Biomod/2011/Harvard/HarvarDNAnos:Protocols
<html> <head>
<style>
- column-one { display:none; width:0px;}
.container{background-color: #f5f5f5; margin-top:50px} .OWWNBcpCurrentDateFilled {display: none;}
- content {width: 0px; margin: 0 auto auto 0; padding: 1em 1em 1em 1em; align: center;}
- column-content {width: 0px; float: left; margin: 0 0 0 0;padding: 0;}
.firstHeading {display:none; width:0px;}
- globalWrapper{width:1280px; margin:auto}
body {background: #F0F0F0 !important;}
- column-one {display:none; width:0px;background-color: #f0f0f0;}
- content{border:none;margin: 0 0 0 0; padding: 1em 1em 1em 1em; position: center; width: 800px;background-color: #f0f0f0; }
.container{ width: 800px; margin: auto; background-color: #f0f0f0; text-align:justify; font-family: helvetica, arial, sans-serif; color:#f0f0f0; margin-top:25px; }
- bodyContent{ width: 1267px; align: center; background-color: #f0f0f0;}
- column-content{width: 1280px;background-color: #f0f0f0;}
.firstHeading { display:none;width:0px;background-color: #f0f0f0;}
- header{position: center; width: 800px;background-color: #f0f0f0;}
- footer{position: center; width:1280px;}
</style>
</head> </html>
<html>
</html>
Laboratory Equipment
Transmission Electron Microscope (TEM)
Sample Prep
- 2% aqueous uranyl formate stain solution (3 mL H20, 0.06 g uranyl formate):
- Uranyl formate (EMS, powder, store with parafilm over lid because seems to degrade in air)
- Weigh out 0.06 g uranyl formate in 10 mL beaker, using the balance in the hood
- Put in magnetic stir bar
- Put on magnetic stirrer under large inverted foil-topped beaker (don't stir it yet)
- Using a pipette, measure out 3 g H20 + 40 uL H20 = 3 mL H20 + 40 uL H20 (nuclease free, Ambion) in another 10 mL beaker
- Boil this water on a hot plate (NOT the same stirrer used for the uranyl formate) and shut off heat right when it boils
- Can carry over to the formate solution on the stirrer with gloves even though the water beaker is hot
- Pour hot water into formate beaker and turn on magnetic stirring; cover with large inverted foil-topped beaker
- Glow discharging the grids
- Formvar/Carbon coated grids (SPI # 3440C)
- Grab glass slide (e.g., VWR micro slides), gently wrap a piece of parafilm (e.g., 1.5 squares long, 1 sq wide) around a section of the glass slide
- Using tiny tweezers, grab grids from the edge and transfer to the parafilm wrapping on the slide
- Tweezers are Dumont N4AC from EMS. Buy your own tweezers; they break easily. Have one tweezer per grid if doing multiple grid preps in parallel.
- Put in EMS glow discharger. Settings: 25 mA, 45 s, 0.1 mBar, negative HT polarity. Press "start".
- Putting sample on grid and staining
- Materials: 15 mL falcon tube (wrap in foil), filter (Acrodisk, 0.2 um) that mounts on 5 mL syringe tip, 5 mL syringe (BD)
- Draw stain solution into the syringe, screw on filter, discharge through filter into fresh 15 mL tube
- Add 1 mL of uranyl formate solution into a fresh 2 mL eppendorf tube using a pipette
- Add 5 uL of 5N NaOH (JT Baker - this is DANGEROUS - DO NOT GET INTO EYES) into the 1 mL of stain solution in the eppendorf
- Vortex; the solution should get slightly darker
- Use a grid mat in a petri dish
- Line up tweezers, 1 tweezer per grid
- Grab grid edge and let rest on tweezers, suspended over air
- Pipette 3.5 uL of sample onto the grid for 4 min
- Take a piece of whatman paper, bend it in the middle
- Wick off sample by bringing whatman paper in contact with the grid edge from the side
- Immediately add 3.5 uL of stain solution onto the grid and let sit for 1 min
- Wick off the stain as before
- Leave for a minute or two to dry
- Transfer grid to mat at an angle: As grid nears mat surface, open tweezers slightly--the grid will still stick to the tweezers, and drag up to a ridge on the mat, at which point the grid will detach from the tweezers and rest up against the ridge on the mat
- It is crucial not to bend the grid or exert any forces on it
- Put stuff that contacted uranyl formate in the radioactive waste bin
Imaging your sample
- Include instructions from Worksheet
Atomic Force Microscope (AFM)
Sample Prep
- Use scotch tape to remove a thin layer of mica from an AFM puck
- Apply 5 uL of 5x buffer and then 5 uL of your sample to the mica; let deposit
- Place puck into magnetic receptacle in AFM
- Carefully add an additional 30 uL of buffer to the mica, making sure not to get liquid onto the AFM
- To "turn on" the AFM, flip switch from STM to AFM
- Fluid Cell and Wafer
- Using tweezers, grab the bottom of a SNL-10 wafer (each has 4 cantilevers with a tip each) and place it into the curvy trough receptacle of a fluid cell
- Secure the wafer by pushing up on the spring, turning the lever, and relaxing the spring, so that it holds the wafer against the fluid cell
- Using tweezers, pick up the fluid cell by a hole on the side, and invert it into the AFM opening; let it settle into the grooves
- Using knob on back-center of AFM, depress the electrodes onto the cell
- Turn on Veeco light
- Adjust laser using knob on the top-near-right and the knob on the top-far-middle so that it hits cantilever, and adjust the mirror slider on the back, maximizing the SUM signal to at least 5
- Adjust the detector using the knob on the top-far-left and the knob on the back-left so that VERT and HORZ are near 0 (+/- 1 is okay)
Imaging your sample
- Open the Nanoscope program; use "ScanAsyst in Fluid"
- Press "Engage" until the tip finds the surface; then "Withdraw" and re-"Engage" a couple of times
- Allow the image to scan
- Generally, 512 samples/line, 0.977 Hz scan rate, 3-30 nm height scale bars, and 1-5 uL scan size is a good place to start
- Alternatively, you can use "TappingMode in Fluid"
- First press "Tune," then "Auto Tune"; wait for straight line
- Set "Sweep Width" to 10 kHz to make sure you're on the upward slope of a nice resonance peak
- Press "Engage" and continue as usual
- Add more fluid if the sample is getting dry (water needs to extend from the puck to the fluid cell)
Dynamic Light Scattering Device (DLS)
Gel Box
Preparing/Running an Agarose Gel
- Notes: Should add magnesium after microwaving since it reacts with the agarose when heated. Good to cool off the gel mixture slightly before pouring so that it does not warp the plastic. Good to run gel in ice water bath to prevent overheating.
