IGEM:IMPERIAL/2009/Assays Protocols/WetLab Plan
From OpenWetWare
Jump to navigationJump to search
WetLab Plan
Current Plan --David Roche 09:57, 6 August 2009 (EDT)
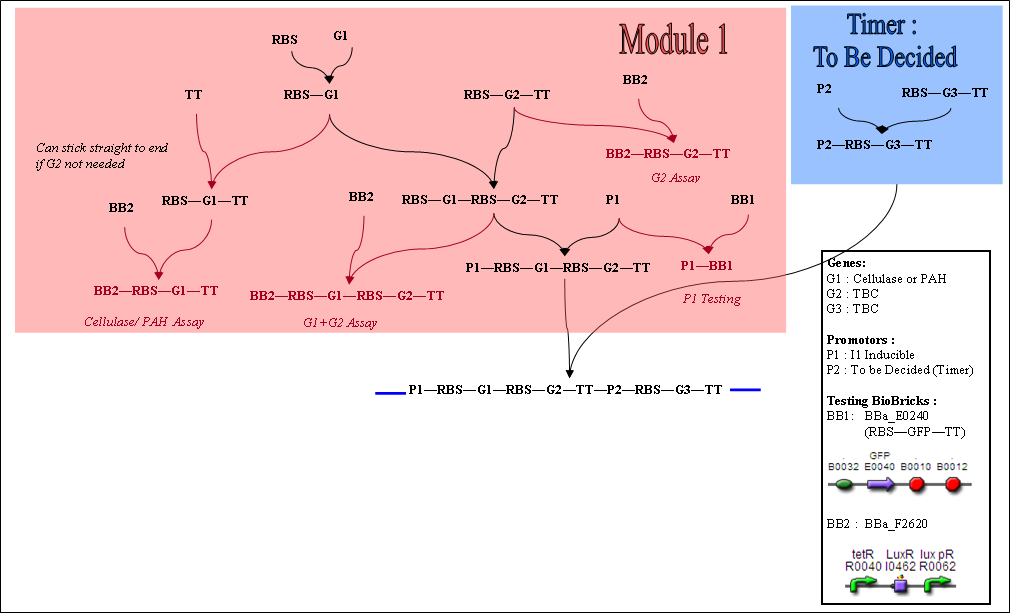
These are including the testing constructs, which are shown in italics. Two standard BioBricks are used for testing - BBa E0240 (BB1) and BBa F2620 (BB2).
- BB1 consists of GFP production device, and can be used to characterise the strength of promoters - both constitutive and inducible.
- BB2 is a promoter sequence, which can give a variable POPs input to a gene sequence of interest, depending on the concentration of HSL inputted. Combining this with a quantitative output assay, we hope to be able to characterise our own BioBricks.
- We have also chosen to use the pBad inducible promoter for the testing of our restriction enzyme constructs. This is because it is tightly regulated, and so this will be useful for the testing.
Updates
- The Testing Constructs and Steps required to reach them are shown in colour. The necessary steps are shown in black.
Module 1
Links to Relevant Assays
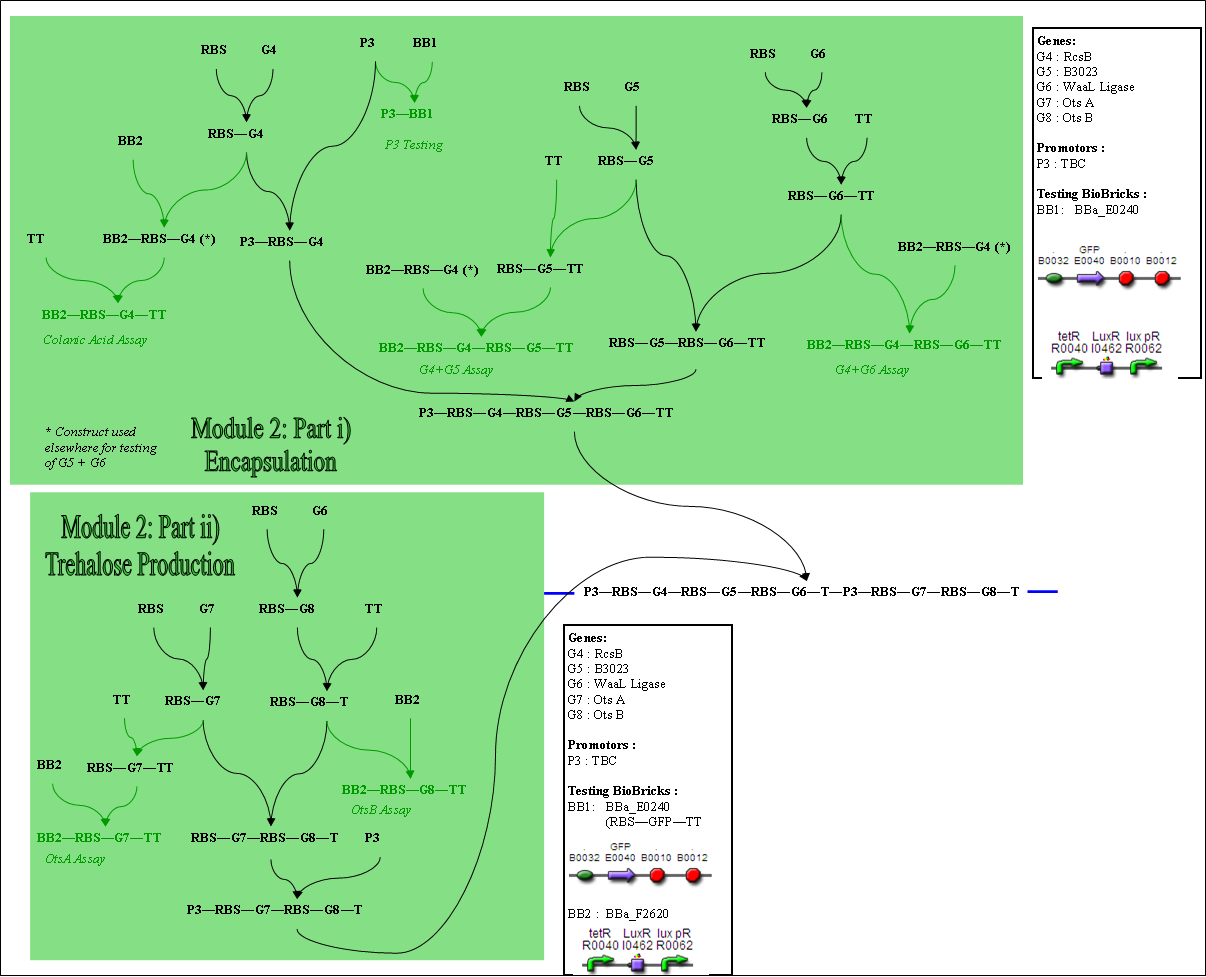
Module 2
Links to Relevant Assays
Colanic Acid Assay: (Testing of RcsB) Testing of G4+G5 and G4+G6 can be done using the same assay.
