SBB12Ntbk-GregFukushima
Greg Fukushima, 7 February 2012
This is my first day of notebooks for 140L! I love wikis!
Greg Fukushima, 16 February 2012
First Day of PCR and cloning.
Construction File for SOEing PCR
PCR ca998/gfRevPR on pBjk2741-jtk2801 (160bp, A) PCR gfForRbsCmR/gfRevRbsCmR on pKGC438 (920bp, B) PCR ca998/gfRevRbsCmR on A+B (1059bp, pcrpdt) Digest pcrpdt (EcoRI/BamHI, L, pcrdig) Digest pBgl00001-Bjk2828 (EcoRI/BamHI, L, plasdig) Ligate pcrdig and plasdig, product is pBgl00001-sbb1220 ---- >ca998 Forward external annealing for purification of P_R part gtatcacgaggcagaatttcag >gfForRbsCmR ggtgataatggttgcGGATCTCCACAACGGTTTCCCTC >gfRevRbsCmR gctagGGATCCTTACGCCCCGCCCTGCCAC >gfRevPR AGATCCgcaaccattatcacc >pcrpdt GTATCACGAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATCTGAAGTGGAATTCATGAGATCTTAAATCTATCACCGCAAGGGATAAATATCTAACACCGTGCGTGTTGACTATTTTACCTCTGGCGGTGATAATGGTTGCGGATCTCCACAACGGTTTCCCTCTAGAAATAATTTTGTTTAACTTTAAGAAGGAGATATACATATGAGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCATGGAGCGAGAAAAAAATCACTGGATATACCACCGTTGATATATCCCAATGGCATCGTAAAGAACATTTTGAGGCATTTCAGTCAGTTGCTCAATGTACCTATAACCAGACCGTTCAGCTGGATATTACGGCCTTTTTAAAGACCGTAAAGAAAAATAAGCACAAGTTTTATCCGGCCTTTATTCACATTCTTGCCCGCCTGATGAATGCTCATCCGGAGTTCCGTATGGCAATGAAAGACGGTGAGCTGGTGATATGGGATAGTGTTCACCCTTGTTACACCGTTTTCCATGAGCAAACTGAAACGTTTTCATCGCTCTGGAGTGAATACCACGACGATTTCCGGCAGTTTCTACACATATATTCGCAAGATGTGGCGTGTTACGGTGAAAACCTGGCCTATTTCCCTAAAGGGTTTATTGAGAATATGTTTTTCGTCTCAGCCAATCCCTGGGTGAGTTTCACCAGTTTTGATTTAAACGTGGCCAATATGGACAACTTCTTCGCCCCCGTTTTCACGATGGGCAAATATTATACGCAAGGCGACAAGGTGCTGATGCCGCTGGCGATTCAGGTTCATCATGCCGTCTGTGATGGCTTCCATGTCGGCAGAATGCTTAATGAATTACAACAGTACTGCGATGAGTGGCAGGGCGGGGCGTAAGGATCCCTAGC
Protocol for PCR:
Made oligo dilutions of ca998/gfRevPR:
9uL Water 1uL 100uM oligo ca998 or gfRevPR
Set up the following reaction in a PCR tube:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL gfRevPR, 10uM 1uL ca998, 10uM 0.5uL Expand polymerase "1" 0.5uL pBjk2741-jtk2801 Template DNA
Actual reaction was done by Zach.
NOTE: I was only able to do the first PCR reaction on my construction file because pKGC438 was not available.
Construction File for Leucine Zipper PCA
PCA1 on o1,o10,o12 (pca1) PCA2 with o1/o12 on pca1 (139 bp, pca2) Digest pca2 (NheI/BamHI, L, 1210dig) Digest pBca9525-Bca1834 (NheI/BamHI, L, vectdig) Ligate 1210dig + vectdig, product is pBca9525-sbb1210 ---- >o1 CCATAgctagcGGCAGTGGATCTGTTAAAGAACTGGAAGACAAAAACGAAGAACTGCTGAGT >o10 CAAAAACGAAGAACTGCTGAGTATCATCTACCACCTGAAAAACGAAGTTGCTCGTCTGA >o12 CAGTAGGATCCTTAGCCGCCACGTTCGCCAACCAGTTTTTTCAGACGAGCAACTTCGTT >pca2 CCATAgctagcGGCAGTGGATCTGTTAAAGAACTGGAAGACAAAAACGAAGAACTGCTGAGTATCATCTACCACCTGAAAAACGAAGTTGCTCGTCTGAAAAAACTGGTTGGCGAACGTGGCGGCTAAGGATCCTACTG
Protocol for PCA:
Added following into PCR tube for the PCA reaction:
- 38 uL ddH2O
- 5 ul 10x expand buffer
- 5 ul 2mM dNTPs
- 1 ul oligo mixture(o1,o10,o12) (100uM total, mixture of oligos after combination of 100uM stocks)
- 0.75 ul Expand polymerase
Actual reaction was done by Zach.