- Running buffer (11 mM MgCl2 in 0.5X TBE): add 0.5X TBE, keep track of how much you added since you will need to add the proportional amount of 1M MgCl2 (11 mL per 1 L of 0.5X TBE); or use the pre-prepared buffer (with Mg already added)
- Gel (2% agarose, large MgCl2 gel): 2.4 g agarose in 600 mL beaker, 0.5X TBE up to 120 g, microwave 2 minutes, swirl. Add 1.3 mL of 1 M MgCl2 and 6 uL of 10 ug/mL EtBr for final concentration of 0.5 ug/mL. Or use 8 uL SYBR-Safe (Invitrogen) in place of the EtBr. Pour and let harden in cold room with comb.
- Loading buffer:
- add to 10 uL of sample: 3 uL 5X LA loading medium, 2 uL H2O
- add to 5 uL of sample: 2 uL 6X AGLB lite, 4 uL H20
- pipette up and down once and then load into gel lane
- DNA ladder (1kb)
- Distilled water - 400 μl
- 6X Blue Loading Dye (600 uL glycerol, 150 uL 1% wt/vol orange G, 220 uL dH2O, 30 uL 1% wt/volume xylene cyanol) - 100 μl
- DNA Ladder from NEB - 100 μl
- Total volume - 600 μl
- Add about 15 uL to ladder lane
- Run at 80V for ~ 2 hours before imaging using the gel imager
- Excision from the gel: use a fresh razor blade on top of plastic wrap on top of safe imager, first excise lane, then band, both vertically and in-plane if possible, giving yourself enough room; take a picture first
- Filtration and recovery with "Freeze 'n Squeeze":
- Crush gel slice w/ pestle in a 1.5 mL eppendorf tube
- Spin for 1 minute at 4000 rcf (for 3 tubes, positions 1, 9 and 17 to balance the centrifuge) at 4C
- cut off tube bottom a few mm above the 100 uL mark, invert it into spin column (actually put the cut off bottom in there upside down) [Bio-Rad 732-6165]
- You can use the tube cutter from USA Scientific
- Spin for 3 minutes at same settings
- Take out the filter, trash it, and close the tube!
- Want around 2 nM concentration, so may need to dilute in 1X folding buffer (10X folding buffer diluted 1:10 in water)
Preparing/Running a Polyacrylamide Gel
- Use 10% TBE (no Urea) gel - take off tape on bottom, and take out the comb on top
- Put into gel box, printed words facing outside of box (wells facing in)
- Put in blank on other side
- Fill with 0.5x TBE buffer to cover both of the electrodes
- Middle section up to top
- Outer section 1/3 the way up
- Clear lanes of gel fluid by using squeeze pipette to displace the gel fluid with buffer
- Load lanes with samples - if possible leave a spacer lane between ladder and samples
- Use 0.5 uL ladder, 20 uL water, and 2 uL loading dye for the ladder lanes; pre-mix and load 11 uL of this solution into each ladder lane
- Use 10 uL sample + 2 uL (of 6X) loading dye for sample lanes
- Attach box head, insert box head wires into electrical source, and run the gel at specified voltage and for specified amount of time. usually, when the yellow dye band (from green loading dye) has reached the bottom of the gel, it should be done.
- After electricity is turned on, you should see small bubbles flowing from the bottom of the gel to the top due to electrolysis of the buffer
- Staining with SYBR-Gold: 5 uL of 10,000X SYBR-Gold in 50 uL water, for 30 mins on slow shaker
Typhoon Gel Imager
- Use gloves with the Typhoon; don't use gloves with the computer
- Pour staining liquid out of staining tub; fill the tub with just enough water to keep the gel wet
- Pull out the Typhoon scanning plate
- Using two hands, gently pull the bottom of the gel against a wall of the tub and slide it up the wall until you can lift the gel (with both hands) out of the tub
- Place the gel on the Typhoon scanning plate squarely, and make note of the region on which it is sitting; insert the scanning plate back into the machine
- On the computer, select the folder in which to save the images; give the image a file name (be sure not to overwrite previously scanned files!)
