Biomod/2011/PUCP/Wiracocha/Results
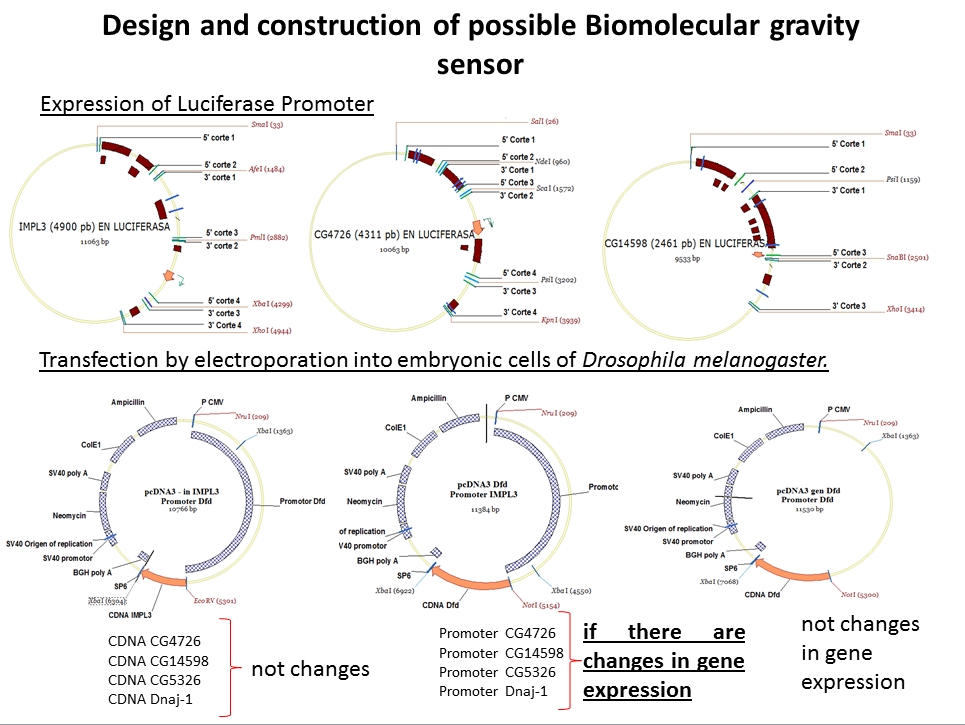
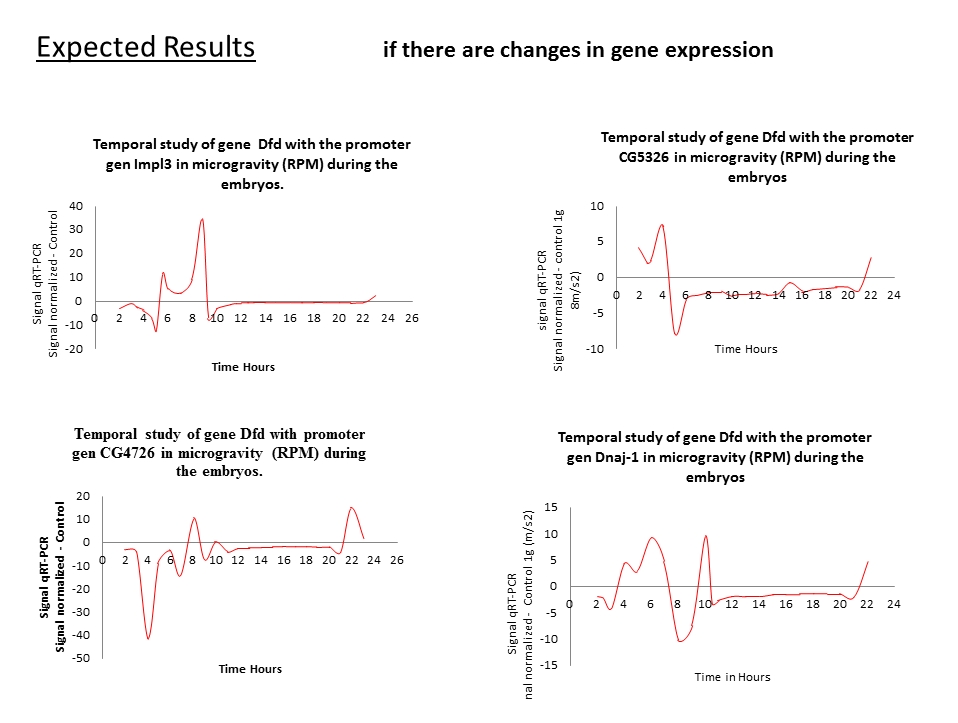
This study shows the proposed method, to demonstrate that regulators factors that have been detected (Laván, David al et., in Press - Thesis) (See general methodology and procedure to be followed) are not the foundation for a process of gene transcription, in microgravity. The study presented is divided into two stages (may be developed in parallel). The first is to demonstrate the possible areas (sequence DNA) that we identified are conserved evolutionarily in the genes IMPL3, CG4726, CG14598, CG5326, Dnaj-1 y CG18431 (called overlapping genes). These areas contain regulatory factors that help the transcription process. These possible regulatory factors are common in the six overlapping genes. The study consists in cloning 2500 bp of the upstream area and the downstream or first intron, of all the six gene, in of the luciferase vector pxp2.seq of 6163 bp. To know which or what are the areas that contain possible regulatory factors sites, we will make serial sections containing the conserved regions of each of the genes under study. To then, the sequence are cloning into the luciferase vector pxp2. Once having obtained the construction of each of our plasmids, We proceed to transfection by electroporation in cells of Drosophila melanogaster. As control we use the betha-galactosidase Assay System with Report Lysis Buffer of Promega. For quantification of luciferase the Luciferase Assay system of Promega protocol will be used. Having identified the conserved regions, each of the six overlapping genes, we will identify the possible binding sites for regulatory factors. For it we will use the EMSA protocol. EMSA, is important technique for studying gene regulation and determining protein. The second study of this proposal is to demonstrate experimentally that the potential regulatory factors that we have detected do not play an important role in microgravity conditions. The study is to express the cDNA of the six genes under study, they are activated by the regulators which are in the upstream areas of the promoter gene Dfd, in microgravity conditions and normal gravity (The gene Dfd expression levels do not change in microgravity, also the Dfd is a possible regulatory factors of the under study genes, (Laván, David al et, in Prees – Thesis)). To quantify your results, the reporter gene to be used is the pcDNA3.1. Once our plasmids are designed we will proceed to transfection into cells of Drosophila melanogaster. We will measure two quantitative variables, the first of these is the mRNA expression levels by qRT-PCR and the second variable to measure is the amount of protein that has been generated, using Western Blot techniques. The control group was performed under the same conditions as the experimental group, with the difference that is subject to conditions of normal gravity.