Greg Fukushima, 17 February 2012
Continued PCR and PCA reactions. Did the second PCR reaction listed on the construction file. Also did amplification step of the Leucine Zipper PCA reaction. Finally did the Small-Frag Zymo Cleanup on "A" from the SOEing PCR construction file.
Protocol for PCR:
Made oligo dilutions of gfForRbsCmR/gfRevRbsCm:
9uL Water 1uL 100uM oligo gfForRbsCmR or gfRevRbsCm
Set up the following reaction in a PCR tube:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL gfRevRbsCmR, 10uM 1uL gfForRbsCmR, 10uM 0.5uL Expand polymerase "1" 0.5uL pKGC438 Template DNA
Actual reaction was done by Zach.
Protocol for PCA:
Added following into PCR tube for the PCA reaction:
- 1 ul of each 10ul outer oligo(o1/o12)
- 1 ul purified pca product(PCA1)
- .5 ul phusion
- 10 ul 5x phusion buffer
- 5 ul 2mM dNTPs
- 32.5 ul H2O
Actual reaction was done by Zach.
Small-Frag Zymo Cleanup Protocol:
- Add 100 uL of Zymo ADB buffer (brown bottle) to the reaction product "A".
- Transfer into the Zymo column
- Add 500uL of Ethanol and pipette up and down to mix
- spin through (15s), discard waste.
- Add 200 uL of Zymo Wash Buffer (which is basically 70% ethanol)
- spin through (15s), discard waste.
- Add 200 uL of Zymo Wash Buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with water (spin 60s) into a fresh Eppendorf tube
Greg Fukushima, 23 February 2012
Did partial EcoRI/BamHI digest of PCA product as well as the gel for the 2 PCR reactions from before.
Protocol for Gels:
- For each PCR, load 6uL of PCR product premixed with 4uL of loading buffer in a single well of a 1% agarose gel.
- Run gel
- Cut out the bands, put them into a single 1.5mL microcentrifuge tube.
- Add 650uL of ADB buffer
- Freeze to store for tomorrow.
My lanes were lanes 1 and 2. Lane 1 was pKGC438 and lane 2 was pBjk2741-jtk2801.
NheI/BamHI Digest of PCA Product:
- Set up the following reaction:
8uL of eluted PCA product 1uL of NEB Buffer 2 0.5uL NheI 0.5uL BamHI
- Incubate at 37 degrees on the thermocycler for 1hr
- Store for gel tomorrow
Greg Fukushima, 24 February 2012
Ran the Zymo Gel Purification on the gel PCR products from yesterday. Also did the gel for the PCA NheI/BamHI Digest from yesterday.
Protocol for Zymo Gel Purification:
- All spins until the drying step are 15 second full speed spins.
- heat the bands + ADB buffer in ependorf tube at 55, shake and/or vortex until the gel has dissolved.
- If the DNA is <300bp add 250uL of isopropanol
- transfer into the Zymo column inside a collection tube (small clear guys)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with water into a fresh Eppendorf tube
Protocol for continuation of NheI/BamHI digest of PCA:
- Mix the digested PCA product with 2uL of loading buffer.
- Put mixture into the gel and run.
- Cut out the bands and melt with 600uL of ADB buffer at 55 degrees.
- Freeze and save for next time.
Greg Fukushima, 8 March 2012
I redid the PCA gel from February 24 because the lanes were not labeled so I did not know which one was mine. Also set up the PCR reaction for A+B to make pcrpdt in the construction file.
Protocol to redo NheI/BamHI gel of PCA:
- Mix 4uL of the digested PCA product with 2uL of loading buffer.
- Put mixture into the gel and run.
- Cut out the bands and melt with 600uL of ADB buffer at 55 degrees.
- Freeze and save for next time.
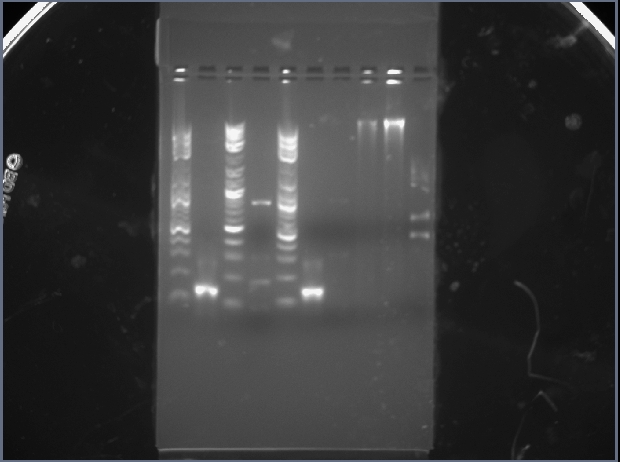
Class Gel 1
Lane 2 contained my PCA.