- Select the region on which the gel is sitting; select the staining agent that was used; adjust the PMT voltage (higher will give darker bands); choose a pixel size
- Press "Scan"; as the image scans, you can adjust the levels
- After the image is done scanning, pressing "Return" on the scanning console will automatically save the image
- Remove the gel and discard; clean the scanning plate with ethanol, water, and Kimwipes
Thermocycler
Centrifuge
Slow Shaker
Nanodrop
- The NanoDrop is a "Micro-Volume UV-Vis Spectrophotometer for Nucleic Acid and Protein Quantitation"
- For 5 nm gold nanoparticles:
- Open the Nanodrop 2000 program
- Click on the "UV-Vis" button
- Adjust wavelength to 520 nm and the pathlength to 1 mm; uncheck "Use cuvette"
- To blank: load 1.5 uL phosphine buffer directly onto the pedestal and lower the arm; press the "Blank" button
- Clean the pedestal with water and a kimwipe
- Load 1.5 uL of your sample onto the pedestal and lower the arm; press "Measure"
- Divide the A520 measurement by appropriate extinction coefficient to obtain concentration in uM
- For oligos:
- Open the Nanodrop 2000 program
- Click on the "Nucleic Acid" button
- Choose the "DNA" option from the dropdown menu in the right panel
- Set the baseline correction to 340 nm (see this reference for why), and uncheck "Use cuvette"
- Proceed as you would for nanoparticles, using water or TE buffer (or whatever you used to resuspend the oligos) to blank
- The measurement that you get is in units of ng/uL; to get concentration in uM, multiply this number by (1000 * moles/g) or by (nmoles/mg * 1/1000) which is molecular weight data you can get off the oligo spec sheet
DNA Origami Assembly
Resuspending Oligos in Solution
- Download ResuspendingDNA.xlsx
- Input requested data from order sheets contained with packing info into Columns B & D (note: for desired concentration in Column D, indicate a concentration that is double your "final desired concentration" as to minimize error and the possibility for over-dilution)
- Add specified volume of DD/DI water to samples as specified in Column F
- Vortex your samples for 15 seconds and microcentrifuge for 90 seconds
- Verify the concentration of your sample using the NanoDrop by selecting Home-->Nucleic Acid-->ssDNA, then blanking with DD or DI water, then beginning to measure samples (taking two measurements for each sample)
- Input the above data into Columns G & H, and then again add the specified volume of DD/DI water, as specified in Column Q, to your sample.
- Vortex your samples for 15 seconds and microcentrifuge for 90 seconds
- Re-nanodrop samples and input data to verify final concentrations
Basic Folding Protocol
- Assume starting with:
- 100 nM scaffold stock
- Applied Biosystems Buffer Kit including:
- 0.5 M EDTA pH 8
- 1 M MgCl2
- 1 M Tris pH 8.0
- DEPC treated water
- Staples at 100 uM concentration
- 1) Make 10X folding buffer (50 mL) -- 50 mM Tris, 10 mM EDTA, 80 mM MgCl2
- 2.5 mL 1 M Tris stock
- 1 mL 0.5 M EDTA
- 4 mL 1 M MgCl2 stock
- Add DI water to 50 mL
- 2) Make oligo pre-stocks (corresponding to different staple pools)
- First spin down plates from manufacturer for 30s at 1000g
- There are i = 1 to N_groups pre-stocks
- For each i, #i is # of staples in group i
- Take 10 uL of each oligo from group i and put in pre-stock i
- 3) Make working stock (mixture of staples from the different pre-stocks)
- For each i, combine the following in a single tube: #i uL from pre-stock i
- Each oligo now has concentration 100 uM / (total # of oligos) within the working stock
- 4) Make folding mixture (here for a 50 uL reaction). Combine in a PCR tube:
- 5 uL 10X folding buffer
- (0.075 uL)*(total # oligos) from the staple working stock
- 15 uL scaffold stock
- Nuclease free water up to 50 uL total volume
- These should give approximate 150 nM concentration of each staple (5 fold excess), and 30 nM scaffold concentration, assuming staples start at 100 uM concentration from the synthesis company and that the scaffold stock is at ~ 100 nM.
- 5) Run the annealing rxn (here for a 6hb nanotube). Run rxn mix in PCR machine with following annealing program:
- 1: 80C for 5 min
- 2: 80C for X min
- -1C per cycle
- Goto 2 60 times
- End
- For a 2D structure, X = 2 minutes will work
- Making 100 nM scaffold stock
- Measuring scaffold concentration on the nanodrop:
- 1.5 uL on the nanodrop, nucleic acid mode, custom extinction coeff = 37
- Nanodrop outputs a value in ng/uL. Convert from ng/uL to nM using spreadsheet that takes into account exact molecular weight of scaffold (see 2010-08-18 folder which has spreadsheet with this value for many of our scaffolds). Nanodrop reading varies over time (due to evaporation?). For concentrated samples, may be wise to consider the actual concentration value as ~ 10% lower than the nanodrop reading converted to nM.
- Measuring scaffold concentration on the nanodrop:
Making 2D Origami
- 10x origami folding buffer (50 mL) -- 50 mM Tris, 10 mM EDTA, 125 mM MgCl2
- 2.5 mL 1 M Tris stock
- 1 mL 0.5 M EDTA
- 6.25 mL 1 M MgCl2 stock
- Add DI water to 50 mL
- Folding mixtures for origami:
- There are ~ 224 staples in the tall rectangle, depending on removal of edge staples for de-stacking etc. Using a multichannel pipette, take 5 uL from each plate well on your plates of staples (we're assuming you ordered your plates hydrated at 100 uM concentration) and combine in a 2 mL eppendorf tube. This is the staple stock.
- Thus for a 50 uL reaction we can use:
- 5 uL 10X folding buffer
- (0.15 uL)*224 = 33 uL from the staple working stock
- 10 uL scaffold stock
- 2 uL nuclease free water (optional, since the exact concentrations don't matter that much)
- These should give 300 nM concentration of each staple (15 fold excess), and ~ 20 nM scaffold concentration, assuming staples start at 100 uM concentration from the synthesis company and that the scaffold stock (e.g., m13mp18 from NEB) is at ~ 100 nM. 15 fold excess is more than enough to get near 100% yield of excellent origami.
