACCGCGCCATGGCTGATGCGCGATTACTAGCGATTCCAGCTTCATAGAGTCGA GTTGCAGACTCCAATCCGAACTGAGACCAGCTTTCGAGATTCGCATCCAGTCGCCTGGTAGCTGCCCTCTGTACTGGCCA TTGTATTACGTGTGTGGCCCAAGGCGTAAGGGCCGTGATGATTTGACGTCATCCCCACCTTCCTCTCTACTTGCGTAGGC AGTCTCACTAGAGTCCCCAACTGAATGATGGCAACTAGTGACAGGGGTTGCGCTCGTTGCAGGACTTAACCTAACACCTC ACGGCACGAGCTGACGACAACCATGCAGCACCTTGAAAAATGTCCGAAGAAAAGTCTATTTCTAAACCTGTCATTTCCCA TTTAAGCCTTGGTAAGGTTCCTCGCGTATCATCGAATTAAACCACATAATCCACCGCTTGTGCGGGCCCCCGTCAATTCC TTTGAGTTTCAGACTTGCGTCCGTACTCCCCAGGTGGCTAACTTATCACTTTCGCTTAGTCTCTGAAGCCAAGGCCCCAA AAACGAGTTAGCATCGTTTACGGCGTGGACTACCAGGGTATCTAATCCTGTTCGCTCCCCACGCTTTCGTCCATCAGCGT CAGTTGTTGCTTAGTAACCTGCCTTCGCAATTGGTGTTCTAAGTAATATCTATGCATTTCACCGCTACACTACTTATTCC AGCTACTTCAACAACACTCAAGACCTGCAGTATCAATGGCAGTTTCACAGTTGAGCTGTGAGATTTCACCACTGACTTAC AGATCCGCCTACGGACCCTTTAAACCCAATAAATNCCGGATAACGCTNCACCCTCCGTATTACCGCGGCTGCTGGCACGG AGTTAGNCNGTGCTTATTCGTATAGTACCTTCAGCTAGATACACGTANNNNNGTTTATCCCTATACAAANAAGTTTANNN CCATANGNNNTCGTCTTCANGCGGANGGCTGGATCNNNTCTNNCCCATNNNCCAATNNTCNCACTGCTGN.
Both of these sequences are of Chryseobacterium.
Figure 1. Is a picture of the PCR product on agarose.
IV. Conclusions: The bacteria identified in my transect, Chryseobacterium, is a gram negative bacteria that is resistant to many antibiotics. This is consistent with the bacteria characteristics from my agar plates, as both are orange colonies and both are gram negative.
--Alison D. Bunnen 23:19, 27 February 2015 (EST)
2/11/15 Lab 5: Invertebrates and Vertebrates
I. Purpose: The purpose of this lab is to learn about the importance of invertebrates and vertebrates. This lab also taught us how simple systems evolved into complex systems. I hypothesize that we will find many invertebrates from our leaf litter collection. I predict that if there were many types of protists and bacteria found, then we will also find a variety of invertebrates.
II. Materials and Methods:
· Break down the Berlese Funnel
· Pour the top 10mLs of the conical tube into a petri dish
· Pour the remaining liquid into a second petri dish
· Observe and identify the organisms at 4x and 10x
III. Data and Observations: The first organism identified at the bottom of the conical tube was in the phylum and class Nematoda Sercerneta, it was 25mm in length when viewed at 4x, it was long slender and worm-like. The second organism identified at the bottom of the conical tube was in the phylum and class Arthropoda Crustacea Malacostraca, it was too big to be measured when viewed at 4x, it was a pillbug, had lots of legs, and a shell like layer. The third organism identified at the bottom of the conical tube was in the phylum and class Arthropoda Insecta, it was too big to be measured when viewed at 4x, it had 6 legs and was an ant. The fourth organism identified at the bottom of the conical tube was in the phylum and class Arthropoda Arachnida, it was 20mm in length when viewed at 4x, it had a large round body and small legs. The fifth organism identified at the bottom of the conical tube was in the phylum and class Arthropoda Entugnatha, it was 40mm in length when viewed at 4x, it had 6 legs and two antennae.
Figure 1. is the table of invertebrates found in my transect.
IV. Conclusions: My hypothesis was correct in that there was a diversity of invertebrates living in my transect and collected in the leaf litter.
--Alison D. Bunnen 21:58, 27 February 2015 (EST)
2/4/15 Lab 4: Plantae and Fungi
I. Purpose: The purpose of this lab was to understand the diversity of plants and their characteristics, as well as learn the importance of fungi. I hypothesize that there will be a diversity of plants in our transect. I predict that if we can visualize the differences of the plants on the outside, then there will be a difference between the plants on the inside.
II. Materials and Methods:
· Take samples of five plants in a non damaging way and collected any seeds in our transect
· Obtain a "leaf litter" sample and place 500g into a bag
· Create a Berlese Funnel
→ Add 25mL of 50:50 ethanol:water solution into a 50mL conical tube.
