IGEM:Imperial/2010/Detection module/2 components systems
Quorum Sensing in B.substilis
"Peptide messages of Gram(+) species are genetically encoded as precursor peptides. These bacteria encode transmembrane enzymes, which proteolytically process the precursors while extruding them into the extracellular space. These proteins thus serve the function of synthases and exporters simultaneously. Two strategies are in use for detection of peptides in the environment: receptor histidine kinases and active importers, usually from the family of ABC transporters. Once imported into the cell, the peptides are sensed by intracellular receptors preventing it from dephosphorylating a transcriptionally active response regulator."ref
Cell surface peptide display and the associated QS of Gram-positive bacteria provides a useful mechanism which we can manipulate.
The idea is to fuse a split protease onto downstream RRs of an engineered QSS artificially introduced into B.substilis.
One concern is that the extracellular proteases produced by B. subtilis may interfere with the system and give false positives. There are two ways we can overcome this problem:
- Use a strain of B. subtilis which does not produce extracellular proteases (but these cells may not be all that healthy!)
- Ensure that the extracellular proteases are not specific for the cleavage site on the membrane-bound protein which, when cleaved, releases the peptide that we want to detect.
The membrane-bound peptide on the surface of our bacterium would have a cleavage site that only the parasite protease (eg elastase) would recognise. The resulting cleavage peptide would bind to a receptor on the extracellular side of the membrane and trigger a response.
NB Newcastle 2008 iGEM team worked on BugBuster which provides some useful info :B
Prof Selkirk suggested using the peptide autoinducer FMLP. However, if elastase always cleaves on the C terminal side of lysine, we would probably keep ending up with an extra lysine on the peptide, meaning that the peptide may no longer fit the receptor. This review has loads of information about peptide autoinducers.
Two Component Systems
Our current idea is two engineer a hybrid 2CS, the sensor receptor will be an endogenous QSS of gram positive bacteria (not naturally occurring in B.subtilis) with the effector domains and RRs of a 2CS which homo or heterodimerize. related review We need 2 downstream interacting proteins which will bind upon external signalling. These two domains will bring together the split protease which will become functional.

List of TCS
NtrC and PhoB: phosphorylation induces dimerization of the receiver modules paper
Dimerization allows DNA target site recognition by the NarL response regulator
paper
KEGG B.subtilis list of 2CS KEGG useful list of all E.coli 2CS
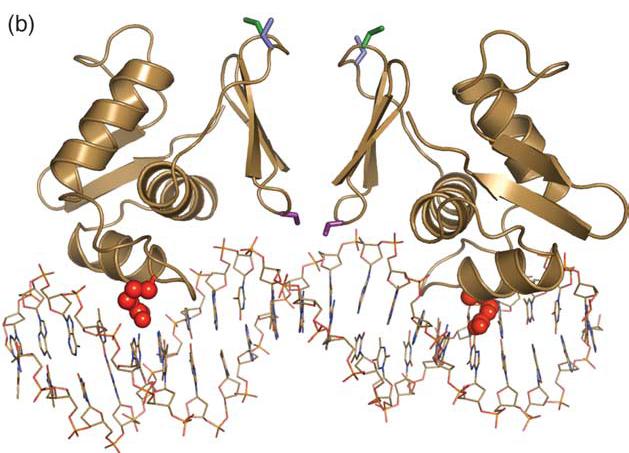
The response regulator OmpR oligomerizes via beta-sheets to form head-to-head dimers EnvZ is a promising downstream 2CS which results in OmpR dimer formation.