- Compare with Paul Rothemund's original protocol: "Staple and remainder strands were purchased unpurified ... in water at 100 uM or 150 uM and stored at -20C. The desired set of up to 273 short strands was mixed with M13mp18 (typically 160 nM of each short strand, 1.6 nM M13mp18 circular or linear, a 100-fold excess of short strands) in a 100 uL volume of 1X Tris-Acetate-EDTA (TAE) buffer with 12.5 mM magnesium acetate (pH=8.3) and annealed from 95C to 20C in a PCR machine (Eppendorf) at a rate of 1C/minute in .1C steps."
- NPRTF folding reaction for 2D origami (the whole anneal takes only ~ 1 hour):
- 95C for 5 min
- 90C for 1 min
- Cool to < 40C at 1 minute per degree (1 degree steps are OK)
- Take out the sample, or continue cooling gradually to room temperature
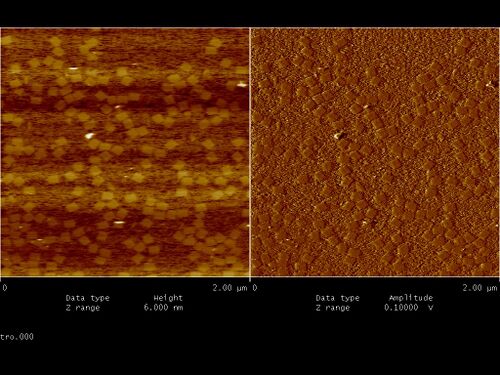
- Right after folding (with no purification) they look like this under fluid tapping AFM (see below):
Making 3D Origami with "Complex Curvatures" (e.g., a sphere)
- Based on Han et al.
- JSON file: Media:Sphere_match.json
- Excel staple file: Media:Sphere_staples.xlsx
- 1x buffer: 5 mM Tris, 1 mM EDTA and 16 mM MgCl2
- Formulation for 10x buffer: 2.5 mL of 1 M Tris, 1 mL of 0.5 M EDTA, 8 mL of 1 M MgCl2 and 38.5 mL H2O
- Scaffold: 10 nm single-stranded M13mp18 DNA
- Staples: 10 times molar excess in buffer
- Cycle: 94°C to 86°C at 4°C per 5 minutes, 85°C to 70°C at 1°C per 5 minutes, 70°C to 40°C at 1°C per 15 minutes, 40°C to 25°C at 1°C per 10 minutes
- Staple pools and reaction mix: Media:Sphere_wells.xlsx
Running Gels
Typhoon Gel Imager
Preparing/Running an Agarose Gel
- Notes: Should add magnesium after microwaving since it reacts with the agarose when heated. Good to cool off the gel mixture slightly before pouring so that it does not warp the plastic. Good to run gel in ice water bath to prevent overheating.
- Running buffer (11 mM MgCl2 in 0.5X TBE): add 0.5X TBE, keep track of how much you added since you will need to add the proportional amount of 1M MgCl2 (11 mL per 1 L of 0.5X TBE); or use the pre-prepared buffer (with Mg already added)
- Gel (2% agarose, large MgCl2 gel): 2.4 g agarose in 600 mL beaker, 0.5X TBE up to 120 g, microwave 2 minutes, swirl. Add 1.3 mL of 1 M MgCl2 and 6 uL of 10 ug/mL EtBr for final concentration of 0.5 ug/mL. Or use 8 uL SYBR-Safe (Invitrogen) in place of the EtBr. Pour and let harden in cold room with comb.
- Loading buffer:
- add to 10 uL of sample: 3 uL 5X LA loading medium, 2 uL H2O
- add to 5 uL of sample: 2 uL 6X AGLB lite, 4 uL H20
- pipette up and down once and then load into gel lane
- DNA ladder (1kb)
- Distilled water - 400 μl
- 6X Blue Loading Dye (600 uL glycerol, 150 uL 1% wt/vol orange G, 220 uL dH2O, 30 uL 1% wt/volume xylene cyanol) - 100 μl
- DNA Ladder from NEB - 100 μl
- Total volume - 600 μl
- Add about 15 uL to ladder lane
- Run at 80V for ~ 2 hours before imaging using the gel imager
- Excision from the gel: use a fresh razor blade on top of plastic wrap on top of safe imager, first excise lane, then band, both vertically and in-plane if possible, giving yourself enough room; take a picture first
- Filtration and recovery:
- Crush gel slice w/ pestle in a 1.5 mL eppendorf tube
- Spin for 1 minute at 4000 rcf (for 3 tubes, positions 1, 9 and 17 to balance the centrifuge) at 4C
- cut off tube bottom a few mm above the 100 uL mark, invert it into spin column (actually put the cut off bottom in there upside down) [Bio-Rad 732-6165]
- You can use the tube cutter from USA Scientific
- Spin for 3 minutes at same settings
- Take out the filter, trash it, and close the tube!
- Want around 2 nM concentration, so may need to dilute in 1X folding buffer (10X folding buffer diluted 1:10 in water)
Preparing/Running a Polyacrylamide Gel
- Use 10% TBE (no Urea) gel - take off tape on bottom, and take out the comb on top
- Put into gel box, printed words facing outside of box (wells facing in)
- Put in blank on other side
- Fill with 0.5x TBE buffer to cover both of the electrodes
- Middle section up to top
- Outer section 1/3 the way up
- Clear lanes of gel fluid by using squeeze pipette to displace the gel fluid with buffer
- Load lanes with samples - if possible leave a spacer lane between ladder and samples
- Use 0.5 uL ladder, 20 uL water, and 2 uL loading dye for the ladder lanes; pre-mix and load 11 uL of this solution into each ladder lane
- Use 10 uL sample + 2 uL (of 6X) loading dye for sample lanes
- Attach box head, insert box head wires into electrical source, and run the gel at specified voltage and for specified amount of time. usually, when the yellow dye band (from green loading dye) has reached the bottom of the gel, it should be done.
