Biomod/2013/IIT-Madras/Project
Background
DNA walkers are nanomachines which exhibit linear motion along a DNA strand. They ware initially constructed with the aim of mimicking the intra-cellular motion of molecular motors such as kinesin so that they could serve as mechanical parts for future Nano-bots. Their structure mainly consists of two strands - a moving strand and a stationary strand along which the moving strand "walks". The major components of the DNA walker system are[1][2]:
- Walker : The moving strand
- Track : The stationary strand
- Attachment strands/Set strands : These hybridize to single stranded segments on the walker and track strands thereby linking them.
- Detachment strands/Unset strands : They have high affinity for the set strands and displace them via toehold mediated strand displacement. This causes the walker strand to move along the track.
There have been several modifications to this construct depending on the strategy of actuation used[1][2][3][4][5][6][7].
In this project, we demonstrate the displacement of DNA in response to stimuli such as pH change.
History of DNA Walkers
DNA Walkers in the past have primarily used other synthetic DNA oligonucleotides[1][2] in order to achieve motion along a track. Other actuation strategies include the use of light[6][7] and restriction enzymes[4][3].
These systems have been extremely effective in controlling motion along a track. However, they may not be able to function in the endogenous cell environment ie. without the addition of "Extra" synthetic biomolecules.
Idea!
In an attempt to overcome this, we embarked upon designing a DNA machine that responds to changes in cell environment by "hopping" along the track.
We intend to control the DNA Walker using
- pH
- Light - Our aim is to protect the recipient DNA strand with photo-activable groups. The binding of the walker strand with the recipient can occur only if both pH and light triggers are present. We have used a short DNA Strand as a proxy for caging the DNA with photo-activatable groups[8]
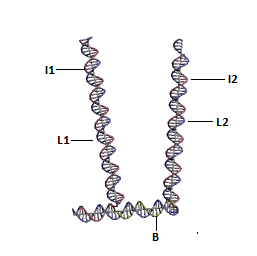
Resting State of DNA "Hopper" - High pH ~8
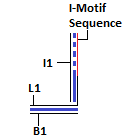
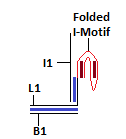
Module I: pH Switch
The strand L1 is involved in the formation of the intra-molecular i-motif at low pH[9]. This will result in the exposure of an overhang of strand I1. This can interact with strand L2 as their sequences are complementary.
The sequences for the strands L1 and I1 have been taken from the following paper[10]. There are a few mismatches in strand I1 (the complement to the I-Motif sequence). These have been incorporated in order to prevent G-quadruplex formation at lower pH. This was done in order to reduce the chances of formation of a secondary structure that would interfere in FRET between the fluorophores in the paper[10]. The duplex below the I-Motif sequence was incorporated to prevent the strands from separation under low pH.
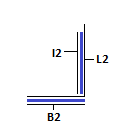
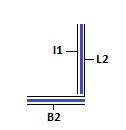
Module II: Receptor/Detector
This module acts as receptor for strand I1 and allows detection of the same. The strand L2 can be caged at dA and dC residues by NDBF (photo-activatable groups)[8]. This will ensure that walking/I1L2 binding occurs only in the presence of light. This approach does not require the binding of I2 and can be done using fluorophores instead. Due to the unavailability of light sensitive caged DNA, we have used a short stretch of complementary DNA, I2 instead. In true spirit, the strand L2 is not caged by I2 as I1 will bind to L2 always. This will enable detection of walking of I1 to the receptor module. Strand I2 is shorter than I1. The unpaired region serves as a toehold for I1 to bind and displace I2 via toehold mediated strand displacement. This can be detected using low-pH Polyacrylamide Gel Electrophoresis. The formation of the above mentioned structure need not occur if all the strands are added into solution as I1 whas greater affinity to L2 as compared to I2. Hence, I1L1 and BL2I2 complexes must be annealed and mixed to obtain the final structure.