- RcsA & B
- PhoP & Q
- SarA & R
- NrI & II
- NtrC & B
- TraA in Agrobacterium
- ComX: linear peptide appears to contain it's activity in contrast to the AgrD linear peptide. A minimal sequence has been identified by Okada et al. (2009): tripeptide [3-5]ComXRo-2. ComQ is the enzyme responsible for isoprenoid addition to a central tryp (Schneider et al 2002)
So far none of the heterodimerising networks can sense a small peptide. As the result we are looking into the feasibility of making a chimeric protein in the network to connect one that uses a signal peptide and one that produces a heterodimeric output (transcription factor)
There has been much literature looking into the feasibility of combining several two-component systems. We are in the process of reviewing past experiments and methods:
Rewiring the Specificity of Two-Component Signal Transduction Systems [1]
Rewiring Bacteria, Two Components at a Time [2]
Computational design of receptor and sensor proteins with novel functions [3]
Using Engineered Scaffold Interactions to Reshape MAP Kinase Pathway Signaling Dynamics [4]
Engineering key components in a synthetic eukaryotic signal transduction pathway [5]
AgR sytem
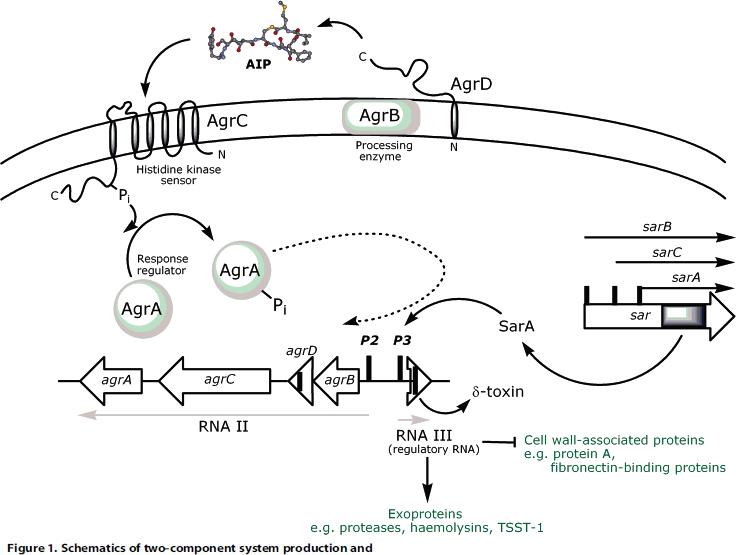
- The AIP used here, AgrD is subjected to intensive posttranslational modification. It is produced as a transmembrane precursor from wich the functional AIP is cleaved (two peptidase steps) to be released into the extracellular space. Furthermore, the AIP is circularized in this step. The fact that those modifications are integrated into the native secretion mechanism prevents us from using this particular AIP in our detector-fusion peptide.
- The 2CS receptor, independent of ligand binding, forms homodimers. Two facts here render the system non-suitable for our approach - we cannot "induce" dimerization and the dimerization occurs on the receptor level, not on the TF level.
We learn from this, that we have to ensure that the AIP released from our detector peptide are active in a non-modified form (e.g. receptor recognizes linear epitope or modification also occurs when AIP is fused in the manner of our detector peptide) and that the receptor-downstream signalling involves dimerizing TF's.
Linear AIPs
CSP-1, BIP-1 dependent pathways in Strep. pneumoniae
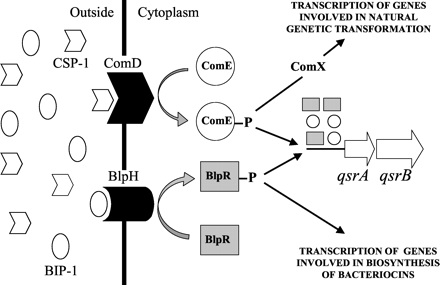
Using linear AIPs would mean that we wouldn’t have to worry about the cyclisation step, which many AIPs need. One example is BlpC from Streptococcus pneumoniae, which is the ligand for a two component signal transduction system. There are 3 different variants of BlpC, called BIP-1, -2 and -3 and their sequences can be found in this paper, which also explains how BlpC upregulates the expression of the same ABC transporter as ComC. It also tells us that The target site of ComE consists of two 9-bp imperfect direct repeats separated by a stretch of 12 nucleotides (34). The consensus sequence of the 9-bp direct repeat is 5-(AGT)CA(GCT)TT(GT)(AG)G-3.