- After electricity is turned on, you should see small bubbles flowing from the bottom of the gel to the top due to electrolysis of the buffer
- Staining with SYBR-Gold: 5 uL of 10,000X SYBR-Gold in 50 uL water, for 30 mins on slow shaker
Using the Typhoon to Image a Gel
- Use gloves with the Typhoon; don't use gloves with the computer
- Pour staining liquid out of staining tub; fill the tub with just enough water to keep the gel wet
- Pull out the Typhoon scanning plate
- Using two hands, gently pull the bottom of the gel against a wall of the tub and slide it up the wall until you can lift the gel (with both hands) out of the tub
- Place the gel on the Typhoon scanning plate squarely, and make note of the region on which it is sitting; insert the scanning plate back into the machine
- On the computer, select the folder in which to save the images; give the image a file name (be sure not to overwrite previously scanned files!)
- Select the region on which the gel is sitting; select the staining agent that was used; adjust the PMT voltage (higher will give darker bands); choose a pixel size
- Press "Scan"; as the image scans, you can adjust the levels
- After the image is done scanning, pressing "Return" on the scanning console will automatically save the image
- Remove the gel and discard; clean the scanning plate with ethanol, water, and Kimwipes
Purifying DNA Origami
Gel Purification: "Squeeze 'n Freeze"
- Excise desired band from the gel
- Use a fresh razor blade on top of plastic wrap on top of safe imager
- First excise lane, then band, both vertically and in-plane if possible, giving yourself enough room
- Take a picture first
- Filtration and recovery:
- Crush gel slice w/ pestle in a 1.5 mL eppendorf tube
- Spin for 1 minute at 4000 rcf (for 3 tubes, positions 1, 9 and 17 to balance the centrifuge) at 4C
- Cut off tube bottom a few mm above the 100 uL mark, invert it into spin column (actually put the cut off bottom in there upside down) [Bio-Rad 732-6165]
- Alternatively, use a pipette tip to scoop crushed gel out of eppendorf tube
- Spin for 3 minutes at same settings
- Take out the filter, trash it, and close the tube!
- Want around 2 nM concentration, so may need to dilute in 1X folding buffer (10X folding buffer diluted 1:10 in water)
- 5 uM Tris
- 1 mM EDTA
- 8 mM MgCl2
Amicon Purification
- Courtesy of Ralf Jungmann
- Buffer = 1X folding buffer
- Amicon Ultra 0.5 ml Centrifugal Filters (100000 MWCO)
- Put folded origami solution into filter
- Add buffer solution to a total volume of 500 µl
- Put filter device into filtrate collection tube
- Spin for 5 min @ 14000 x g. This gives ~ 20 µl volume in the filter device
- Fill it up again to 500 µl total volume and repeat the last step. Make sure to empty the filtrate collection tube beforehand!
- After the second filtration step, turn the filter device upside down and spin for 3 minutes @ 1000 x g into a new filtrate collection tube
- Fill up to the original sample volume with buffer
Gold Nanoparticles
Using Gold Nanoparticles as Cargo attached to DNA Origami
- Synthesize or purchase AuNPs of a particular size
- Use the DLS Device to measure hydrodynamic size and extent of aggregation of the particles
- If the particles are mono-disperse and appear to be the appropriate size under DLS, continue to step 3
- If your solution appears to contain aggregates either return to step 1 or use a vacuum filter to remove the appropriate size range of particles and then remeasure using DLS
- Else, return to step 1
- Use TEM and ImageJ to calculate the average size of particles
- If the particles appear to be the appropriate size, continue to step 4
- Else, return to step 1
- Stabilize your particles by coating them with phosphene and then concentrating them in solution
- Conjugate your a particular DNA Strand(s) to your Gold Nanoparticles
- Purify your conjugates via Sucrose Gel Extraction
- Further stabilize particles via PEG or Poly-T Addition
- Attach your conjugated particles to your Origami structure
Synthesizing 5nm Gold Nanoparticles
Procedure adopted from Polysciences, Inc.: Technical Data Sheet #787, please see this sheet if you wish to make particles with either a smaller or larger diameter than 5 nanometers
- To make 100 mL of gold solution, two stock solutions have to be prepared and then mixed:
- Prepare Solution A
- 80 mL distilled water and 1 mL 1% aqueous gold chloride
- Prepare Solution B
- 4 ml 1% tri-sodium citrate. 2H2O + 16 ml H2O + 1 mL of 1% tannic acid
- Note: These two solutions must be made fresh each day!