Protocol for setting up PCR:
Set up the following reaction in a PCR tube:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL gfRevRbsCmR, 10uM 1uL ca998, 10uM 0.5uL Expand polymerase "1" 0.5uL "A" Template DNA 0.5uL "B" Template DNA
Greg Fukushima, 13 March 2012
Did zymo gel purification on PCA digest from last time and a regular zymo cleanup on pcrpdt. Also did digestion of pcrpdt with EcoRI and BamHI. Need to run gel on the digested pcrpdt next time.
Protocol for Zymo Gel Purification:
- All spins until the drying step are 15 second full speed spins.
- cut out bands minimizing extra gel matter.
- put in ependorf tube and add 600uL of Zymo ADB buffer (brown bottle).
- heat at 55, shake and/or vortex until the gel has dissolved.
- add 250uL of isopropanol
- transfer into the Zymo column inside a collection tube (small clear guys)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with 8uL of water into a fresh Eppendorf tube
Protocol for Regular Zymo Cleanup:
- Add 180 uL of Zymo ADB buffer (brown bottle) to the 33uL reaction
- Transfer into the Zymo column (small clear guys)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with 33uL of water into a fresh Eppendorf tube
Protocol for EcoRI/BamHI Digest of PCR Products:
- Set up the following reaction:
8uL of eluted PCR product (pcrpdt) 1uL of NEB Buffer 2 0.5uL EcoRI 0.5uL BamHI
- Incubate at 37 degrees on the thermocycler for 1hr
- Save for gel next time
Greg Fukushima, 20 March 2012
Ran the pcrpdt gel and did gel purification. Started a ligation but threw away product because temperature for heat shock was not correct.
Protocol for Zymo Gel Purification:
- All spins until the drying step are 15 second full speed spins.
- cut out bands minimizing extra gel matter.
- put in ependorf tube and add 600uL of Zymo ADB buffer (brown bottle).
- heat at 55, shake and/or vortex until the gel has dissolved.
- transfer into the Zymo column inside a collection tube (small clear guys)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of Zymo Wash Buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with 6uL of water into a fresh Eppendorf tube
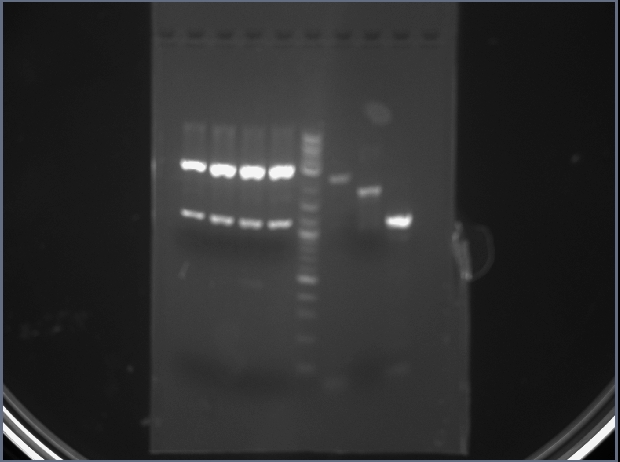
My pcrpdt was in lane 9.
Greg Fukushima, 22 March 2012
Did ligation and transformation for both PCR product and PCA product.
Protocol for Ligation of EcoRI/BamHI digests
- Set up the following reaction:
6.5uL ddH2O 1uL T4 DNA Ligase Buffer (small red or black-striped tubes) 1uL Vector digest 1uL Insert digest 0.5uL T4 DNA Ligase
- Pound upside down on the bench to mix
- Give it a quick spin to send it back to the bottom of the tube
- Incubate on the benchtop for 30min
- Put on ice and proceed to the transformation
Protocol for Transformation by heat-shock
1. Thaw a 200 uL aliquot of cells on ice 2. Add 30 uL of KCM to the cells 3. Put your ligation mixture on ice, let cool a minute or two 4. Add 70 uL of the cell cocktail to the ligation, stir to mix 5. Let sit on ice for 10 min 6. Heat shock for 120 seconds at 42 7. Put back on ice for 1 min 8. Add 100uL of 2YT, let shake in the 37 degree incubator for 1 hour 9. Plate 70+ uL on selective antibiotics, let incubate at 37 degrees overnight
Followed these protocols twice, once for the PCR and once for the PCA product.