BIOL368/F14:Nicole Anguiano Week 4: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
(→Exploring HIV Evolution In-Class Activity: added activity 1 part 2) |
(→Exploring HIV Evolution In-Class Activity: Added activity 1, part 3) |
||
| Line 4: | Line 4: | ||
===Methods=== | ===Methods=== | ||
=====Activity 1, Part 2===== | =====Activity 1, Part 2===== | ||
*First, I opened the link to the NCBI website and searched for the Markham, et al article. Once found, I switched the search type to nucleotide and searched again using the same search criteria that had led me to the original paper (Markham RB[AUTHOR]). Then, I selected a [http://www.ncbi.nlm.nih.gov/nuccore/AF016818.2 gene] from the list. | *First, I opened the link to the [http://www.ncbi.nlm.nih.gov NCBI website] and searched for the Markham, et al article. Once found, I switched the search type to nucleotide and searched again using the same search criteria that had led me to the original paper (Markham RB[AUTHOR]). Then, I selected a [http://www.ncbi.nlm.nih.gov/nuccore/AF016818.2 gene] from the list. I saved it in fasta format. | ||
*I selected 3 other files, [http://www.ncbi.nlm.nih.gov/nuccore/AF089153.2 one from subject 4], [http://www.ncbi.nlm.nih.gov/nuccore/AF016768.2 one from subject 1], and a second [http://www.ncbi.nlm.nih.gov/nuccore/AF016767.2 from subject 1], and saved all of them in fasta format. | |||
=====Activity 1, Part 3===== | =====Activity 1, Part 3===== | ||
*I navigated to the [http://workbench.sdsc.edu/ Biology Workbench] site and created an account, then entered the workbench and selected "nucleic tools". I selected "Add New Nucleic Sequence" and hit "Run". I uploaded the fasta file I had created before and saved it to my workbench under the label "S2V4C2". | |||
*I then imported the rest of the files that I had saved, and saved them under the same labeling scheme (Subject, Visit, Clone number). | |||
*I selected all of the files I'd uploaded into my workbench, chose the option "ClustalW - Multiple Sequence Alignment", and ran it. I submitted it with the default settings. | |||
=====Activity 2, Part 1===== | =====Activity 2, Part 1===== | ||
=====Activity 2, Part 2===== | =====Activity 2, Part 2===== | ||
===Results=== | ===Results=== | ||
=====Activity 1, Part 3===== | =====Activity 1, Part 3===== | ||
[[Image:Sequence_Alignment_ClustalW.png]] | |||
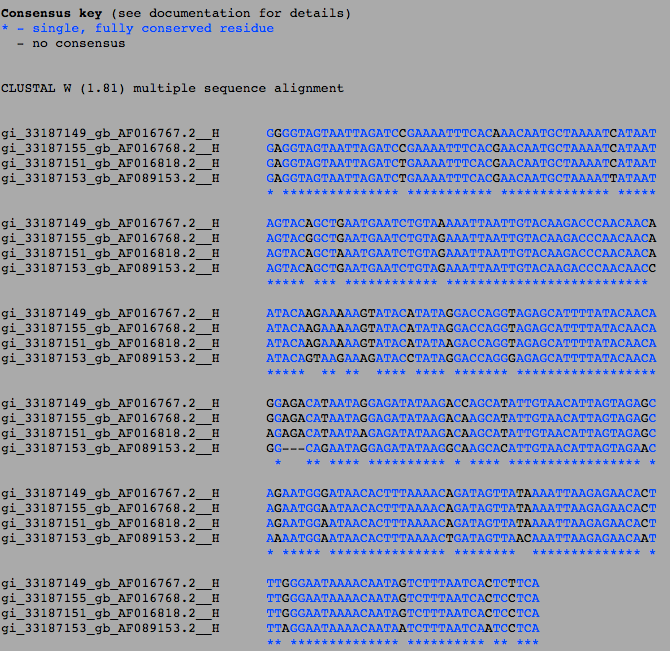
*Figure 1: The results from the sequence alignment from the Clustal W. | |||
*AF016767.2 and AF016768.2 come from subject 1, but AF016768.2 is slightly more similar to AF016818.2 than AF016767.2, having 6 differences with AF016767.2 and 5 differences with AF016818.2 (Fig. 1). AF016767.2 and AF016818.2 are different, having 11 differences between the two of them (Fig. 1). AF089153.2 is the most different from all of them, having 21 locations where it is completely different from the other 3, three of them being areas that do not exist in the gene (Fig. 1). | |||
[[Image:A1P3_ClustalW_Dendrogram.png]] | |||
*Figure 2: The unrooted tree resulting from the ClustalW. | |||
*Of the four genes used, three of the four (AF016768.2, AF016767.2, and AF016818.2) have a relatively high degree of similarity, which can be seen in the unrooted tree. Those three are clustered relatively closely together, with AF016768.2 having a closer similarity to AF016818.2 and AF016767.2 than AF016818.2 and AF016767.2 have with each other. AF089153.2 is far away from the others, indicating a large amount of dissimilarity between it and the other three genes. | |||
=====Activity 2, Part 1===== | =====Activity 2, Part 1===== | ||
=====Activity 2, Part 2===== | =====Activity 2, Part 2===== | ||
| Line 28: | Line 36: | ||
#Which subject of the study was that HIV sequence from? Which section of the record contains information about who the HIV was collected from? | #Which subject of the study was that HIV sequence from? Which section of the record contains information about who the HIV was collected from? | ||