- 4 ml 1% tri-sodium citrate. 2H2O + 16 ml H2O + 1 mL of 1% tannic acid
- Add 1 mL of 25mM potassium carbonate for pH adjustment
- Warm up solutions A and B to 60°C on a hot plate (or water bath) with a thermometer, and mix them while stirring with a magnetic stir bar
- Note: for fast mixing (which insures proper formation of particles, use a syringe to transfer solution B to solution A, therefore when preparing solution B you should make a little extra to insure you are able to add a total volume of 22 mL to solution A
- After initial mixing, allow the solution to reach 95°C (it will look like it is about to boil) and then cool the solution on ice until it appears to have returned to room temperature
- Then use the DLS device to analyze the size of your particles, under the intensity measurement they should appear to be between 9 and 11 nanometers in size (a measure of the hydrodynamic radius of the particles)
Using the DLS to Determine Size and Aggregation
- Load cuvette with room temperature sample to appropriate height after mixing, or your data may convey that your sample contains extremely large aggregates (the cuvettes available at the Wyss Institute either require 1mL or 40 uL of sample)
- Open the Zetasizer program on the computer, and open a new measurement file to record data [File, New, Measurement File]
- Create a new Standard Operating Procedure (SOP), if needed, to specify cuvette size and run conditions
- Run the appropriate SOP [Measurement, Start SOP] (at the Wyss Institute: "Colloidal Gold 1 mL Cuvette.sop" for 1mL cuvetter or "Gold Particle Project.sop" for 40 uL cuvette)
- Useful Notes:
- Forgetting to mix your samples can lead to misrepresented data; here is a comparison of the same sample before mixing and after mixing
- A poly-dispersity index (PDI) of less than or equal to 0.1 means that a given sample has a uniform peak (low aggregation)
- Intensity (first-pass measurement) is independent of Refractivity Index and Absorption (properties of the material) and is the most reliable measurement for the size of particles because it makes the least assumptions/inferences about the collected data
- Second-pass measurements can translate intensity to relative volume of the particles of different sizes, but are less accurate
- Each measurement will run between 10 and 30 trials
- Use this website to convert .xps files to PDFs (to convert the graphs that the program outputs to a readable file type)
Using TEM and ImageJ to Determine Average Size of Particles
- Add 3uL of a dilute sample of your gold nanoparticles (the solution should be a light pinkish color) to a copper grid for imaging under TEM
- Glow discharge isn't specifically necessary in preparing your samples, but may be helpful in the case of a dilute sample
- No staining is necessary, as AuNP alone provide enough contrast, additionally staining may lead to phosphene appearing in your TEM images, which would cause an over-approximation of the size of your particles
- You may either leave the sample until the water dries, or wick off remaining water after about five minutes, either procedure is valid but the first will result in more particles left on your grid (which is a good thing if you placed a highly dilute sample on the grid)
- Take multiple images of the particles at about 100,000x zoom that give you a view of approximately 500 particles:
- It is not essential to use 100,000x zoom, but you do need well focused images and a uniform zoom across all images
- Open your multiple images using stacks in ImageJ
- Set the scale on your images in ImageJ by:
- Zooming in
- Selecting the draw line tool
- Drawing a line over the scale bar
- Clicking analyze, then set scale, then typing into the dialog box the size of your scale bar in the appropriate unit of measurement
- Crop your image to remove the labels at the bottom
- Cropping one image will crop all
- Clear image of weird looking noise by selecting the region and then selecting edit, clear (you will have to do this on each layer)
- Set the contrast by selecting image, adjust, threshold, set contrast, then manipulating the slide bars until it appears that the majority of your particles are highlighted red, click apply, and then make sure that neither of the two checkboxes in the dialog box that appears are checked before proceeding to click okay
- Set the measurement to analyze the size of your particles by clicking analyze, set measurements, and deselecting all check boxes except fit ellipses and ferric diameter
- Analyze the size of your particles by selecting analyze, analyze particles, and then adjusting to an appropriate size range (as square units) that you are expecting for your particle (for 5nm, use 4-400 … aka a 2-20nm particle range), select exclude edges, display results, and show outlines, click OK
- Save the image obtained and export your data table to excel
- Determine the average size of your particles by taking the average of the major axes of your ellipses
- Determine the sphericity of your particles by dividing the average of your minor axes lengths by the average of your major axes lengths (1 will represent a perfect sphere)
- Useful Notes:
- Microsoft Excel 2011 does not come with the capability to create histograms, but this website provides an applet that allows you to do so
Stabilizing and Concentrating Gold Nanoparticles
- Take roughly 100 mL (80 mL is okay too) of gold nanoparticle solution
- Add 20 mg of phosphine powder, cover tube with aluminum foil, stir slowly for 12-24 hours
- Add about 10 g NaCl powder until the color changes from red to purple/blue, such that the NaCl concentration is roughly 3M in solution
- Centrifuge at 1000g in 2 50 mL conical tubes, splitting the volume equally between the two tubes
- After centrifuging you will see black powder attached to the sides of your tubes, and a clear-ish supernatant left in the tube; remove all but approximately the last mL of the supernatant using an electronic pipet
- Centrifuge your two tubes again for 5 mins at 1000g
- Add 2 mL of 2.5 mM phosphine buffer to each tube (66.9 mg phosphine powder in 50 mL H2O: see here)
- Resuspend black powder (your nanoparticles) by pipetting or manually swirling, solution will be red
- Add 5 mL methanol to each tube, swirl, and then combine your two tubes into one tube
- Centrifuge at 1000g for 30 minutes, remembering to balance your single tube with an extra tube filled with the appropriate volume of water
- Remove supernatant with a pipet and re-suspend your particles 1 mL of 2.5 mM phosphine and transfer to 1.5 mL tube, which you should cover in foil and refrigerate
- Useful Notes:
- After stabilized, this solution can sit at 4C in the fridge for several months
- The stability is light sensitive, so the particle tubes should be covered in foil whenever possible to prevent degradation
Conjugating Gold Nanoparticles to DNA
This is applicable for attaching 1 or a few DNA strands per nanoparticle. This is a draft protocol based on one from Chenxiang Lin, with additional info from Shawn Douglas and Steve Perrault.
