McClean: Matlab Code for Analyzing FACS Data
Summary
This is a collection of Matlab functions for plotting flow cytometry data.
All of the code requires the function fca_readfcs written by Laszlo Balkay available from the Mathworks file exchange at: FCS data reader
Information about flow cytometry at Princeton can be found at: Flow Cytometry Resource Facility
Example: GFP Induction Timecourse
The following example code is for visualization of a time series of GFP fluorescence (expressed from pGAL1-GFP in yeast cells) collected using flow cytometry. The full m-file (Example_165.m) along with the individual functions can be found here: Flow Cytometry Timecourse Code
Variables to pass to the individual functions:
%% Some variables that can be quickly changed to modify this for another timecourse NameFiles='165'; %Name common to all the files in the timecourse (For this timecourse the files are named '_165T0.fcs','_165T1.fcs',....'_165T7.fcs') Time=[0 1 2 3 4 5 6 7]; %Timepoints Output=0; %Print the output (1) or don't (0) to the screen from the various functions column=3; %Need column 4 for the mCherry data, 3 for GFP, this may be different for your FACS experiment Bins=linspace(0,10,100); %Bins to use for plotting histograms
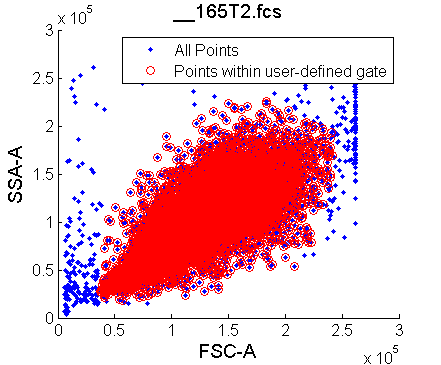
User gates on the side- and forward-scatter:
%% 1. User visually defines the gates based on side-scatter and forward-scatter using the mouse to map out a region
[Gates,FNames]=FindGate(strcat('*',NameFiles,'*','fcs'), Output);
save(strcat(NameFiles,'_GateVariables'),'Gates','FNames'); %Save gate info so analysis can be reproduced later
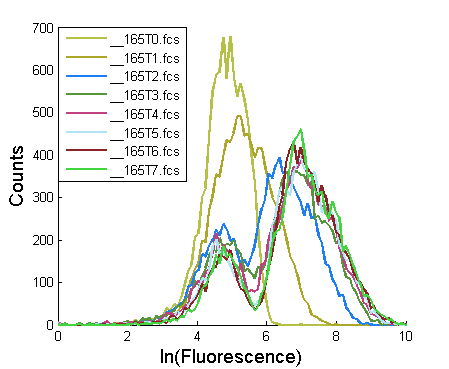
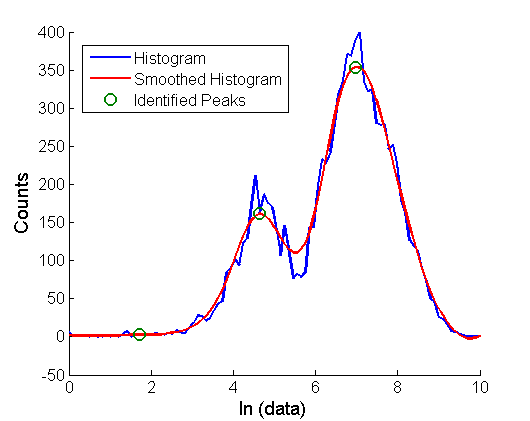
Plot histograms of the logarithm of the GFP intensities at each time point:
%% 2. Histograms of the timecourses are plotted (FACSSTC also saves a plot of the histograms)
[LOGM]=FACSTC_PlotHist(FNames, Gates, column, Bins,strcat('h',NameFiles,'_histograms'));
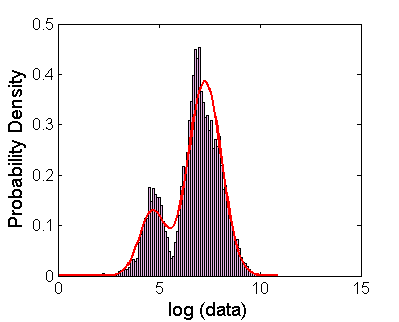
Peaks in the distribution are found by fitting to one or two normal distributions (this assumes that the logarithm of the data is distributed approximately log-normally)
%% 3. Find peaks by fitting logarithm of intensities to normal distributions (this assumes that the intensity data is originally lognormal distributed) (FACSTimecourse) %%Fit to normals or two normals (the logarithm of the data) and then plot %%the centers of the distributions as a function of time %P1 contains the 1 gaussian fit parameters, P2 contains the 2 gaussian fit parameters [P1 P2]=FACSTimecourse(FNames, column, Gates,Output);