The above image shows that ComE homodimers and BlpR homodimers can form, as well as ComE-BlpR heterodimers. This could be used to bring our TEV protease together. However,the crystal structures of these proteins may not be available and without them, making fusion proteins would probably be very difficult.
The CSP-1 and BIP-1 AIPs do contain an internal lysine, but the schistosoma protease would be unlikely to cleave them internally because the lysines are not contained within the appropriate recognition motif.
We'll hopefully use CSP-1, which has the following primary sequence: N-EMRLSKFFRDFILQRKK-C
At the C-terminus, we'll add on SWPL, so that the cercariae elastase will recognise the C-terminal lysine and cleave at the correct position, yielding CSP-1, which can then diffuse to the receptor (ComD).
Homology searches using ComE and BlpR
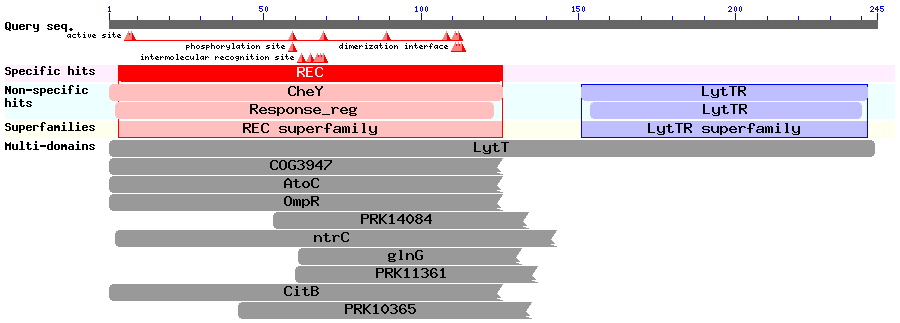
Results of a BLAST search using BlpR (Accession no. CAC03515.1):
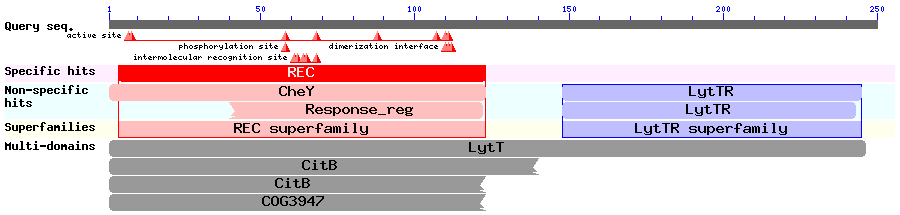
Results of a BLAST search using ComE (Accession no. AAC44897):
These results might help us decide where to fuse our split protease to the response regulators.
This review talks about the structures of response regulators. It says that the DNA-binding domain of the LytTR superfamily of RRs hadn't been solved at that time (it was published in 2006).
The LytTR superfamily of RRs binds to DNA via a 'novel' DBD. This paper describes and also mentions BlpR and ComE, but doesn't discuss dimerisation of RRs much. This article describes how they solved the crystal structure of the RR AgrA from S. aureus (a member of the LytTR superfamily) and show how it binds to DNA. The two proteins bind to DNA at a distance of 12bp from each other, but it is unclear when exactly the dimerisation occurs.
NB ComD accession no AAC44896.1, ComC accession no AAC44895.1
Threshold levels of CSP
This paper show that only 10ng/ml of synthetic CSP was needed for competence-stimulating activity. It also explains what parameters affect the response to CSP, such as pH and growth phase.
ComX/ComP/ComA system in B. subtilis
ComX is one of the two AIP's signalling density depentend induction of competende in B.subtilis (together with CFS). Different strains of B. sub use different ComX variants. For example the laboratory strain Ro-2 does not recognize ComX released by wildtype forms of B.sub (Pottathil et al 2008) - the dangers of false positives would therefore be limited.