- Use the average size of your particles to determine your extinction coefficient using the internal file provided by Steve Perrault
- In general, a 5nm particle has an extinction coefficient of about 10^7
- Use the nanodrop, under a UV-Vis A520 setting to determine the concentration with comparison to your extinction coefficient
- Before measuring, blank the nanodrop with 2.5 mM phosphine buffer
- Use the nanodrop, under the nucleic acid setting, to determine the concentration of your oligo to be added to your nanoparticles
- Blank your sample with distilled water
- Using your above acquired concentrations, mix your oligo in a 1:1 ratio with your nanoparticles
- Add 10x TBE to get ~0.5x TBE final concentration (20x dilution)
- Add e.g. 5M NaCl to get 50 mM NaCl final concentration (100x dilution)
- Leave for 40 hours on slow shaker, covered in foil
- Useful notes:
- Under the UV-Vis setting, absorbance levels should fall between .1 and 5; if the absorbance comes in above 5, dilute your sample until you get a reading within the .1-5 range
- Save some of these unconjugated AuNPs for comparison with the conjugated ones on your gel
- Another protocol for AuNP-Gold conjugation can be found here: PDF
Nanoparticle Sucrose Extraction
This is a draft protocol courtesy of Steve Perrault.
- Pour a gel for sucrose extraction
- Pour 50 mL of 4% agarose to produce a base, without a comb
- Once set, pour the top using 140mL of 3% agarose
- Use standard 0.5X TBE for these, no Mg is needed
- Use an appropriate comb with wells taped to give you enough space for the volume of product.
- You don't need to use DNA loading buffer, rather add 1/5 volume of 40% sucrose solution to increase the density, and add it to the gel
- Include unconjugated and conjugated lanes; you can load up to 1 mL of sample per lane.
- Run the gel at 100V for however long you need to get good separation. Again, it helps to start with a highly concentrated solution as you can see the bands really nicely.
- If you're using a short DNA strand (e.g., 15-mer), then the fastest band is generally BOTH conjugated and unconjugated nanoparticles. Higher bands are just aggregation in that case. If you're using a long strand (e.g., 60-mer), then the 1st band is unconjugated and the 2nd band is monoconjugated which is what we want...
- Once finished, you're going to cut the gel for sucrose extraction, but only the top of the gel, leaving the bottom base intact. Cut a well in front of the band that you want to extract. It should be slightly wide (5mm) on either side, and about 1 cm wide in the direction of electrophoresis. Then, make cuts behind the band all the way up to the top, so that you remove all the other bands and wells. You should have just a single bridge of agarose with your band in it, with a pool of buffer behind.
- Add the sucrose solution into the well in front of the band (0.5X TBE and I think with 30 or 40% sucrose, check with Chenxiang).
- Run the gel for 5-10 min intervals, keeping an eye on the band. The gold NP make it really easy to see when you need to extract the solution in the well and add new. We typically find that ~8 min, 100V intervals work well.
- Do this as many times as necessary to retrieve the entire band product. Purify it and do a buffer exchange with water using an Amicon column. I recommend doing at least 2X good washes with H2O.
- Finally, you should do absorption spectroscopy, EM and DLS to characterize your product. I've found (and Wei told me this) that having an NaCl solution of ~15 mM is necessary in your EM prep, to keep the particles from aggregating as they dry on the grid. DLS should show a population of (hopefully) monodispersed particles in the 10nm range, but that depends on the ligand you added to the surface. This characterization is important b/c it will give you a comparator for down the road, if you think your particles might be aggregating.
- For long term storage, you can powder fractions of the product using the freeze-dryer, replace the atmosphere with Argon and throw them in the freezer. They'll be good for a long time that way, probably only weeks to a couple months otherwise.
- Determine their concentration using absorption. I have a function that will tell you the extinction coefficienct of AuNP based on their size, but it's at work so I can't get it to you until I'm back. If they look pretty close to 5 nm by TEM (the cores), you can use the coefficient provided by Sigma or Ted Pella.
- Useful Notes:
- The gel-based purification is more efficient (higher recovery) if you start with a higher concentration of a smaller volume of product. This allows fewer wells to be used. Anything up to about 1 mL can be put into a gel but if you can get it down to 100-200 uL, the job will be easier. The solution can be concentrated by freeze-drying or by use of an Amicon column.
Adding PEGs to the surface of Gold Nanoparticles
- Re-measure the concentration of your Gold Nanoparticles and then determine the number of particles you have in solution
- Calculate the average surface area of your particles given the diameter you measured
- Determine what concentration of PEGs must be added to your solution such that there are 10 PEGs per square nanometer of each gold nanoparticle
- Add the specified amount of thiolated polyethylene glycol (PEG) molecule to your solution, thus stabilizing your particles with their attached oligo for use in DNA Origami
- Useful Notes:
- It is also possible to use a thiolated poly-t in place of a PEG
- On TEM, AuNPs with PEGs appear to have halos
Other
Photocleavable Spacers
- AmberGen protocol for photocleavage of PC-Biotin (same as for internal photocleavable spacer): "Photocleavage can be achieved with near-UV light and the recommended light source is a 365 nm peak Blak-Ray Lamp (Model XX-15, UVP, Upland, CA) or similar, used at a 5 cm distance for 5-15 minutes (ideally with mixing/shaking). The power output under these conditions is roughly 3 mW/cm2 at 360 nm, 1 mW/cm2 at 310 nm and 0.2 mW/cm2 at 250 nm. With this light source, most detergent and protein carriers in the photocleavage buffer are compatible, presuming they do not significantly absorb at the peak wavelength. Photocleavage through clear, preferably thin-walled, polypropylene tubes/vials (e.g. PCR tubes) is possible, since polypropylene transmits the necessary light. Shorter photocleavage wavelengths are possible, such as from conventional UV transilluminators, and in fact, PC-biotin absorbs maximally at ~275 nm. Light transmission through buffers and barriers should always be verified."
