Biomod/2011/TUM/TNT/Methods/Data Analysis
<html>
<style>
- column-one { display:none; width:0px;}
.container{background-color: #f5f5f5; margin-top:50px} .OWWNBcpCurrentDateFilled {display: none;}
- content { width: 0px; margin: 0 auto auto 0; padding: 1em 1em 1em 1em; align: center;}
- column-content {width: 0px; float: left; margin: 0 0 0 0;padding: 0;}
.firstHeading {display:none; width:0px;}
- globalWrapper{ width:1280px;}
div.tleft { border-bottom-width: 0em; border-left-width: 0px; border-right-width: em; border-top-width: 0em; clear: left; float: left; margin-right: 0.5em;
}
div.tnone { border-bottom-width: 0em; border-left-width: 0px; border-right-width: em; border-top-width: 0em; }
- column-one {display:none; width:0px;background-color: #ffffff;}
- content{ margin: 0 0 0 0; padding: 1em 1em 1em 1em; position: center; width: 800px;background-color: #f5f5f5; }
.container{ width: 800px; margin: auto; background-color: #ffffff; text-align:justify; font-family: helvetica, arial, sans-serif; color:#555555; margin-top:25px; }
- bodyContent{ width: 1267px; align: center; background-color: #ffffff;}
- column-content{width: 1280px;background-color: #f5f5f5;}
.firstHeading { display:none;width:0px;background-color: #ffffff;}
- header{position: center; width: 800px;background-color: #ffffff;}
- footer{position: center;}
div.thumb { border-bottom-color: rgb(255, 255, 255); border-bottom-style: solid; border-left-color: rgb(255, 255, 255); border-left-style: solid; border-right-color: rgb(255, 255, 255); border-right-style: solid; border-top-color: rgb(255, 255, 255); border-top-style: solid; margin-bottom: 0.5em; width: auto;
}
div.tright { border-bottom-width: 0em; border-left-width: 0px; border-right-width: 0px; border-top-width: 0em; clear: right; float: right;
}
body {
font-family: helvetica, arial, sans-serif; color: black;
background:#f5f5f5;
}
ul#topnav {
margin: 0 0 0 0px ; padding: 0 0 0 0px; float: left; width: 800px; list-style: none; position: relative;
font-family: helvetica, arial, sans-serif;
font-size: 12px; background: #f5f5f5;
height: 5px;
} ul#topnav li { float: left; margin: 0; padding: 0; border-right: 1px solid #ffffff; /*--Divider for each parent level links--*/ } ul#topnav li a { padding: 11px 15px; display: block; color: #333333; text-decoration: none; } ul#topnav li:hover { background: #f5f5f5;} /*--Notice the hover color is on the list item itself, not on the link. This is so it can stay highlighted even when hovering over the subnav--*/
ul#topnav li span { float: left; padding: 15px 0; position: absolute; left: 0; top:30px; display: inline; /*--Hide by default--*/ width: 800px; background: #cccccc; color: #333333; /*--Bottom right rounded corner--*/ -moz-border-radius-bottomright: 5px; -khtml-border-radius-bottomright: 5px; -webkit-border-bottom-right-radius: 5px; /*--Bottom left rounded corner--*/ -moz-border-radius-bottomleft: 5px; -khtml-border-radius-bottomleft: 5px; -webkit-border-bottom-left-radius: 5px; } ul#topnav li:hover span { display: block; } /*--Show subnav on hover--*/ ul#topnav li span a { display: inline;} /*--Since we declared a link style on the parent list link, we will correct it back to its original state--*/ ul#topnav li span a:hover {text-decoration: underline;}
- abstractandlinks{
display:block; width:800px; background-color:#f5f5f5; text-align:justify;font-family: helvetica, arial, sans-serif; color:#555555; margin-top:0px; }
- innerBox
{ display: block; padding: 5px; }
- abstract{
font-family: helvetica, arial, sans-serif; width: 490px; margin-top: 20px; padding:0px 5px; float: left;background-color:#f5f5f5; ; }
- links{
font-family: helvetica, arial, sans-serif; width: 290px; margin-top: 20px; padding:0px 5px; float: left; background-color:#f5f5f5;; } .clear { clear:both; }
- tour{
font-family: helvetica, arial, sans-serif; width: 790px; padding:0px 5px; margin-top: 1em, margin-bottom: 1em; float: left;background:#f5f5f5; font-size: 2em; }
- struktur{
font-family: helvetica, arial, sans-serif; width: 790px; padding:0px 5px; float: left;background:#f5f5f5; }
- toctitle {align: left; text-align: left; }