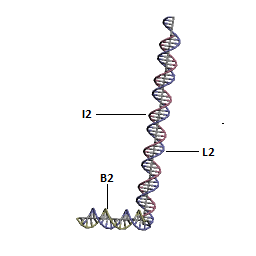
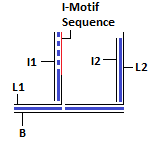
Final Structure of DNA Walking Platform
These modules are linked to form the DNA Walking Platform. 2 nucleotides are inserted between the base and other double helices in order to make the joints flexible. The sequence of the bases was rationally designed using sequence symmetry minimization taking into account the constraint posed due to the fixed i-motif sequence using the ideas given in the following paper[11].
DNA "Hopper" in motion - low pH ~5
At Low pH (pH<5.5), L1 folds into an intra-molecular i-motif due to the formation of CHC+ (hemi-protonated Cytosine base pairs). This folding doesn't release the Input (I1) into the solution, but assists toehold mediated strand-displacement to module 2.
Module 1
Module 2
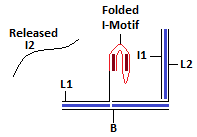
Complete Picture :: DNA "Hopping"
Translation of Input (I1) from Module 1 to Module 2 triggered by pH (change in cellular environment). This causes release of I2 from the receptor region.
Note:Blue line represent normal Watson-Crick base pairing. Brown lines represent abnormal or Hoogsten base pairing.
Experiment Design
The experiments that we have designed for demonstration of DNA walking can be found here Link.
References
-
A Precisely Controlled DNA Biped Walking Device William B. Sherman and and Nadrian C. Seeman, Nano Letters 2004 4 (7), 1203-1207 [1]
-
A Synthetic DNA Walker for Molecular Transport Jong-Shik Shin and and Niles A. Pierce, Journal of the American Chemical Society 2004 126 (35), 10834-10835 [1]
-
Yin, P., Yan, H., Daniell, X. G., Turberfield, A. J. and Reif, J. H. (2004), A Unidirectional DNA Walker That Moves Autonomously along a Track. Angew. Chem., 116: 5014–5019. doi: 10.1002/ange.200460522
-
Bath, J., Green, S. J. and Turberfield, A. J. (2005), A Free-Running DNA Motor Powered by a Nicking Enzyme. Angew. Chem. Int. Ed., 44: 4358–4361. doi: 10.1002/anie.200501262
-
Molecular Gears: A Pair of DNA Circles Continuously Rolls against Each Other, Ye Tian and and Chengde Mao, Journal of the American Chemical Society 2004 126 (37), 11410-11411[1]
-
Building a Nanostructure with Reversible Motions Using Photonic Energy Mingxu You, Fujian Huang, Zhuo Chen, Ruo-Wen Wang, and Weihong Tan, ACS Nano 2012 6 (9), 7935-7941 [1]
-
You, M., Chen, Y., Zhang, X., Liu, H., Wang, R., Wang, K., Williams, K. R. and Tan, W. (2012), An Autonomous and Controllable Light-Driven DNA Walking Device. Angew. Chem. Int. Ed., 51: 2457–2460. doi: 10.1002/anie.201107733
-
Wavelength-Selective Uncaging of dA and dC Residues Florian Schäfer, Khashti Ballabh Joshi, Manuela A. H. Fichte, Timo Mack, Josef Wachtveitl, and Alexander Heckel [1] Organic Letters 2011 13 (6), 1450-1453
-
pH-Induced Intramolecular Folding Dynamics of i-Motif DNA Jungkweon Choi, Sooyeon Kim, Takashi Tachikawa, Mamoru Fujitsuka, and Tetsuro Majima, Journal of the American Chemical Society 2011 133 (40), 16146-16153 [1]
-
Two DNA nanomachines map pH changes along intersecting endocytic pathways inside the same cell Souvik Modi, Clément Nizak, Sunaina Surana, Saheli Halder & Yamuna Krishnan, Nature Nanotechnology 8, 459–467 (2013) doi:10.1038
-
Seeman NC, Kallenbach NR. Design of immobile nucleic acid junctions. Biophys J. 1983;44(2):201–209. doi: 10.1016/S0006-3495(83)84292-1 [1]