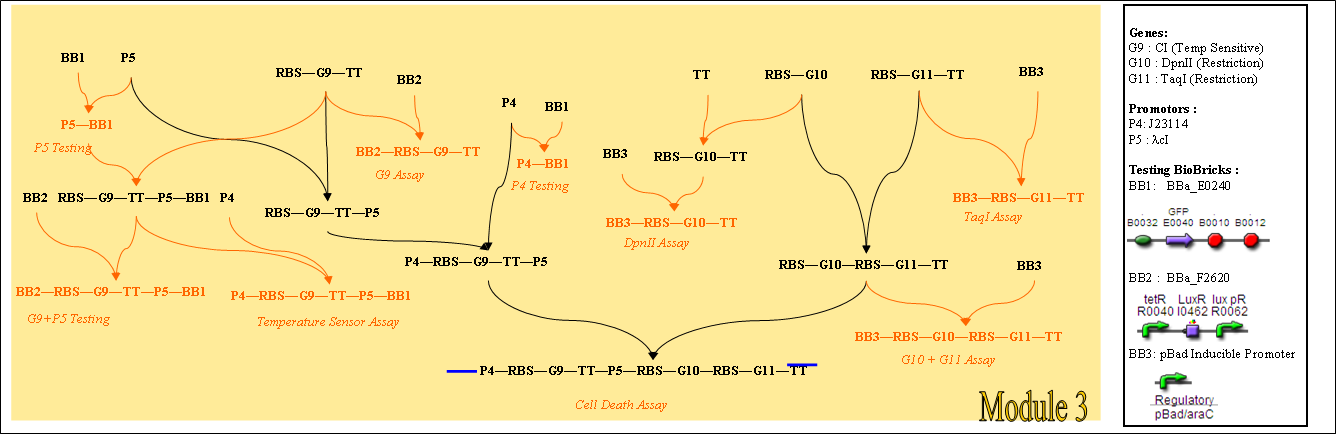
Module 3
Links to Relevant Assays
Heat Induction Assays
Cell Death Assays
- Dam testing should be performed to check whether the resultant methylation of the DNA is sufficient to protect from the basal levels of restriction enzyme.
- Firstly, testing should be done with the restriction enzymes on a Dam -ve strain. In this case the basal levels of restriction enzymes should cleave the DNA, as there is no protection.
- Then, a test should be performed with the same Dam -ve strain, but with our Dam BioBrick inserted. This should result in the expression of Dam, and consequently the methylation of the DNA. This should be sufficient to protect the genetic material from cleavage due to basal levels of restriction enzyme expression.
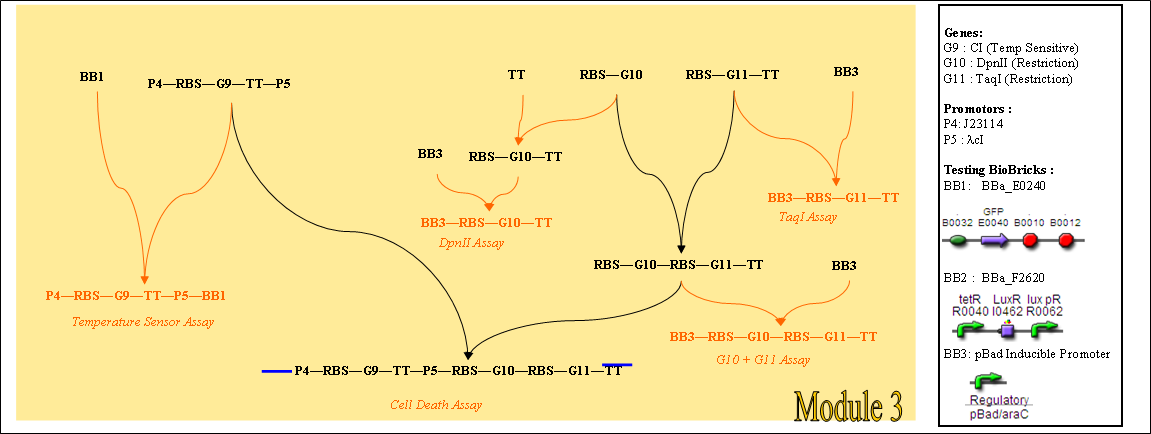
Alternatively, if we can isolate a well characterised temperature sensor BioBrick, several steps can be skipped, giving a cloning strategy as shown below.