#*The sequence was from subject 2. The "Definition" section of the record contained the information about where the HIV was collected from. | #*The sequence was from subject 2. The "Definition" section of the record contained the information about where the HIV was collected from. | ||
=====Activity 2, Part 1===== | =====Activity 2, Part 1===== | ||
=====Activity 2, Part 2===== | =====Activity 2, Part 2===== | ||
Revision as of 15:25, 17 September 2014
Exploring HIV Evolution In-Class Activity
Methods
Activity 1, Part 2
- First, I opened the link to the NCBI website and searched for the Markham, et al article. Once found, I switched the search type to nucleotide and searched again using the same search criteria that had led me to the original paper (Markham RB[AUTHOR]). Then, I selected a gene from the list. I saved it in fasta format.
- I selected 3 other files, one from subject 4, one from subject 1, and a second from subject 1, and saved all of them in fasta format.
Activity 1, Part 3
- I navigated to the Biology Workbench site and created an account, then entered the workbench and selected "nucleic tools". I selected "Add New Nucleic Sequence" and hit "Run". I uploaded the fasta file I had created before and saved it to my workbench under the label "S2V4C2".
- I then imported the rest of the files that I had saved, and saved them under the same labeling scheme (Subject, Visit, Clone number).
- I selected all of the files I'd uploaded into my workbench, chose the option "ClustalW - Multiple Sequence Alignment", and ran it. I submitted it with the default settings.
Activity 2, Part 1
Activity 2, Part 2
Results
Activity 1, Part 3
- Figure 1: The results from the sequence alignment from the Clustal W.
- AF016767.2 and AF016768.2 come from subject 1, but AF016768.2 is slightly more similar to AF016818.2 than AF016767.2, having 6 differences with AF016767.2 and 5 differences with AF016818.2 (Fig. 1). AF016767.2 and AF016818.2 are different, having 11 differences between the two of them (Fig. 1). AF089153.2 is the most different from all of them, having 21 locations where it is completely different from the other 3, three of them being areas that do not exist in the gene (Fig. 1).
- Figure 2: The unrooted tree resulting from the ClustalW.
- Of the four genes used, three of the four (AF016768.2, AF016767.2, and AF016818.2) have a relatively high degree of similarity, which can be seen in the unrooted tree. Those three are clustered relatively closely together, with AF016768.2 having a closer similarity to AF016818.2 and AF016767.2 than AF016818.2 and AF016767.2 have with each other. AF089153.2 is far away from the others, indicating a large amount of dissimilarity between it and the other three genes.
Activity 2, Part 1
Activity 2, Part 2
Interpretations
Activity 1, Part 2
Activity 1, Part 3
Activity 2, Part 1
Activity 2, Part 2
Questions
Activity 1, Part 2
- What was the accession number of the sequence you chose?
- The Accession number was AF016818.
- Which subject of the study was that HIV sequence from? Which section of the record contains information about who the HIV was collected from?
- The sequence was from subject 2. The "Definition" section of the record contained the information about where the HIV was collected from.
Activity 2, Part 1
Activity 2, Part 2
Conclusions
Activity 1, Part 2
Activity 1, Part 3
Activity 2, Part 1
Activity 2, Part 2
Overall Conclusions
Electronic Lab Notebook
Links
Nicole Anguiano
BIOL 368, Fall 2014
Assignment Links
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 13 Assignment
- Week 15 Assignment
Individual Journals
- Individual Journal Week 2
- Individual Journal Week 3
- Individual Journal Week 4
- Individual Journal Week 5
- Individual Journal Week 6
- Individual Journal Week 7
- Individual Journal Week 8
- Individual Journal Week 9
- Individual Journal Week 10
- Individual Journal Week 11
- Individual Journal Week 12
- Individual Journal Week 13
- Individual Journal Week 15