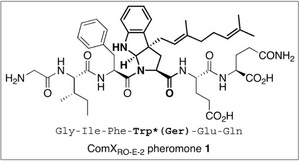
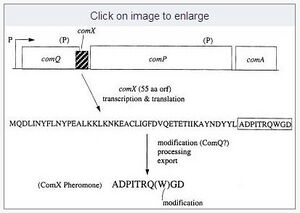
The active AIP ComX of the strain Ro-2 is a linear hexapeptide, including a central Tryptophan, which is modified by addition of an isoprenoid, either Farnesyl or Geranyl (see figure) (Okada et al, 2008).
ComX is produced as a 55aa precursor which is modified by cleavage and addition of the isoprenoid residue to the central Trp. The hexapeptide hereby is positioned at the C-terminal end (?) of the precursor. While it is unkown which peptidase performs the cleavage, trp modification is most likely carried out by ComQ. ComQ appears to bind pre-synthesised isoprenoid residues (hepS, hepT and uppS seem to be involved here) via conserved aspartates in region II of the enzyme and transfere it to the ComX trp. Potentially we could engineer our detector protein with a sequence which would still allow ComQ to recognize the ComX sequence in the fusion protein. Information on the reaction mechanism is avaliable (Okada et al, 2008), however on the specific enzyme-substrate interactions no informaion has so far been published.
The sequence of the hexapeptide has been analysed by Okada et al (2007) who found only the residues 3-5 necessary for close-to normal activity. Interestingly, addition of Alanine residues to both ends did not significantly alter the activity of the AIP. Thus it appears that the central, modified Trp is both necessary and sufficient for specific recognition by the cognate receptor ComP.
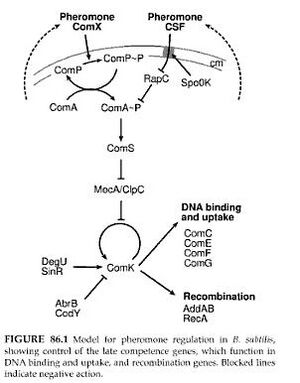
The downstream signalling invovles ComA, which dimerizes and binds to DNA to act as a TF (Griffith et al 2008)
The ComA binding domain crystal structure: NMR Solution Structure and DNA-binding Model of the DNA-binding Domain of Competence Protein A (ComA), 2010
 |
 |
 |
EnvZ
Altering the specificity of the EnvZ receptor
If we could change the specificity of EnvZ by changing the extracellular domain (eg by replacing it with that of an extracellular domain of a receptor which recognises AIPs), we could still use the downstream signaling pathways of EnvZ, but we'd still be able to detect AIPs.
This paper shows that it has been possible to fuse the sensory domain of a sugar binding protein receptor to the cytoplasmic domain of EnvZ. This is because both receptors had similar transmembrane domains.
We looked into the feasibility of using this system to detect a quorum sensing peptide. The main problem we encountered is that there is no natural mechanism to detect anything bigger than a di-peptide. Moreover none of the QS molecule receptors were sufficiently similar to be reliably fused with EnvZ to produce a functional receptor. In addition the outer membraine of E.coli would be impermeable to small peptides such as ComE (signalling peptide)
Periplasmic Binding Protein Receptors-EnvZ hybrids
Tsr Tar Tgr and Tap are receptors involved in E.coli sensory response producing a chemotactic output. "Each of the four chemoreceptors recognizes different ligands. Three interact with ligand occupied periplasmic binding proteins. The receptor Trg binds galactose and ribose-binding proteins. Tap recognizes dipeptide-binding protein, and Tar interacts with maltose-binding protein. In addition, Tar binds aspartate directly, and the fourth receptor, Tsr, binds serine. Chemoreceptors are homodimers (25) in which each subunit has a transmembrane domain of two transmembrane segments, a periplasmic, ligand recognition domain and a cytoplasmic, signalling domain. There is substantial identity among the amino acid sequences of the four receptors in their cytoplasmic domains but little in their periplasmic or transmembrane domains (13), yet mutational analyses of ligand binding imply that there is a common three-dimensional organization."