- table.toc {background-color:"#5f5f5f";align: left; text-align: left;}
- toc #toctitle, .toc #toctitle, #toc .toctitle, .toc .toctitle {
text-align: left;
}
- toc, .toc, .mw-warning {
background-color: #f5f5f5; border-bottom-color: rgb(170, 170, 170); border-bottom-style: solid; border-bottom-width: 0px; border-left-color: rgb(170, 170, 170); border-left-style: solid; border-left-width: 0px; border-right-color: rgb(170, 170, 170); border-right-style: solid; border-right-width: 0px; border-top-color: rgb(170, 170, 170); border-top-style: solid; border-top-width: 0px; font-size: 105%; padding-bottom: 5px; padding-left: 5px; padding-right: 5px; padding-top: 5px; align: left; text-align: left; display: left
}
</style>
<head>
<script type="text/javascript"> var fullDates = new Array('08/11/2011','08/12/2011','08/16/2011','08/17/2011','08/23/2011','08/24/2011','08/25/2011','08/29/2011','08/30/2011','08/31/2011','09/01/2011','09/02/2011','09/06/2011','09/12/2011','09/13/2011','09/14/2011','09/15/2011','09/16/2011','09/19/2011','09/20/2011','09/21/2011','09/27/2011','09/28/2011','09/29/2011','09/30/2011','10/05/2011','10/06/2011','10/07/2011','10/10/2011','10/12/2011','10/13/2011','10/14/2011','10/17/2011','10/18/2011','10/19/2011','10/20/2011','10/21/2011','10/26/2011','10/27/2011','10/28/2011','10/30/2011');
$(document).ready(function() {
$("ul#topnav li").hover(function() { //Hover over event on list item $(this).css({ 'background' : '#cccccc'}); //Add background color and image on hovered list item $(this).find("span").show(); //Show the subnav
} ,
$("ul#topnav li").oneclick(function() { //Hover over event on list item $(this).css({ 'background' : '#cccccc'; 'display' : 'inline'}); //Add background color and image on hovered list item $(this).find("span").show(); //Show the subnav
} ,
function() { //on hover out...
$(this).css({ 'background' : 'none'}); //Ditch the background
//$(this).find("span").hide(); //Hide the subnav });
}); </script>
<script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.3/jquery.min.js"></script>
<meta http-equiv="Content-Type" content="text/html; charset=utf-8" />
<script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.4.2/jquery.min.js"></script> <script type="text/javascript"> $(document).ready(function(){ $('#lside1').mouseover(function() { $('#lside1').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside1').mouseleave(function() { $('#lside1').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); $('#lside2').mouseover(function() { $('#lside2').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside2').mouseleave(function() { $('#lside2').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); $('#lside3').mouseover(function() { $('#lside3').stop().animate({backgroundPosition: '0px 0px'}, 1000); }); $('#lside3').mouseleave(function() { $('#lside3').stop().animate({backgroundPosition: '-250px 0px'}, 1000); }); }); </script>
<style type="text/css">
#side1 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside1 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
#side2 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside2 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
#side3 {
width: 250px;
height: 100px;
position: relative;
top: 0px;
left: 20px;}
#lside3 {
width: 250px;
height: 100px;
background: url(http://www.tftshop.net/Testbild_100ppi.jpg) -250px 0px no-repeat;
position: absolute;
top: 0px;
left: 0px;}
</style>
<title>
TUM NanU - Home
</title> </head>
<body background-color="#F0F0F0">
</body> </html>
Data Analysis
http://openwetware.org/index.php?title=Biomod/2011/TUM/TNT/Methods/Data_Analysis&action=edit
Thermal fluctuation of the arms
Introduction
In order to estimate how broad a broad spread of the data points around the mean angle in TEM angle measurements must be expected, we calculated the width of a Boltzmann distributed fluctuation of the arms. Hereby we assumed the DNA helices to be cylinders with a certain stretch modulus E, persistence length p and to fluctuate independently. Within these measurements we always quantified the angle between the end points of the arms end the other end (at the base).
