Registry of Standard Biological Models/Basic Component Models/Constitutive Promoter
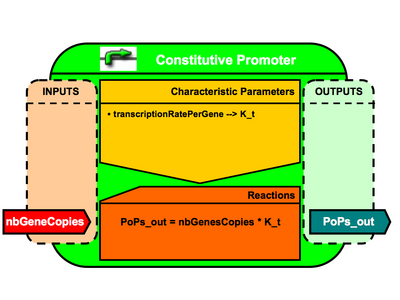
Constitutive Promoter Architecture
CellML structure (CellML 1.1 spec)
- Component: ConstitutivePromoter
- Units:
- TBD
- Variables:
- nbGeneCopies (public interface = in / init value = 1.0)
- transcriptionRatePerGene (public interface = none / init value = XXX)
- PoPs_OUT (public interface = out / init value = 0.)
- MathML
- <amsmath>PoPsOUT = nbGenesCopies*transcriptionRatePerGene</amsmath>
CellML File
<syntax type='xml'> <?xml version="1.0"?>
<model xmlns="http://www.cellml.org/cellml/1.0#"
xmlns:cmeta="http://www.cellml.org/metadata/1.0#" xml:base="file:///C:/CellML_models/constitutive_promoter.cml" cmeta:id="constitutive_promoter" name="constitutive_promoter">
<component name="constitutive_promoter">
<variable name="mRNAsynthesisRate" initial_value="" public_interface="out" units="moles_per_second"/> <variable name="nb_gene" initial_value="1" public_interface="none" units="dimensionless"/> <variable name="mRNAsynthesisRatePerGene" initial_value="1" public_interface="none" units="moles_per_second"/>
[math]\displaystyle{ <apply id="mRNAsynthesisRate"> <eq/> <ci>mRNAsynthesisRate</ci> <apply> <times/> <ci>mRNAsynthesisRatePerGene</ci> <ci>nb_gene</ci> </apply> </apply> }[/math]
</component>
<import xmlns:xlink="http://www.w3.org/1999/xlink"
xlink:href="file://C:\CellML_models\units.cml">
<units name="moles_per_second" units_ref="moles_per_second"/>
<units name="per_second" units_ref="per_second"/>
</import>
</model>
</syntax>