The Tap (taxis toward peptides) receptor and the periplasmic dipeptide-binding protein (DBP) of Escherichia coli together mediate chemotactic responses to dipeptides. Perhaps we could use this dipeptide signal in our system.
OR because the binding protein must first bind its ligand and then bind the receptor, perhaps we could tether this protein onto a cell surface expressed peptide or flagella or pilli or something via a cleavage site for the parasitic protease. Its receptor would exist on the chassis membrane and could only be activated in the presence of the parasite. An endogenous TCS pathway could be used in this scenario, coupled with our engineered output. As it would be the binding protein which activates the receptor, we would have to supply to the medium its ligand (Ribose etc)so that it is primed to bind its receptor upon release from the surface. Need to check the feasibility of whether the PBP can pass through the membrane as it's quite large?
a mutant Ribose binding protein has been used in iGEM before ref EnvZ-Periplasmic Binding Peptide Receptors
Periplasmic binding proteins: a versatile superfamily for protein engineering REF
Chimeric chemoreceptors in Escherichia coli: signaling properties of Tar-Tap and Tap-Tar hybrids. [6]
List of previous iGEM teams that have used EnvZ
- Valencia 2006 Main Wiki Page, System diagram including EnvZ fusion protein
- ETH Zurich 2006 PowerPoint Presentation
- Edinburgh 2009 - TNT sensing
- Tokyo 2009 Light-receptor
There is more, but can't remember which.
OmpR
OmpR is a molcule phosphorylated by the EnvZ.
Crystal structure of its parts is:
Split-Protein Sensors
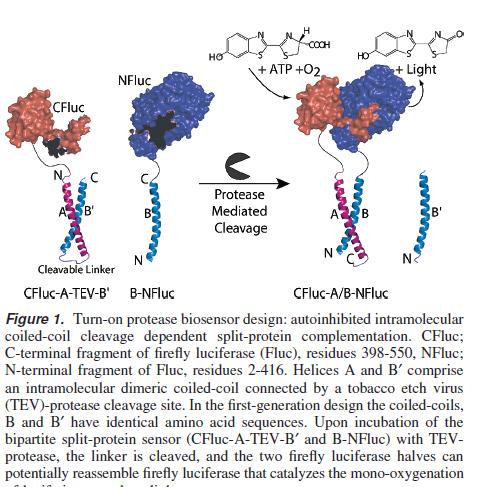
An Autoinhibited Coiled-Coil Design Strategy for Split-Protein Protease Sensors Paper
For an amplification step after the initial transcription event has occured, these sensors containing enzyme and or proteases could exist in a deactivated state within the cytoplasm, to be activated by TEV protease cleavage.
This application would form our system output module. The modularity comes from the engineered linker region separating two halves of a protease/enzyme containing any cleavage site but also the split enzyme/ protease itself can be changed. Split proteins that exist and have been widely used include Luciferase, Beta-lactamase, TEV protease amongst others.
QSS 2CS
The gram positive QSS we are hoping to use is from the agr-QSS locus of S.aureus. This has been used before in iGEM by Cambridge 2008. The HK (AgrC) activates the RR (ArgA) which binds the DNA to activate transcription of downstream genes. The system relies on AIP (auto-inducing peptide) which the cell produces and utilises as an external signalling peptide. We would add our specific protease tag onto the base of the cell surface displayed AIP such that it can be cleaved off when parasitic proteases are present in the environment. Agr-QSS
an article on AgrC: [Transmembrane topology and histidine protein kinase activity of AgrC, the agr signal receptor in Staphylococcus aureus.]
Com-QSS of E.coli is the other system that we have considered.