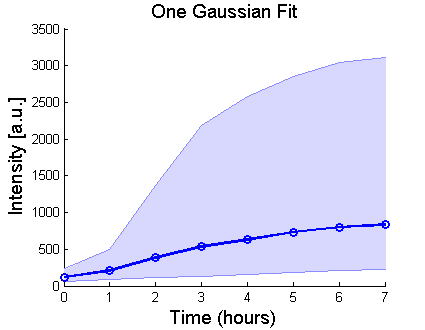
%Plot the peaks from the single gaussian fit:
figure; hold;
H=shadedLogErrorBar(Time, exp(P1(:,1)),exp(P1(:,2)),{'bo-','LineWidth',2});
title('One Gaussian Fit','FontSize',14);
xlabel('Time (hours)','FontSize',14); ylabel('Intensity [a.u.]','FontSize',14);
H=gcf;
saveas(H,strcat(NameFiles,'_low'),'fig')
saveas(H,strcat(NameFiles,'_low'),'png')
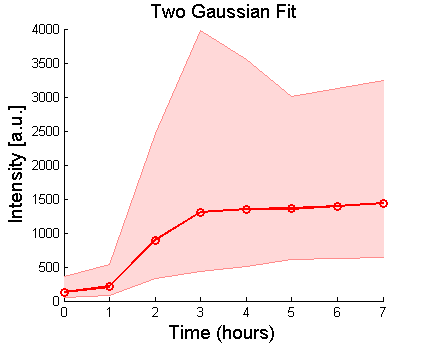
%Plot the peak of the induced population from the two-gaussian fit:
figure; hold;
H=shadedLogErrorBar(Time, max(exp(P2(:,2)),exp(P2(:,3))),max(exp(P2(:,4)),exp(P2(:,5))),{'ro-','LineWidth',2});
title('Two Gaussian Fit','FontSize',14);
xlabel('Time (hours)','FontSize',14); ylabel('Intensity [a.u.]','FontSize',14);
H=gcf;
saveas(H,strcat(NameFiles,'_high'),'fig')
saveas(H,strcat(NameFiles,'_high'),'png')
pause; close all;
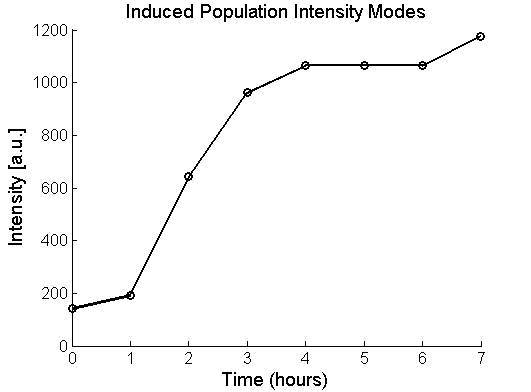
%% 4. Find peaks using a different function (FACSTC_PeakFind) based on Matlab's findpeaks %Find the midpoints of populations by using a different function based on %Matlab's findpeaks function: [PeakLocations]=FACSTC_PeakFind(FNames, column,linspace(0,10,100),Gates,Output); P3=PeakLocations; close all;
figure; hold;
plot(Time,exp(P3(:,1)),'ko-','LineWidth',2);
plot(Time,exp(P3(:,2)),'bo-','LineWidth',2);
title('Induced and Uninduced Population Intensity Modes','FontSize',14);
xlabel('Time (hours)','FontSize',14); ylabel('Intensity [a.u.]','FontSize',14);
h=gcf;
saveas(h,strcat(NameFiles,'_peakfind'),'fig')
saveas(h,strcat(NameFiles,'_peakfind'),'png')
pause; close all;
%% %%Save the fitted peak centers to be used later for plotting P1_low=P1; P2_high=P2; P3_peaks=P3; save(strcat(NameFiles,'_Peaks'),'P1_low','P2_high','P3_peaks');
Notes
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input!
- Megan N McClean 17:27, 22 February 2012 (EDT): I wrote most of this to be able to take a timecourse of induction of some fluorescent protein of interest and plot the midpoint of the induced population. As usual, it isn't perfect, but maybe it will serve as a useful starting point for someone else in the lab.
References
Mathworks Online Help: http://www.mathworks.com/help
Contact
- Megan N McClean 14:01, 22 February 2012 (EDT)
or instead, discuss this protocol.