Erman's Lab
About Lab Members
Principal Investigator
- Batu Erman, PhD
Current Lab Members
Postdoctoral Fellows
- Ceren Tuncer, PhD
- Belkis Atasever, PhD
- Jitka Eryilmaz, PhD
PhD Students
MSc Students
Previous Lab Members
MSc Students
- Pinar Ayata , Rockefeller University
- Sema Kurtulus, University of Cincinnati
- Ibrahim Aksoylar, University of Cincinnati
- Pinar Onal, Max Plank Institute Berlin
- Serkan Belkaya, UT Southwestern
- Abdalsalam Kmail, Arab American University-Jenin
- Gamze Gunal, Dresden University of Technology
- Meral Ilcim Ozlu, King's College London
Undergraduate students
- Sine Yaganoglu , ETH
- Basak Burcu Cicek , Charite Universitat Medizin
- Emir Tinaztepe , NYU School of Medicine
- Begum Irmak On , Bosphorus University
- Duygu Saatcioglu , University of Uppsala
- Erkin Kuru University of Indiana
Research Topics
T lymphocytes are essential for the functioning of the adaptive immune system. This is demonstrated by genetic or virus induced human immunodeficiencies resulting from a lack of T-cells. T-cells recognize changes in their environment through a specific T-cell receptor (TCR) protein expressed on their surface. This receptor binds to peptide antigens presented in the context of major histocompatibility complex (MHC) proteins on the surface of antigen presenting cells. TCR binding to its ligand causes signaling that re-programs the cell by turning on new genes. TCR first gets expressed on the surface of T cell precursors that also have two co-receptors on their surface named CD4 and CD8. These double positive (DP) thymocytes have the business of generating all the T lymphocytes in the body. Re-programming by new gene expression is important for the development of the thymocyte precursors of T cells in the thymus and for the response of T lymphocytes in peripheral lymphoid organs. We are interested in the structure of the TCR on the surface of T lymphocytes, the cytoplasmic signaling proteins that convey the signal to the nucleus and how new genes get turned on by TCR signals.
Our group is trying to understand how TCR subunits come together to form a receptor that can signal T lymphocytes. The stoichiometry of the TCR is a matter of debate. The widely accepted model of TCR structure is that there is one molecule each of the TCR alpha, beta, CD3 gamma, delta proteins and two molecules each of CD3 epsilon and TCR zeta . Whether TCR stoichiometry is static during different developmental stages and during activation is not known. We are performing fluorescence resonance transfer (FRET) experiments with fluorescently labelled TCR subunits using flow cytometry and confocal microscopy to address the stoichiometry of this receptor complex during and in the absence of signaling. Here we mark one protein with a cyan fluorescent protein (CFP) and another protein with yellow fluorescent protein (YFP). If the two proteins interact, CFP and YFP come together to generate fluorescence with a unique wavelength, which we measure in live cells. We also use a related technique called bioluminescence resonance energy transfer (BRET). Here we tag protein #1 wtih renilla luciferase (rLuc- a bioluminescent protein) and protein #2 with YFP, if and only if protein 1 and 2 interact, the bioluminescent energy generated by rLuc protein is absorbed by the YFP protein, resulting in fluorescence, which we detect in a fluorimeter machine.
The role of the CD3 delta subunit in the TCR is not clearly defined. CD3 delta knockout mice have double positive (DP) T cell precursors that express both CD4 and CD8 molecules, whereas knockouts of the other TCR subunits cannot advance to this stage of development. Morever, we have generated data which demonstrates that TCR lacking CD3 delta can signal in vitro. So what cytoplasmic subunit does this partially defective TCR activate? We are trying to identify if TCR subunits associate with signaling molecules using protein-protein interaction trap experiments. It is difficult to perform classical yeast two hybrid interaction traps with membrane proteins because in such experiments interactions happen in the nucleus rather than near the plasma membrane. So we are using a novel split ubiquitin interaction trap where interactions happen in the plasma membrane. For this purpose we use a novel yeast two hybrid system developed by a Swiss company called Dualsystems. We have identified various membrane proteins that interact with CD3delta, but the real trick is to express all of the TCR subunits in yeast cells. The problem is internal ribosome entry site (IRES) sequences do not seem to work in yeast cells, and we are limited to only one plasmid that has an auxotrophic marker (URA3) for selection in yeast cells, so we are experimenting with different methods to express more than one protein in yeast.