- Adapted protocol by Evan
- Place 10 uL of sample into thin-walled PCR tube
- Place PCR tube flat on the reaction surface (i.e. on surface of the pipette tip box, not in a hole) in the reaction chamber
- Insert the UVP UVGL-59 Handheld UV Lamp (Cambridge, UK) into the reaction chamber receptacle - there should now be approximately 3 cm between the PCR tube and the lamp


- Close the box and tape down the lids
- Extend your hand slightly into the hole where the UV lamp enter the reaction chamber and flip the switch to the right to "LONG WAVE"
- Place outer shield box over the reaction chamber

- Set timer for one hour
- After thirty minutes, turn off the UV lamp, flip the PCR tube so its other side faces up, and turn the UV lamp back on; allow the hour of UV exposure to finish
- When running gel of resulting sample against an uncleaved sample, be sure to cover the gel box with aluminum foil.
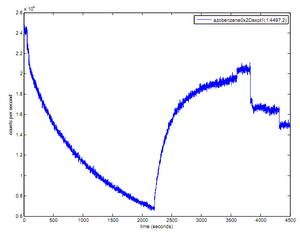
Azobenzene
- Note: this is a draft protocol
- General notes:
- Run rxns at ~1 nM concentration, up to around 10 nM
- Azobenzene strand: 1-2, domain 1 contains azobenzene, domain 2 is a toehold, starts at 50 nM
- Rox fluorophore strand has the sequence of domain 1, starts at 41.3 uM
- RQ quencher strand has the sequence 2*-1*, starts at 43.91 uM
- 1-100 dilutions of Rox and RQ strands were used as starting materials
- excitation 584, emission 602
- when PMT machine is first turned on: wait 30 minutes for PMT to stabilize, should go down to about 90 counts per second
- temp = 30C
- mix by inversion between steps
- Reaction: this was carried out in 1X folding buffer with 12 mM MgCl2.
- 1 mL total reaction volume
- 100 uL 10x buffer
- 1 nM of Rox strand (2.42 uL of 1-100 dil of Rox strand)
- got about 23000 counts per second
- added 5 uL 1-100 dilution of RQ strand
- counts dropped below 8000 after about 30 minutes
- added 20 uL azobenzene strand which led to recovery of counts to about 20000
- fluorimeter stopped this acquisition, so I re-started a new one here
- exposed to short-wave UV at 3 cm distance for 15 minutes in Evan's UV box in the dark
- fluorescence decrease to around 17000 counts (up to time ~500 sec on the second acquisition)
- exposed to visible for 5 minutes in the ambient room light
- measured no increase in fluorescence
- exposed to short-wave UV for another 5-10 minutes in the dark
- fluorescence decrease to around 15000 counts
- exposed to visible for 5 minutes
- no increase in fluorescence
- So, maybe the azobenzene is working and the switch back with visible is just slow at these intensities
- Or maybe we're just chewing up the DNA by exposure to the short-wave UV? but how would that decrease the fluorescence? I'd expect it to lead to unquenching, if anything.
- after use, wash cuvette with water, then ethanol, then water, then ethanol, then dry in an air stream
- 1 mL total reaction volume
- raw data:
- results:
Software Programs
SphereCAD
- See here
Using ImageJ to Analyze Particle Size
- Add 3uL of a dilute sample of your gold nanoparticles (the solution should be a light pinkish color) to a copper grid for imaging under TEM
- Glow discharge isn't specifically necessary in preparing your samples, but may be helpful in the case of a dilute sample
- No staining is necessary, as AuNP alone provide enough contrast, additionally staining may lead to phosphene appearing in your TEM images, which would cause an over-approximation of the size of your particles
- You may either leave the sample until the water dries, or wick off remaining water after about five minutes, either procedure is valid but the first will result in more particles left on your grid (which is a good thing if you placed a highly dilute sample on the grid)
- Take multiple images of the particles at about 100,000x zoom that give you a view of approximately 500 particles:
- It is not essential to use 100,000x zoom, but you do need well focused images and a uniform zoom across all images
- Open your multiple images using stacks in ImageJ
- Set the scale on your images in ImageJ by:
- Zooming in
- Selecting the draw line tool
- Drawing a line over the scale bar
- Clicking analyze, then set scale, then typing into the dialog box the size of your scale bar in the appropriate unit of measurement
- Crop your image to remove the labels at the bottom
- Cropping one image will crop all
- Clear image of weird looking noise by selecting the region and then selecting edit, clear (you will have to do this on each layer)
- Set the contrast by selecting image, adjust, threshold, set contrast, then manipulating the slide bars until it appears that the majority of your particles are highlighted red, click apply, and then make sure that neither of the two checkboxes in the dialog box that appears are checked before proceeding to click okay
- Set the measurement to analyze the size of your particles by clicking analyze, set measurements, and deselecting all check boxes except fit ellipses and ferric diameter
- Analyze the size of your particles by selecting analyze, analyze particles, and then adjusting to an appropriate size range (as square units) that you are expecting for your particle (for 5nm, use 4-400 … aka a 2-20nm particle range), select exclude edges, display results, and show outlines, click OK
- Save the image obtained and export your data table to excel
- Determine the average size of your particles by taking the average of the major axes of your ellipses
- Determine the sphericity of your particles by dividing the average of your minor axes lengths by the average of your major axes lengths (1 will represent a perfect sphere)
- Useful Notes:
- Microsoft Excel 2011 does not come with the capability to create histograms, but this website provides an applet that allows you to do so
<html> <a href="http://www2.clustrmaps.com/counter/maps.php?url=http://openwetware.org/wiki/Biomod/2011/Harvard/HarvarDNAnos:Protocols" id="clustrMapsLink"><img src="http://www2.clustrmaps.com/counter/index2.php?url=http://openwetware.org/wiki/Biomod/2011/Harvard/HarvarDNAnos:Protocols" style="border:0px;" alt="Locations of visitors to this page" title="Locations of visitors to this page" id="clustrMapsImg" /> </a>
</html>