General methodology and procedure to be followed
Biochemical study
Oligo design and Taqman probes
1. Design of the area downstream and upstream of promoter of the gene
2. Design of successive cuts in the area downstream and upstream of promoter of the gene under study
3. Semiquantitative study of expression levels mRNA of the genes under study (IMPL3, CG4726, CG14598, CG5326 and DnaJ-1) For semiquantitative studies, the plasmids were designed using oligos that recognize the sequence around the cDNA for each of the genes under study. In the case of those whose length cDNA was greater than 1000 bp it fragment into two parts and then be united.
Name of the Oligos Sequence 5' to 3'
IMPl3 5' atggccgccattaaggacagtc
IMPl3 3' ttagaacttcagaccagcctggacatc
CG14598 5' atgtcactgaaatacttactcattgcttgtgc
CG14598 3' ctacagtctgacgaagcgtggaaaag
Dnaj-1 5' atgggcaaagacttctacaagattctgggc
Dnaj-1 3' ctagttgggcagcagctcggacagc
CG5326 5' atgtcggtcagactgaacgagacga
CG5326 3' ctactcctttttcgccgccagc
CG4726 5' atgacggacgaagtgcccgc
CG4726 3' ttagggcttagttggcagctccttc
CG4726 5'_Corte atgcacgtgatgatcgcctcc
CG4726 3'_Corte cccaacaagcagcgtttcatca
4. Quantitative study of mRNA expression levels of genes under study (IMPL3, CG4726, CG14598, CG5326 and DnaJ-1).
Name of the Gene Exon Length Stock Sequence
IMPL3 _03 - _04 93 Dm01841229_g1 ggcgaacatggcattgacaaggatgtgttcctct cgctgccctgcgttctcaatgccaacggtgtgacat ccgtggtcaagcagatcctgact
CG4726 _1 - _2 64 Dm01800475_m1 gcaaactcgtgcccgcccgctatgtgctggccctcc tggggtccatcggcatggccattgtgta
CG14598 2 134 Dm02368409_s1 aaaagtggaagaagcgcaaaattaattcctaaaa taatatttatcgtaaaggaaattcatacacccttctt agcgaccaatggatagcgttttaccttttccagagc ttcatcggcatgtttttaattgtagta
CG5326 _03 - _04 81 Dm02143291_g1 tggctcgtgccgtgtggctgtactacattgccaagat cacggagctgttggacaccgtgttctttgtgctgcgca agaaac
Dnaj-1 1 101 Dm02362419_s1 aacacggggaggttacatcagttagtagaacacttat agatttatacgccgtaaggacatagtacgttttagtac caattctattatgttataaacaataa
For the qRT-PCR will be requested to AB Applied Biosystems, the following probes:
5. Analysis of common regulatory factors
5.1 Collection of pupae with 4.5 days of development.
5.2 RNA extraction using Trizol.
5.3 Checking in silico of the different cuts of the areas upstream and downstream of the promoter.
5.4 Checking the design of primers for different DNA amplifications.
5.5 Amplification of DNA fragments by PCR from each of the sections downstream and upstream of the promoter.
5.6 Ligation of each one of the fragments using restriction enzymes.
5.7 Checking the fragments linked by digestion techniques. (Results observed with techniques of agarose gel electrophoresis).
5.8 Cloning of all promoter regions in the luciferase vector.
5.9 Transformation in bacteria Dh5alpha.
5.10 Purification of plasmid by miniprep techniques and maxiprep following the protocol of Qiagen.
Transfection
5.11 Check that the primers to be used for transfections are well designed
5.12 Amplification of DNA fragments by PCR.
5.13 Ligation of DNA fragments
5.14 Cloning of each of the fragments in the Luciferase vector
5.15 Transformation in bacteria Dh5alpha.
5.15 Purification of plasmid by miniprep techniques and maxiprep following the protocol of Qiagen.
5.16 Checking the fragments linked by digestion techniques. (Results observed with techniques of agarose gel electrophoresis).
5.17 Transfection of the plasmids by electroporation into cells of Drosophila melanogaster.
5.18 Quantification of Luciferase. The protocol experiment group is: Luciferase Assay system of Promega and the group control is Betha Galactosidase Enzyme Assay System with reporte Lysis Buffer of Promega.
5.19 Study of factors of regulation by ENSA.
6. Expression of mRNA of genes under study, by means of the region downstream of the promoter Dfd.