→ Place a small square of screening into the bottom of the funnel to prevent the leaf litter from falling into our tube.
→ Add the leaf litter to the funnel and place 2 inches from a 40 watt lamp and cover in foil.
III. Data and Observations: The five samples plants had different mechanisms of reproduction. All of the sample plants had phloem and xylem as vascularization. All sample plants had cuticles and stomata for specialized structures. Transect sample plant #1 and #2 were hard to identify and therefore I was unable to recognize the mechanisms of reproduction. Transect sample plant #3 was a hollie shrub and the mechanism of reproduction is asexual. Transect sample plant #4 was an flowering because of the flowering bud that it had. Transect sample plant #5 was identified as a maple leaf and maple trees are angiosperms.
The fungus growing on my 10-3 nutrient agar plate + tet plate was identified as white mold. White mold is ascomycota which is the largest division of fungi. See figure 2. for a picture of the white mold.
Figure 1. is a table of the transect sample plants collected during lab.
Figure 2. is of the white mold found on my 10-3 nutrient agar plate + tet plate
IV. Conclusions: My hypothesis was wrong. Just because the external features of our plants was different, the internal features were very similar. However, the mechanisms of reproduction varied regardless of the internal structures.
--Alison D. Bunnen 21:19, 27 February 2015 (EST)
1/28/15 Lab 3: Microbiology and Identifying Bacteria with DNA Sequences
I. Purpose: The purpose of this lab was to understand and identify the characteristics of bacteria, observe antibiotic resistance in the nutrient agar + tet plates. I hypothesize we will find both gram positive and gram negative bacteria in our agar plates. If we take samples from our Hay Infusion Culture, where we already identified protists and algae, then there will be all sorts of bacteria.
II. Materials and Methods:
· Observe agar plates
· Record the 100-fold serial dilutions results
· Make a wet mount and also make gram stains of 2 nutrient agar plates and 2 nutrient agar + tet plates > we chose to use nutrient agar plates 10-3 and 10-9, and used nutrient agar + tet plates 10-3 and 10-9
· Record description of cells and motility
· Set up PCR for 16S sequencing
→ Pick a colony of bacteria and transfer it to 100μL of water in a sterile tube
→ Incubate for 10 minutes at 100°C in a heat block
→ At 13,400 rpm, centrifuge for 5 minutes
→ While the sample is centrifuging, add 20μL of primer/water mixture into a PCR tube. Mix to disolve the PCR bead. Label tube
→ After centrifuging, transfer 5μL of supernatant to the 16S PCR reaction.
→ Place the tube in the PCR machine
III. Data and Observations:
The Hay Infusion Cultures changed slightly in appearance and smell. The water is more murky and smells worse, this is probably due to all the bacteria, fungi, and algae in the jar. I hypothesize everyone's Hay Infusion Cultures smell worse. If the organisms inside the jar are multiplying, then the jar will be filled with more organisms and will smell worse.
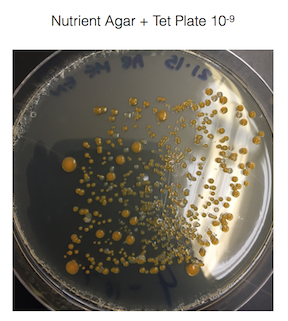
The agar plates with tetracycline grew the gram - bacteria, but not the gram + bacteria and also grew fungi. It would seem that gram + bacteria is susceptible to tetracycline. However, I know from in a pharmacology class that I took at Bel-Rea Insititute of Animal Technology that tetracycline is a broad spectrum antibiotic that affects both gram positive and gram negative bacteria. This leads me to believe the gram negative bacteria that has grown on the nutrient agar + tet plates are antibiotic resistant.
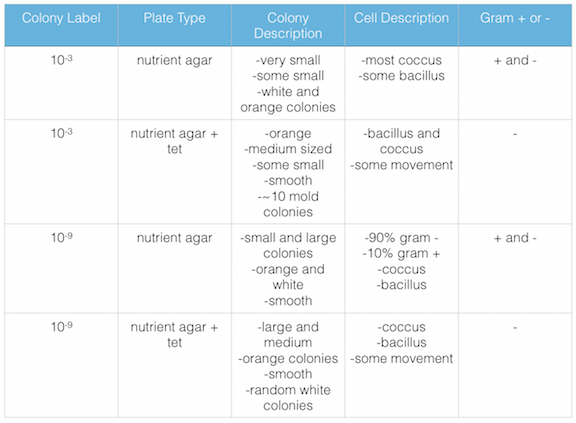
Figure 1. shows the 100-fold serial dilutions results with the plates that we chose to gram stain and do further testing on starred.
Figure 2. shows the bacteria characteristics
IV. Conclusions: My hypothesis was correct. Both gram positive and gram negative bacteria are present in the cultures that we took.
