|
Objective
- Continue laboratory work from yesterday, collecting absorption data for adenosine and inosine solutions
- Calculate calibration curves using linear regression
- Conduct data analysis of the class' data: Means, Standard Deviations, Grubb's test for outliers, 95% and 90% Confidence Intervals
Procedure
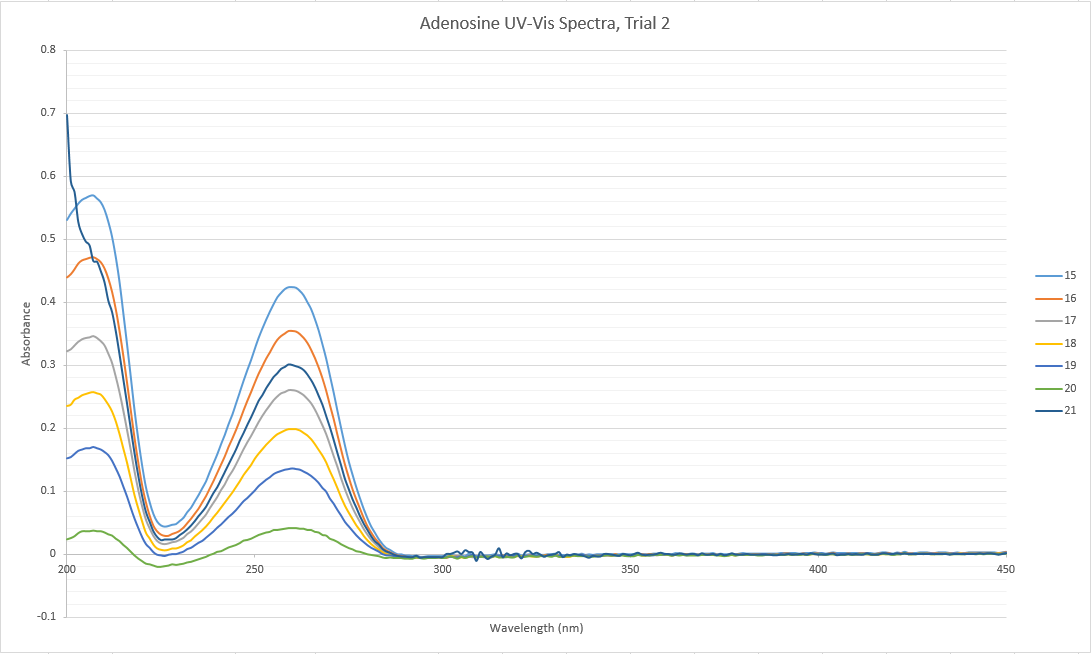
- Procedure from yesterday was used for a second trial of adenosine. A new stock solution and set of dilutions were made. The stock solution was made from .0809 grams of solid adenosine:

Data: CMJ
Data: Group
- Group Inosine Calibration

- Note: None of the data points proved to be outliers for inosine.
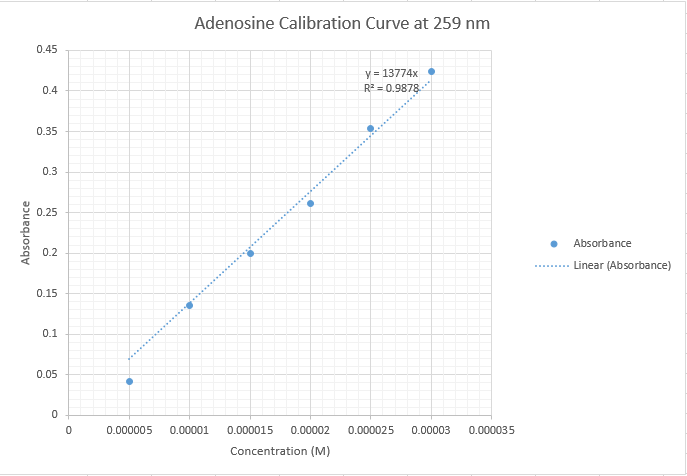
- Group Adenosine Calibration With Outliers

- Group Adenosine Calibration Without Outliers

- Note: Three data points were removed (2×105, 0.406; 2.5×105, 0.442; 3×105, 0.558). Removing the outliers decreased the slope from 14138 to 13411 and improved the r-squared value from .8921 to .9435.
- Statistical Analysis of Group Data


Determining the Unknown Inosine Sample
- A=εbc, therefore c=A/ε where b=1 cm.
- Excel Linear regression analysis calculates the standard error of the Inosine Group Calibration (ε=11007.433792783 M-1cm-1) to be 247.476791358845. Thus, the inherent error of the calibration curve is ±200, resulting in ε=11000±200 M-1cm-1 for inosine at 249 nm.
 - According to the spectrum collected, at 249 nm the unknown has an absorbance of .047.
- c=A/ε --> c=.047/(11000 M-1cm-1) --> c= 4.27×10-6M
- The error in the unknown concentration measurement based on the error inherent in the calibration curve is c*sqrt((200/11000)2). Therefore, the error in unknown concentration measurement is ±.08×10-6M.
- The resulting range is 4.19×10-6M to 4.35×10-6. HAT's actual concentration for this sample was 4.0×10-6M, which does not fall in the range calculated. Our concentration measurement is not in agreement with the actual concentration.
Notes
All of the data collected yesterday was found to be systematically lower than the rest of the class' data. Thus, the data from the inosine trial was not included in the overall group data. Furthermore, a trial 2 of adenosine dilutions was run, using a new stock solution. This second set of data fit with the overall group data, and our first trial was discarded. It is most probable that there was an error in making the first set of stock solutions yesterday that resulted in the incomplete transfer of the massed solids into the 100 mL volumetric flasks. Having a smaller amount of solute in the solvent than calculated would result in systematically lower absorbance results.
|
 Biomaterials Design Lab
Biomaterials Design Lab