Image taken from 'The desk encyclopedia of microbiology' By Moselio Schaechter, Joshua Lederberg
Alternative approaches to activating a non-functional protease
Alternative approach to fusing the split protease directly to two PPI RRs could be to use interacting proteins that have been used before see Section 2.4and fuse these onto the RRs. C1 and C2 coiled coils are an example. The advantage would be that these constructs have been made and the resulting heterodimer produces a functional protease.
Allosteric Activation of DegS, a Stress Sensor PDZ Protease paper
Evolution of the ssrA degradation tag in Mycoplasma: Specificity switch to a different protease. paper In the same way Slovenia iGEM 2008 used a split ubiquitin tagging enzyme
there is a AAA+ Lon protease which would cleave at a tagged site (between two fret pairs for e.g?)
Phosphate-inducible proteases
In order to create a fast responding system we are hoping to bypass a translation step. Therefore ideally we are looking for a protease which will exist in the cell in a deactivated state. Upon our selected stimulus the protease will become activated at the end of the 2CS relay. So far alternative scenarios for fast activation include using phosphorylation. Two proteases that work by this mechanism include Caspase 9 1 and PAP 2
The problem with these two proteases include PAP being a serine protease and Caspase 9 being part of an elaborate signalling mechanism which would be hard to re-engineer. Some work on PAP is being carried out by Alia Cloutier-Bosworth (BSc student)who is attempting to characterize the protease.
Alternative system without downstream protease + FRET
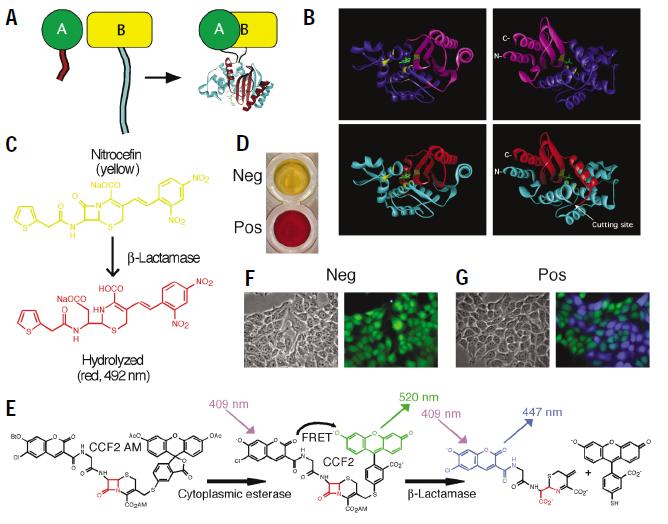
β-lactamase and β-galactosidase
Using β-lactamase and or β-galactosidase with cephalosporin nitrocefin and X-Gal respectively
They have split β -lactamase: spit B-lactamase and this hydrolyses chromogenic substrate nitrocefin that changes from yellow to red. We would get a visual output. The system would involve having the bacteria in a medium which contains the nitrocefin, they’d take it up and so it would be present intracellularly. Only upon induction of our artificial 2CS in response to our parasite would the β-lactamase become active and hydrolyse the cephalosporin nitrocefin. We could either create a fusion protein of the enzyme onto receptors which dimerize or onto homo/hetero dimer cytosolic interacting proteins of the TCS.
Shannon, K., and I. Phillips. 1980. Beta-lactamase detection by three simple methods: intralactam, nitrocefin and acidimetric. J. Antimicrobe. Chemother.
Papers 1
Secondly in a similar concept: β-galactosidase can be split in two peptides, LacZα and LacZΩ, neither of which is active by itself but both spontaneously reassemble into a functional enzyme wiki this enzyme hydrolyses X-gal to create colourless to blue pigment. We would have to include X Gal in our bacterial solution, but again the system would only become active upon QSS/TCS activation and dimerization of receptor/ downstream components (achieved by parasitic protease cleavage of a cell surface peptide.)