--Alison D. Bunnen 21:15, 4 February 2015 (EST)
1/21/15 Lab 2: Identifying Algae and Protists
I. Purpose: The purpose of this lab is to examine and understand the characteristics of algae and protists from my transect that is found in the Hay Infusion Culture. Please see Lab 1 for how the Hay Infusion Culture was created. I hypothesize the transect have copious amounts of bacteria living in the niche. If water is added to the gathered materials and is allowed to incubate, then the bacteria from the niche (in the soil) will grow in the new ecosystem created.
II. Materials and Methods:
· Make a wet mount from the niche at the top of the jar and make a wet mount from the niche at the bottom of the jar
· View at 4x, 10x, and 40x.
· Make observations of at least 3 different organisms from each niche
· Use a dichotomous key to identify the organisms
III. Data and Observations: My Hay Infusion Culture has a foul smell, almost like rotten garbage. The top of the jar has a layer of film, maybe mold, with a gray/green color. The water is yellow/brown tinged. The bottom of the jar has a thin layer of dirt on it.
Figure 1. is the top view of my Hay Infusion Culture for Group 3
Figure 2. is the side view of my Hay Infusion Culture for Group 3
The organisms below were found at the top of the jar niche:
Figure 1. is of Organism 1. In lab I established this organism to be a Didinium Cyst, however after further research, that is not what Organism 1 is. At the moment, Organism 1 is still unknown. It is ~30μm , almost circular with two indentations at the top and bottom of the cell, little oval specks inside moving, and colorless.
Figure 2. is of Organism 2. In lab, I established this organism to be Colpidium. The organism varied in size, one I measured was ~50μm, packman-like face, spins, type of motility undetected, and colorless.
Figure 3. is of Organism 3. In lab, I established this organism to be Euglena, however after extensive google image searches, I believe it to be Fusarium graminearum. It is ~50μm, peapod/canoe shaped, long, thin, some are segmented inside, and colorless.
Figure 4. is of Organism 4. In lab, I could not determine the genus and species. It is ~75μm, disc-like, pods in the middle and colorless.
The organisms below were found at the bottom of the jar niche:
Figure 5. is of Organism 5. In lab, my lab partners could not determine the organism. It is colorless at ~60μm. Photograph by Emma Niden.
Figure 6. is of Organism 6. In lab, my lab partners could not determine the organism. It is colorless, and kidney bean shaped. Photograph by Emma Niden.
Figure 7. is of Organism 7. In lab, my lab partners determined this to be algae. Photograph by Emma Niden.
IV. Conclusions: After observing and identifying 6 or more organisms residing in the two niches of my Hay Infusion Culture, I have determined my hypothesis to be true. There were multiple organisms growing in my group's Hay Infusion Culture that we observed and we were able to identify a few of them.
--Alison D. Bunnen 18:33, 29 January 2015 (EST)
1/14/15 Lab 1: Biological Life at AU
I. Purpose: The purpose of today's lab was to observe and record characteristics of an assigned transect. We took samples from our niche to create a Hay Infusion Culture. I hypothesize the transect have copious amounts of bacteria living in the niche. If water is added to the gathered materials and is allowed to incubate, then the bacteria from the niche (in the soil) will grow in the new ecosystem created.
II. Materials and Methods:
· 20 x 20 meter transect to study
· Use a sterile 50 mL conical tube to gather materials such as soil and vegetation from transect
· Place 10-12 grams of gathered materials into a large jar
· Add 500 mLs of deerpark water to the jar
· Add 0.1 gm of dried milk into the jar and screw on lid
· Mix contents gently for 10 seconds
· Place the jar in designated lab section area with the jar labeled and lid off
III. Data and Observations: My assigned transect was group 3: Tall Bushes. I recorded macroscopic observations of abiotic and biotic components in my transect. Then my group made a Hay Infusion Culture from materials gathered from our niche. The Hay Infusion Culture was a mix of ~10 grams of soil and vegetation from my transect, 500 mL of deerpark water, and 0.1 gm of dried milk.
The abiotic components inside the transect included:
· Light
· Water/snow
· Concrete
· A metal lamp post
· Dirt
· Rocks
The biotic components inside the transect included:
· Grass
· Shrub
· Bushes
Figure 1. is an aerial-view diagram of my transect labeled with it's abiotic and biotic components.
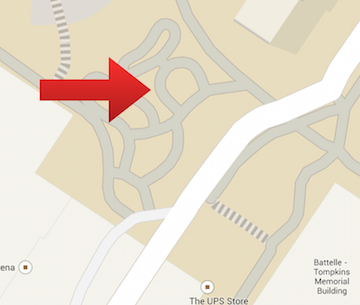
Figure 2. is a satellite view of Group 3's transect.
Figure 3. is a map view of Group 3's transect.
IV. Conclusions: There are abiotic and biotic components in Group 3's transect. I will not know if my hypothesis is correct until further examination of the incubated Hay Infusion Culture is done.
--Alison D. Bunnen 22:24, 26 January 2015 (EST)
References
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 19-23.