Derivation
Fluctuation of one arm
The probability of finding a rod with a 10 helix bundle cross section at a certain angle [math]\displaystyle{ \phi }[/math] is given by a Boltzmann distribution
[math]\displaystyle{ p(\phi - \phi_0) = \frac{1}{Z} e^{- \frac{E(\phi - \phi_0)}{k_B T}} }[/math]
where the energy is given by
[math]\displaystyle{ E(r ) = \frac{1}{2} \frac{E_{DNA} I_{10 hb} L}{r^2} }[/math]
with r the radius of curvature, L the contour length of the arms and I10 hb the area moment of inertia of a 10 helix bundle. If we assume a constant curvature of the arms, the radius of curvature can be translated in an angle [math]\displaystyle{ \phi }[/math] (see Fig. 1).

[math]\displaystyle{ 2 \phi = 2π \frac{L}{2\pi r} }[/math]
[math]\displaystyle{ E(\phi- \phi_0) = \frac{1}{2} \frac{4 E_{DNA} I_{10 hb} (\phi - \phi_0)^2}{ L } }[/math]
The area moment of inertia of a rod is given by
[math]\displaystyle{ I = \frac{R^4}{4} \pi }[/math]
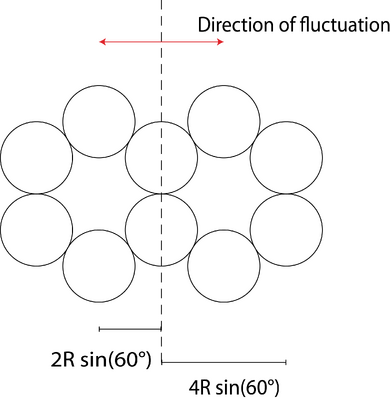
Now we're able to calculate the area moment of inertia of the 10 helix bundle with the parallel axis theorem (see Fig. 2).
[math]\displaystyle{ I = \sum_n (I_n + {z_n}^2 A_n) = I_{DNA} (10 + 320 sin (\frac{\pi}{6})) }[/math]
and therefore the energy needed to bend one arm by the angle [math]\displaystyle{ \phi }[/math] as well as the probability.
Fluctuation of the measured angles
Since the fluctuation of the arms is stochastically independent, we have to take the convolution of the two probabilities:
[math]\displaystyle{ p(\Delta\phi) = \int p_1 (\phi_1 ) p_2 (\Delta\phi - \phi_1) d\phi_1 }[/math]
with [math]\displaystyle{ \Delta\phi = \phi_2 - \phi_1 }[/math]
With all this follows
[math]\displaystyle{ p(\Delta\phi) = \frac{1}{Z^2} \sqrt{\frac{π k_B T}{E_{eff}}} e^{-\frac{E_{eff}}{2 k_B T}(\Delta \phi - \phi_0)^2} }[/math]
with
[math]\displaystyle{ E_{eff} = \frac{2 E_{DNA} I_{DNA} (10 + 320 sin(\frac{\pi}{6}))}{L} }[/math]
Therefore we get a standard deviation of the measured angles of
[math]\displaystyle{ \sigma = \sqrt{\frac{k_B T}{E_{eff}}} = \sqrt{\frac{L}{2 p (10 + 320 sin(\frac{\pi}{6}))}} }[/math]