Papers which have played around with β-galactosidase fragment fusions (im still looking into this!! But I thought to put up what I’ve found so far) ref
Previous iGEM team which re-engineered and utilized the β-galactosidase enzyme to produce a colour change: Synthetic biology: Engineering Escherichia coli to see light paper
No idea of feasibility….but ‘Use of a protein scaffold’: engineer a protein scaffold that contains SH2 and another homology type binding domain e.g. PDZ SH3 (some scaffolds naturally exist ‘POSH’ etc) If we attach an SH2 homologous domain to one half of the split protease and modify the SH2 binding domain to have a phosphor-tyr in the consensus sequence, one half of the split protease would be attached to this site. The other half could be fused to PDZ/SH3. Through receptor tyrosine kinase signalling (some RTKs exist in bacteria) the SH3 binding domain could become phosphorylated allowing the SH3-split protease to bind adjacent the already bound half allowing the two halves to interact and reform. Phosphorylation would occur upon RTK activation, it would require modification/ creating a chimeric receptor that could respond to a signal that we choose, unless an endogenous signal exists which we can manipulate or use. RTKs in bacteria ref
Outcome of Research
We explored several avenues for combining a fast response module with a two component system:
We have looked into a number of Quorum sensing systems from Gram positive bacteria. We identified ComABCDE system from S.pneumonae as one of the few systems that uses a small peptide molecule that is linear and does not undero any post-translational modifications. This is an important requirement if we are to attach this signalling peptide to a protein that is localised to the cell wall. This in turn will allow us to present the peptide for cleavage by the paracite elastase.
Our initial hope was to by-pass transcription entirely and bring together a Split protease when signalling molecules dimerise on phosphorylation. As the two halves come together that could produce a functional protease which could then activate a response mechanism.
Unfortunately this required complex protein engineering the outcome of which was dependent on many unknown factors making it too unpredictable for an iGEM project. This part of the project had to be dropped in favour of more traditional transcriptional activation, the module should still be faster as the output is produced by a pre-made inactive enzyme that is activated by a transcribed protease.
The output module relies on substrate being pre-made in the cell and triggered into active form by protease cleavage or being brought together by dimerisation of a signalling protein.
We have considered a number of options, pros and cons of which are shown in the Output module section.
References
A useful review on quorum sensing. look at the middle of the article for gram positive bacteria Quorum sensing]
Cell wall peptidoglycan architecture in Bacillus subtilisref
http://www.ncbi.nlm.nih.gov/Complete_Genomes/RRcensus.htm Response regulators encoded in bacterial and archaeal genomesl The MiST (Microbial Signal Transduction) database uses a domain-based approach for the identification of signal transduction molecules and represents a comprehensive electronic catalog of signaling components in microbial genomesMiST P2CS Database
phoP ompR dimers really good paper on the fusion of FPs and dimerization of numerous RRs also info on hetero pairs.
Some papers on the Structural Biology of (re-engineering)2CS receptors
1. Cheung J, Bingman CA, Reyngold M, Hendrickson WA,Waldburger CD: Crystal structure of a functional dimer of the PhoQ sensor domain. J Biol Chem 2008, 283:13762-13770.
2. Cheung J, Hendrickson WA: Crystal structures of C4–dicarboxylate ligand complexes with sensor domains of histidine kinases DcuS and DctB. J Biol Chem 2008,283:30256-30265.
3. Zhou YF, Nan B, Nan J, Ma Q, Panjikar S, Liang YH, Wang Y,Su XD: C4–dicarboxylates sensing mechanism revealed by the crystal structures of DctB sensor domain. J Mol Biol 2008, 383:49-61.
4. Reinelt S, Hofmann E, Gerharz T, Bott M, Madden DR: The structure of the periplasmic ligand-binding domain of the sensor kinase CitA reveals the first extracellular PAS domain. J Biol Chem 2003, 278:39189-39196.
paper on protease assays(may be useful later)
Design principles of the bacterial quorum sensing gene networks