Biomod/2013/IIT-Madras/Results
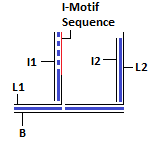
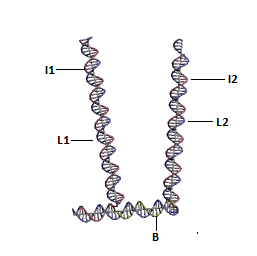
Structure of DNA Walker
PAGE Results
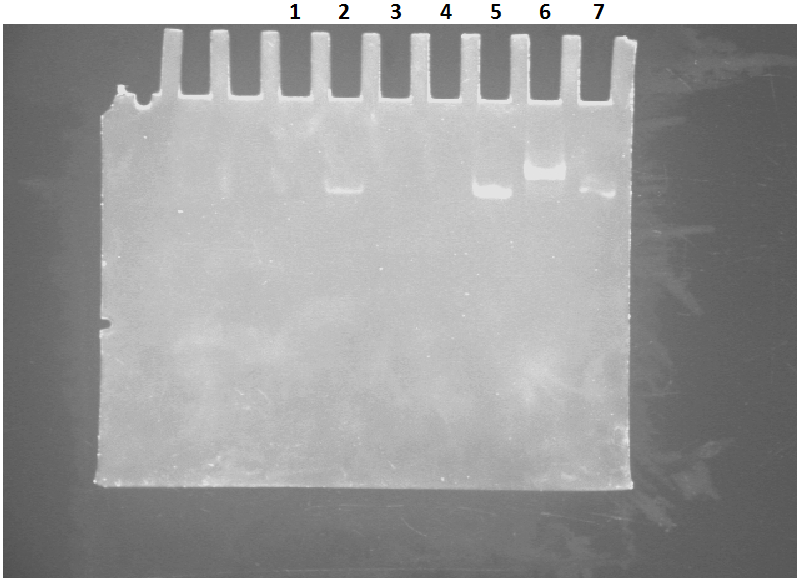
The lane for I1L1 shows a band higher than I1 and L1 lanes. However, both bands for I1 and L1 can not be at the same level.
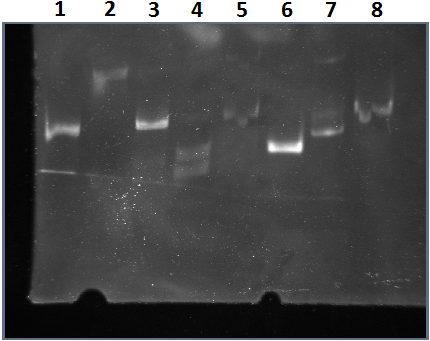
The lane for BI1I2 shows multiple bands, indicating the absence of secondary structures.
The band for I1L1 is significantly higher than L1 and I1. The difference between I1 and L1 lanes can be seen.
Sequence Design
The sequences for the strands L1 and I1 have been taken from the following paper[1]. The sequence of the bases was rationally designed using sequence symmetry minimization taking into account the constraint posed due to the fixed i-motif sequence using the ideas given in the following paper[2].
Programs were written in Python to design the sequencesLink. They do not take into consideration energetics/thermodynamics. The results of the program were then fed to NUPACK for secondary structure analysis (at Na+ concentrations of 0.05 owing to the absence of Na+ in the annealing mixture)Link.
The sequence set has been selected as it formed the desired structures with high affinity.Link
The NUPACK result of B+I1+I2 showed the absence of interactions between the three strands. This confirms the efficiency of strand design
The formation of the desired structure requires I1L1 and BI2L2 complexes to be formed with high stability.
The NUPACK result for I1L1 complex showed 100% formation of the structure even at temperatures of 40C Link. Analysis of strands B, L2, I2 showed high turnover to form the annealed complex too Link.
References
-
Two DNA nanomachines map pH changes along intersecting endocytic pathways inside the same cell Souvik Modi, Clément Nizak, Sunaina Surana, Saheli Halder & Yamuna Krishnan, Nature Nanotechnology 8, 459–467 (2013) doi:10.1038
-
Seeman NC, Kallenbach NR. Design of immobile nucleic acid junctions. Biophys J. 1983;44(2):201–209. doi: 10.1016/S0006-3495(83)84292-1 [1]