Signaling by the TCR activates various transcription factors that translocate to the nucleus to bind to transcriptional control elements of TCR responsive genes. NF-AT, and NF-kappaB, are two such transcription factors that wait in an inactive form in the cytoplasm before signaling happens. Nuclear translocation of these factors results from cytoplasmic phosphorylation and dephosphorylation events. If non-mamalian species have signaling proteins that are similar to but not identical to NF-AT, and NF-kappaB activating proteins, we may be able to introduce these factors into mammalian cells and modulate TCR signal transduction. We are generating reporter cell lines that transcribe green fluorescent protein (GFP) in response to NF-AT, and NF-kappaB activation. In such cells we introduce plant cDNA's and identify novel factors that modulate TCR signal transduction. Such factors may be used as drugs that activate or inhibit normal T lymphocyte signaling.
One of the genes that is activated by TCR signaling is the gene for the receptor of the pro-survival cytokine, IL-7. IL-7 receptor is expressed in T lymphocytes as well as B lymphocytes. Signaling by this receptor is critical for lymphocyte survival as demonstrated by immunodeficiencies observed in IL7R gene knockout mice and by the X-linked severe combined immunodeficiency (X-SCID) syndrome that develops when a gene encoding a component of the IL7R signaing complex (the cytokine receptor common gama chain) is mutated on the X chromosome. The transcription regulatory regions of the IL7-R gene are not clearly defined, and whether the same transcription factors activate the transcription of this gene in these different lineage cells is not known. We are trying to identify transcriptional enhancers of the IL7-R gene locus using bioinformatics and gene reporter assays. We are modifying the IL7-R gene locus by “recombineering” bacterial artificial chromosomes (BAC) that we re-introduce into tissue culture cell lines that express IL7-R. We perform structure-function studies in re-engineered mutant IL7-R loci to identify the gene regulatory regions.
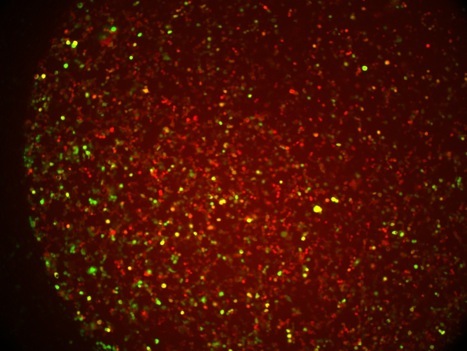
Here are some of our tissue culture cells that express two fluorescent proteins, Jred and TurboGFP. We use these cells as reporters of microRNA’s. (Serkan Belkaya)
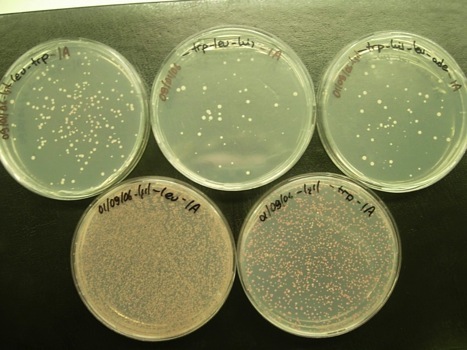
Here are some of the results of our yeast two hybrid screens, where we identify proteins that interact with CD3delta and with c-krox. (Isil Nalbant)
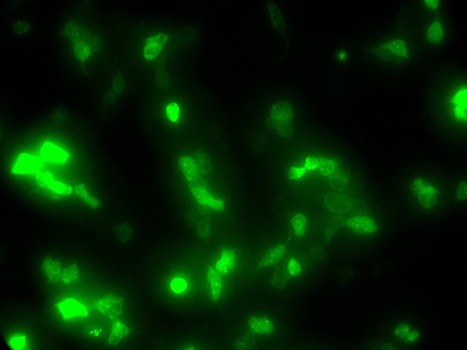
Here are some of our tissue culture cells that fluoresce green because they express a GFP reporter protein, because of the activity of a transcription factor called NFAT2. (Abdalsalam Kmail)
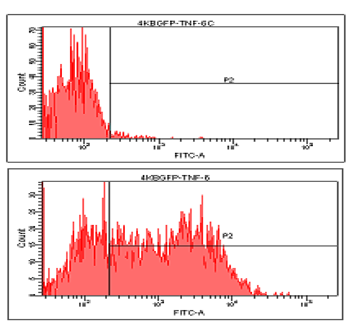
Here are some of our tissue culture cells that fluoresce green because they express a GFP reporter protein, because of the activity of a transcription factor called NFkB. The top histogram is before stimulation, and the bottom histogram is 24 hours after stimulation with TNFalpha. (Belkis Atasever)
Funding Sources
E.C. 6th FW Program Marie Curie IRG
E.C. 6th FW Program Marie Curie RTN(Chromatin Plasticity)
TUBITAK
U.S. National Institutes of Health
Fogarty International Center GRIP Fellowship
Fogarty International Research Collaboration
Award FIRCA